A Recurrent ERCC3 Truncating Mutation Confers Moderate Risk for Breast Cancer
|
|
|
- Josephine Joseph
- 5 years ago
- Views:
Transcription
1 Published OnlineFirst September 21, 2016; DOI: / CD A Recurrent ERCC3 Truncating Mutation Confers Moderate Risk for Breast Cancer Joseph Vijai1,2, Sabine Topka1,2, Danylo Villano1,2, Vignesh Ravichandran1,2, Kara N. Maxwell3, Ann Maria1,2, Tinu Thomas1,2, Pragna Gaddam1,4, Anne Lincoln1,4, Sarah Kazzaz1,2, Brandon Wenz3, Shai Carmi5, Kasmintan A. Schrader6, Steven N. Hart7, Steve M. Lipkin8, Susan L. Neuhausen9, Michael F. Walsh1,4,10, Liying Zhang11, Flavio Lejbkowicz12, Hedy Rennert12, Zsofia K. Stadler1,4,8, Mark Robson1,4,8, Jeffrey N. Weitzel13, Susan Domchek3,14,15, Mark J. Daly16,17, Fergus J. Couch7,18, Katherine L. Nathanson3,14,15, Larry Norton1, Gad Rennert12, and Kenneth Offit1,2,4,8 ABSTRACT Known gene mutations account for approximately 50% of the hereditary risk for breast cancer. Moderate and low penetrance variants, discovered by genomic approaches, account for an as-yet-unknown proportion of the remaining heritability. A truncating mutation c.325c>t:p.arg109* (R109X) in the ATP-dependent helicase ERCC3 was observed recurrently among exomes sequenced in BRCA wild-type, breast cancer affected individuals of Ashkenazi Jewish ancestry. Modeling of the mutation in ERCC3-deficient or CRISPR/Cas9-edited cell lines showed a consistent pattern of reduced expression of the protein and concomitant hypomorphic functionality when challenged with UVC exposure or treatment with the DNA alkylating agent IlludinS. Overexpressing the mutant protein in ERCC3-deficient cells only partially rescued their DNA repair deficient phenotype. Comparison of frequency of this recurrent mutation in over 6,500 chromosomes of breast cancer cases and 6,800 Ashkenazi controls showed significant association with breast cancer risk (ORBC = 1.53, ORER+ = 1.73), particularly for the estrogen receptor positive subset (P < 0.007). SIGNIFICANCE: A functionally significant recurrent ERCC3 mutation increased the risk for breast cancer in a genetic isolate. Mutated cell lines showed lower survival after in vitro exposure to DNA-damaging agents. Thus, similar to tumors arising in the background of homologous repair defects, mutations in nucleotide excision repair genes such as ERCC3 could constitute potential therapeutic targets in a subset of hereditary breast cancers. Cancer Discov; 6(11); AACR. 1 Department of Medicine, Memorial Sloan Kettering Cancer Center, New York, New York. 2Cancer Biology and Genetics Program, Memorial Sloan Kettering Cancer Center, New York, New York. 3Division of Hematology/ Oncology, Department of Medicine, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, Pennsylvania. 4Clinical Genetics Service, Department of Medicine, Memorial Sloan Kettering, New York, New York. 5Braun School of Public Health and Community Medicine, The Hebrew University of Jerusalem, Jerusalem, Israel. 6British Columbia Cancer Agency, Canada s Michael Smith Genome Sciences Centre, Vancouver, Canada. 7Department of Health Sciences Research, Mayo Clinic, Rochester, Minnesota. 8Department of Medicine, Weill Cornell Medical College, New York, New York. 9Department of Population Sciences, Beckman Research Institute of City of Hope, Duarte, California. 10Department of Pediatrics, Memorial Sloan Kettering Cancer Center, New York, New York. 11Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, New York. 12Clalit National Israeli Cancer Control Center and Department of Community Medicine and Epidemiology, Carmel Medical Center and B Rappaport Faculty of Medicine, Haifa, Israel. 13Clinical Cancer Genetics (for the City of Hope Clinical Cancer Genetics Community Research Network), City of Hope, Duarte, California. 14 Division of Translational Medicine and Human Genetics, Department of Medicine, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, Pennsylvania. 15Abramson Cancer Center, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, Pennsylvania. 16Broad Institute of Harvard and Massachusetts Institute of Technology, Cambridge, Massachusetts. 17Center for Human Genetic Research and Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts. 18Department of Laboratory Medicine and Pathology, and Health Sciences Research, Mayo Clinic, Rochester, Minnesota. Note: Supplementary data for this article are available at Cancer Discovery Online ( J. Vijai and S. Topka share first authorship of this article. Corresponding Author: Kenneth Offit, Memorial Sloan Kettering Cancer Center, 1275 York Avenue, Box 192, New York, NY Phone: ; Fax: ; offitk@mskcc.org. doi: / CD American Association for Cancer Research. NOVEMBER 2016 CANCER DISCOVERY 1267
2 Vijai et al. INTRODUCTION Genetic susceptibility to breast cancer has been shown to be strongly associated with rare coding gene mutations and common noncoding genomic variants ( 1 ). Loss-of- function mutations in well-characterized genes, such as those in BRCA1/2 and other members of the homologous recombination pathway, are routinely used for clinical risk assessment in cancer, but account for a small fraction of excess familial risk ( 2 ). Common single-nucleotide polymorphisms (SNP) are yet to demonstrate clinical utility, but over 90 SNPs account for over 37% of familial relative risk ( 3 ). An as-yet-to-be-defined proportion of the remaining familial risk is accounted for by variants of moderate risk, including coding or noncoding variants in the DNA repair pathways such as nucleotide excision repair (NER), mismatch repair (MMR), and base excision repair (BER), which play an important role in checkpoint and genomic integrity maintenance ( 4, 5 ). In addition to serving as candidate cancer susceptibility genes, members of the NER pathway also play a critical role in DNA-damage repair caused by chemotherapeutic agents and radiation exposure. ERCC3 is an ATP-dependent DNA helicase that is part of the TFIIH transcription factor complex. In this report, we characterize the functions of a recurrent founder mutation in ERCC3 and demonstrate its association with breast cancer risk. RESULTS Identification of a Recurrent ERCC3 R109X Mutation in Ashkenazi Probands with BRCA1/2 Wild-Type Breast Cancer Exome sequencing was carried out on DNA extracted from peripheral blood/saliva of 46 early-onset (<45 years) and 13 familial BRCA wild-type (WT) breast cancer probands (33 with known ER-positive status) of Ashkenazi Jewish ancestry. We filtered for rare protein truncating variants using public data sources such as the Exome Sequencing Project (ESP), 1000 Genomes, and the Exome Aggregation Consortium (ExAC) without The Cancer Genome Atlas. We further filtered for recurrent mutations in the DNA-repair genes with low background mutation load using the smallest residual variation intolerance score percentage ( 6 ) amongst 70 genes, calculated based on the ExAC data. We observed three individuals in the same BRCA WT kindred ( Fig. 1 ) with the same protein truncating mutation, R109X in ERCC3 (HGMD: c.325c>t: p.r109*; chr2: ; rs ). Sanger sequencing was used to confirm the next-generation sequencing findings (shown in Supplementary Fig. S1). Two of the siblings were affected with ER-positive breast cancer, and the third was a male with breast cancer. The same mutation was also observed in two other individuals at relatively young age (<40 years), from unrelated kindreds with multiple cases of breast cancer. Details of the families and cancer incidence are described in Supplementary Table S1. The variant is almost absent in other world populations, and relatively rare in Caucasians in public data sources such as the ExAC Consortium (Non-Finnish EUR MAF = ; ESP-EUR MAF = ). Analysis of the structure of the first 300 amino acids of ERCC3 shows that R109 is most likely part of a righthanded alpha helix (Supplementary Fig. S2). However, no crystal structure is available for high confidence prediction. Phenotype Rescue by Complementation Assay To understand whether this variant has a deleterious effect on the gene function, we carried out a series of in vitro functional assays. Using a previously well-characterized ERCC3- deficient cell line derived from a patient with combined xeroderma pigmentosum/cockayne syndrome (XP/CS; ref. 7 ), phenotype rescue after DNA damage was assessed by cell viability assay. The parental XPCS2BA cell line and derived lines stably overexpressing the ERCC3 R109* mutant (R109X) or the WT ERCC3 ( Fig. 2A and B ) showed varying degrees of sensitivity to DNA damage inducing agents such as UVC and a fungal sesquiterpene IlludinS. We observed that cells overexpressing ERCC3 R109X showed significantly higher cell death compared with the ERCC3 WT, whereas the untransfected cells all I Lung 88 Lung 67 Breast 66 Pancreas Pancreas II Ovarian 60 Breast 74 Breast 68 Breast 74 Breast 56 R109X R109X R109X Figure 1. Identification of germline mutations in ERCC3 in a family with multiple breast cancer cases. Sequencing was performed on three individuals affected with breast cancer, confirming identification of ERCC3 R109X in all three affected siblings. Pathology reports for individuals II-3 and II-5 show both with well-differentiated (low-grade) invasive ductal carcinoma diagnosed at Stage IA. One of the tumors was ER + PR + HER2 + (II-3) and the other one was ER + PR HER2 (II-5). Double-digit numbers indicate age at diagnosis. For individuals I-4 and I-5, age at diagnosis was not available CANCER DISCOVERY NOVEMBER
3 ERCC3 R109X Mutation and Breast Cancer A Relative transcript level 1, D E F Relative cell viability XPCS2BA WT R109X Relative luciferase signal B 1.5 XPCS2BA WT R109X XPCS2BA WT WT ERCC3 (89 kda) R109X R109X (~12 kda) GAPDH Time after UVC XPCS2BA irradiation XPCS2BA WT ERCC3 WT WT R109X R109X pchk1 γ-h2ax ERCC3 R109X ,000 UVC [J/m 2 ] UVC [J/m 2 ] C Relative cell viability IlludinS [ng/ml] GAPDH XPCS2BA WT R109X Figure 2. Functional evaluation of the ERCC3 mutant via overexpression in an ERCC3-deficient cell line. A, transcript and B, protein levels in XPCS2BA parental cell line and cell lines stably overexpressing a lentiviral construct containing WT ERCC3 (WT) or the R109X mutant (R109X) cdna. Expression from the mutant cdna produces a truncated protein fragment of about 12kDa. C D, relative cell viability of XPCS2BA and ERCC3 WT and mutant overexpressing cell lines at 72 hours following treatment with increasing doses of IlludinS ( C ) or UVC ( D ). E, host cell reactivation assay showing reduced DNA repair ability of the mutant as opposed to WT ERCC3 -overexpressing cell lines. Data represent the mean of three experiments with error bars representing the SD ( C ) or SEM ( D and E ). F, phosphorylation of H2AX and CHK1 in response to UVC-induced DNA damage. XPCS2BA, WT, or R109X cell lines were harvested at different time points following exposure to 20 J/m 2 UVC, and activation of H2AX and CHK1 was assessed by Western blotting. died upon exposure to higher doses of both IlludinS and UVC ( Fig. 2C and D ). Exposure at 2 ng/ml of IlludinS ( Fig. 2C ), a fungal toxin from the Jack O Lantern mushroom ( Omphalotus illudens ) that creates DNA damage, which blocks transcription with demonstrated antitumor activity ( 8 ), showed that 3%, 42%, and 77% of the XPCS2BA, R109X, and WT cells survived, respectively ( P < ). At a dose of 4 ng/ml, most of the WT cells survived, demonstrating that the mutant cells were transiently and partially able to withstand UV-induced DNA damage, while failing to adequately repair DNA damage to ensure survival. Similarly, following UVC irradiation at a dose of 10 J/m 2, 72 hours after exposure, 42% of the R109X cells survived compared with 63% of WT cells ( P < ). At a dose of 4 ng/ml, these numbers reduced more drastically for the mutant, showing the same trend as the IlludinS exposure experiment (Fig. 2D ). Host Cell Reactivation Assay Using an exogenous source of DNA, namely a luciferase reporter plasmid, we measured the ability of intact live cells to repair DNA damage. Through measurement of the reporter, the extent of reactivation of the damaged plasmid is quantitated and is a readout of DNA-repair activity. Relative luminescence measured after cotransfection of the reporter plasmid (exposed to 600 J/m 2 ) showed a 1.5-fold difference between mutant and WT overexpression cell lines, suggesting a markedly reduced reporter reactivation in the R109X ( Fig. 2E ; P < ). We observed almost no reactivation in the parental cell line at this dose. Activation of the DNA-Damage Response Pathway Following UVC Irradiation DNA-damage response is marked by activation of H2AX (γh2ax) and leads to the induction of cell-cycle checkpoint initiation by activation of CHK1. To explore the ability of ERCC3 R109X to trigger the checkpoint cascade in response to DNA damage, we examined γh2ax and phosphorylated CHK1 following UVC-mediated DNA-damage induction in XPCS2BA cells. UV-induced phosphorylation of H2AX is reduced in cells deficient in the nucleotide excision repair pathway ( 9 ). In agreement with this, we found lower levels of activated H2AX in the XPCS2BA cell line compared with cells where WT ERCC3 expression had been restored. In the mutant, we observed lower γh2ax levels, almost as low as in the parental cell line. Activation of the checkpoint kinase CHK1 was also strongly reduced in the mutant as compared with the WT-overexpressing cells, and similar to the untransfected XPCS2BA ( Fig. 2F ). Functional Characterization of Genome-Edited ERCC3 Mutants in Human Mammary Epithelial Cells Overexpression of ERCC3 -mutant constructs does not recapitulate physiologic levels of protein in a germline heterozygous mutation state within cells. Therefore, using CRISPR/ Cas9, we engineered several heterozygous mutations in the mammary epithelial cell line HMLE that mimic the site of the originally discovered recurrent mutation in individuals NOVEMBER 2016 CANCER DISCOVERY 1269
4 Vijai et al. A B Relative transcript level 1.5 HMLE P106fs V107fs T111fs R109X **** ERCC3 **** **** **** GAPDH 0.0 HMLE P106fs V107fs T111fsB5 R109X C D E Relative cell viability 1.5 +/+ +/P106fs +/V107fs +/T111fs +/R109X IlludinS [ng/ml] Relative cell viability 1.5 +/+ 150 ERCC3 CRISPR ERCC3 CRISPR + WT 100 * IlludinS [ng/ml] Relative % γh2ax-positive cells 50 0 HMLE HMLE ERCC3 CRISPR * 6h 12h 16h Figure 3. Modeling of ERCC3 R109X by CRISPR/Cas9 in a mammary epithelial cell line and functional analysis. A, ERCC3 transcript and ( B ) protein levels in HMLE and CRISPR/Cas9-edited cell lines modeling the heterozygous R109X mutation. C, relative cell viability of HMLE control and CRISPRedited cell lines at 72 hours following treatment with increasing doses of IlludinS. Data represent the mean of three experiments with error bars representing the SEM. D, relative cell viability of HMLE control and combined CRISPR-edited cell lines (with and without reexpression of the WT ERCC3) following treatment with IlludinS. E, quantification of phosphorylated H2AX by flow cytometry in HMLE control and CRISPR-edited cell lines harvested at different time points after DNA damage. Asterisks refer to the P values determined by a 2-way ANOVA comparing the reduction in H2AX fluorescence at the respective time points between the WT HMLE and CRIPSR-edited HMLE cell lines ( P 0.05). with breast cancer (Supplementary Table S2; Supplementary Fig. S3). Quantitative real-time PCR of ERCC3 transcripts (exons 1, 2 and exons 9, 10) showed relative transcript levels reduced by half in these cells when compared with the parental HMLE cell line ( Fig. 3A ). Western blotting also showed the reduction in total ERCC3 protein levels ( Fig. 3B ), suggesting that in the CRISPR clones, ERCC3 is transcribed mainly from the remaining WT allele. Because we did not observe any homozygous mutations amongst the surviving CRISPR clones, we are unsure if a homozygous mutation is viable. The ERCC3 CRISPR clones generally showed substantial reduction in relative cell viability 72 hours after exposure to IlludinS ( Fig. 3C ). At 2 ng/ml IlludinS, all the parental HMLE cells survived, whereas the CRISPR edited cell lines showed significantly reduced survival ( P < ). These data suggest that the mutants function at a much lower efficiency compared with WT and hence may be classified as hypomorphs. To demonstrate that the observed effect of IlludinS on the viability of the CRISPR-edited HMLE is specific to the resulting ERCC3 deficit, we performed rescue experiments by overexpressing WT ERCC3 in these cells. We observed that the ERCC3 CRISPR cell lines, after stable overexpression of WT ERCC3, showed the same response to IlludinS as the WT HMLE cell line ( Fig. 3D ), thereby showing complete rescue of the phenotype, confirming that this phenotype was indeed a result of the engineered ERCC3 deficiency. To assess the induction and removal of phosphorylated H2AX in response to DNA damage, we performed immunostaining and flow cytometric analysis of γh2axpositive cells following treatment with IlludinS. Although this led to a similar fold induction of γh2ax-positive cells in the WT and CRISPR clones (not shown), the reduction in γh2ax was significantly lower in the ERCC3 CRISPR mutants, indicating less efficient DNA-damage repair in these cells ( P < 0.05; Fig. 3E ). ERCC3 R109X as a Risk Allele for Breast Cancer in Ashkenazim Using breast cancer affected individuals and controls from the Memorial Sloan Kettering Cancer Center (MSKCC), the University of Pennsylvania, and the Clalit National Israeli Cancer Control Center, we performed a matched case control association of the ERCC3 variant using TaqMan genotyping and sequencing across 3,286 cases and 2,716 controls. A third group of 705 controls was sourced from The Ashkenazi Genome Consortium (TAGC) project (ref. 10 and in preparation ) control resource ( Table 1A ). All control groups showed similar allele frequencies. A total of 101 heterozygote carriers were present in the combined data. A 1.53-fold increased risk was observed for breast cancer (OR = 1.53, lower CI = 7; P = 0.023) after examining over 6,500 chromosomes in cases and 6,800 chromosomes in controls ( Table 1A ). A stronger 1270 CANCER DISCOVERY NOVEMBER
5 ERCC3 R109X Mutation and Breast Cancer Table 1A. Association of ERCC3 R109X risk allele to breast cancer Center Cases Controls Ca-Alt Ca-Ref Ctrl-Alt Ctrl-Ref P OR Lower CI MSKCC/PENN 1,968 1, , ,248 CLALIT 1,318 1, , ,152 TAGC ,401 Total 3,286 3, , , Table 1B. Association of ERCC3 R109X risk allele to ER-positive breast cancer Center Cases Controls Ca-Alt Ca-Ref Ctrl-Alt Ctrl-Ref P OR Lower CI MSKCC/PENN 1,288 1, , ,248 CLALIT 992 1, , ,152 TAGC ,401 Total 2,280 3, , , association was observed with the ER-positive subtype (OR = 1.73, lower CI = 1.19; P = 0.007; Table 1B ). The majority of the tumors in the carriers had an ER-positive status. These data show that the ERCC3 c.325 T allele is a moderate risk factor for breast cancer in individuals of Ashkenazi ancestry. Unphased haplotype analyses from the TAGC heterozygote carriers suggested a founder mutation (Supplementary Fig. S4). In the chromosomal region 2q14.3, a 3.4 cm long haplotype was observed in the carriers which was significantly longer than in noncarriers ( P < ). DISCUSSION Here, we identified a truncating germline mutation in the first putative helicase domain of the DNA-repair gene ERCC3 and report a strong genetic association with risk of breast cancer in individuals of Ashkenazi Jewish ancestry. The R109X variant, seen in 1.83% of Ashkenazi Jewish individuals with breast cancer and 2.06% in the ER-positive subset, confers moderate risk (OR BC = 1.53; OR ER+ = 1.73). The same mutation was also observed twice as a somatic event in a breast carcinoma and a soft-tissue sarcoma (Supplementary Fig. S5A). Mutations within the ERCC3 helicase domain are also often seen in tumors such as melanoma, in addition to non small cell lung cancer and colorectal, esophagogastric, and bladder cancers. In general, ERCC3 is seldom disrupted in somatic tissue by genomic integrity loss such as amplification or deletions (Supplementary Fig. S5B). In the LOVD database, the variant was seen in 6 of 8,600 individuals of European ancestry, a similar frequency to public databases. As a core component of the TFIIH basal transcription factor, ERCC3 ATP-dependent DNA helicase has key functions in both RNA transcription by RNA polymerase II and in NER following DNA damage ( 11 ). It has previously been associated with Mendelian DNA repair disorders such as XP, XP/CS, and trichothiodystrophy (TTD) with hallmarks of increased photosensitivity, cancer predisposition, and impaired development ( 12, 13 ). Hypomorphic mutations in the DNA-damage repair gene NBN cause the autosomal recessive chromosomal instability disorder Nijmegen breakage syndrome (NBS) in homozygous individuals, but have also been shown to lead to increased cancer incidence in heterozygous relatives of patients with NBS, thus providing a plausible example of a hypomorphic mutation that is also a susceptibility gene ( 14, 15 ). Similarly, ERCC3 R109X behaves as a hypomorph in our functional assays. In experiments using an ERCC3-deficient cell line, R109X was partially successful at phenotype rescue, with the cells exhibiting intermediate repair capability, suggesting that they are more likely to harbor a second hit following genotoxic events. Importantly, ERCC3 transcript expression was shown to be lower in the proband s cells compared with a non mutation carrier (Supplementary Fig. S6). Dosage dependency of key molecular components of the DNAdamage repair pathway, such as haploinsufficiency, has been previously demonstrated ( ), leading to tumorigenesis in in vitro and in vivo models of H2AX, BLM, and CHEK1. ERCC3 homozygous knockout mice have been shown to be embryonic lethal, indicating that the gene is necessary for development ( 21 ). Additionally, very few ERCC3 mutations have been reported, even amongst patients with XP, suggesting the gene is intolerant to common mutational mechanisms. Analysis of gene-based mutability from the ESP and ExAC exome data has shown that quantitative metrics that predict gene conservation and mutation tolerance rank ERCC3 as bearing low background mutational load ( 22 ). In light of the rarity of known mutations in ERCC3, there have thus far been no studies elucidating cancer susceptibility in heterozygous carriers. The only mouse model that has been described modeled the XP/CS hereditary DNA repair deficiency syndromes ( 21 ). Animals with heterozygous knockout of the NER genes XPC and XPE/ DDB2 showed increased cancer incidence after exposure to UV irradiation ( 23, 24 ), whereas heterozygous XPC knockout mice also showed elevated frequency of spontaneous NOVEMBER 2016 CANCER DISCOVERY 1271
6 Vijai et al. mutations as a function of age ( 25 ). Also, a mouse model harboring a mutation in the other helicase of the TFIIH complex, XPD, showed a strongly increased cancer incidence in response to UV exposure ( 26 ). This is the first study that shows a specific truncating mutation in ERCC3 conferring increased risk of breast cancer. However, polygenes in DNA repair and/or other pathways, acting epistatically with ERCC3, are likely contributors to and modifiers of the heritable risk of breast cancer, complicating attempts to demonstrate cosegregation of single variants in multiplex kindreds. We have previously demonstrated a modestly elevated breast cancer risk associated with specific mutations of CHEK2 and APC in the Ashkenazi Jewish genetic isolate ( 27, 28 ), and other recurring mutations of CHEK2, HOXB13, PALB2, and RAD51C have been variably associated with breast cancer risk in diverse populations ( ). Although ERCC3 R109X is more frequent in Ashkenazi Jews, we have observed it as well as other ERCC3 mutations in non- Ashkenazi individuals (data not shown), but because of the rarity of these ERCC3 mutations, further population-based studies will be required to assess their role as breast cancer susceptibility alleles. Although the relative risks of 1.5 to 2.0 shown here are higher than those associated with most genome-wide association study SNPs, the clinical utility of variants associated with risks in this range remains to be established. Nonetheless, these discoveries provide insight into molecular pathways of breast cancer pathogenesis and potentially identify therapeutic targets ( ). The enhanced susceptibility of ERCC3 -mutant cells to reagents of the fungal sesquiterpene class, demonstrated here, suggests additional in vivo studies of the therapeutic efficacy versus toxicity of these compounds against breast tumors in an ERCC3 -mutant genetic background. METHODS Next-Generation Sequencing of Breast Cancer Probands and Controls One microgram of germline DNA extracted from peripheral blood was used for whole-exome capture using the Agilent SureSelect 38-Mb paired-end sequencing with the Illumina HiSeq Sequence reads in the form of FASTQ files were aligned to the human decoy reference (GRCh37) to generate BAM files using Burrows-Wheeler Aligner (BWA) v Picard tools were used for quality metric calculation and marking duplicate reads. The Genome Analysis Tool Kit (GATK) version g37228af was used for variant calling using the haplotype caller algorithm. Variant quality score recalibrated (VQSR) data were used for filtering variants. Variant-level and interval-level annotations were performed using SNPEff, ANNOVAR, and CAVA programs. Downstream analysis consisted of filtering out low-quality variant calls and those already reported as common in public databases. The TAGC has sequenced the whole genomes of 128 and 577 individuals of Ashkenazi Jewish ancestry using the Complete Genomics and Illumina X10 sequencing platform at the New York Genome Center, respectively. The paired-end libraries were generated using the TruSeq DNA Nano Kit, sequenced to an average depth of 30, and reads were aligned using a standard pipeline involving BWA version and GATK version Extensive quality control is performed on all whole-genome sequencing samples, including alignment rates (>97%), median and mean library insert size (>350bp), percent duplication (typically <20%), mean genome coverage (>30 ) and uniformity of coverage, TiTv and Het-Hom ratios. An automated concordance check is also performed against the SNP array genotyping data, to ensure against sample swap at any step during the sequencing process, and to further validate the quality of the single-nucleotide variant calls from the sequencing data. All research participants provided written consent to an Institutional Review Board approved research protocol to allow for the collection and use of biospecimens. Haplotype Analysis Using only high-quality biallelic SNPs, the haplotype lengths carrying the ERCC3 R109X mutation were calculated to either side of the mutation in the TAGC carriers as the length until an opposite homozygous genotype. This is an overestimate, due to the lack of accurate phasing information. The mean length was 3.4 cm, estimated by using the HapMap recombination rates for the region. The mean lengths of haplotypes shared between noncarriers were also computed. The significance of the difference between the distributions of haplotype lengths at the carriers and at 100 random noncarriers was calculated using the Kolmogorov Smirnov test. Sanger Sequencing Sanger sequencing of the ERCC3 R109X variant was performed using the following primers: ERCC3_F1: CATGGAGCACCTATGCCTATT ERCC3_R1: CTGCAACTCATGTTTCCTTGTC TaqMan Genotyping The allelic discrimination assay C _10 for genotyping was done using TaqMan (Life Technologies). The assay was run on an ABI HT7900 machine and automatically clustered and manually reviewed. Confirmed heterozygotes were run as positive controls, and duplicate concordances were checked per plate. Statistical Analyses Allele counts were tabulated from the TaqMan genotyping after QC. Statistical analysis was performed using the R statistical package using the fisher.test module. Because the truncating genetic variant was hypothesized and shown by functional assays to lead to increased risk associated with reduced DNA repair efficiency, we report onesided Fisher exact test results. Nevertheless, two-sided tests for both breast cancer and ER status were also significant at the 5% level (breast cancer overall: P = 0.036, OR = 1.53, 95% CI = ; ER + : P = 0.011, OR = 1.73, 95% CI = ). Plasmids The pentr221 plasmid containing human ERCC3 ORF was purchased from TransOMIC. The ERCC3 ORF was cloned into the plx302 lentiviral expression plasmid, a gift from David Root, Broad Institute of MIT and Harvard (Addgene # 25896). The ERCC3 R109X mutant was generated from the WT ERCC3 plasmid using the QuickChange II XL Site-Directed Mutagenesis Kit (Agilent). pspcas9(bb)-2a-gfp (PX458) was a gift from Feng Zhang, Broad Institute of MIT and Harvard (Addgene # 48138). Guide RNAs were designed using the CRISPR Design Tool ( 37 ). The 24-mer oligonucleotides were synthesized (ref. 37 ; Integrated DNA Technologies), annealed, and cloned into px458. Cell Culture and Transfections The HMLE cell line, established in the laboratory of Robert A. Weinberg at the Whitehead Institute for Biomedical Research, Cambridge, MA, was grown in mammary epithelial growth medium (MEGM) and supplements as recommended by Lonza. The XPCSBAsv40 cell line was grown in RPMI-1640 HEPES medium (Invitrogen), supplemented with 10% FBS and 1% penicillin streptomycin and glutamate. HEK293T cells (ATCC; CRL-3216) were maintained in 1272 CANCER DISCOVERY NOVEMBER
7 ERCC3 R109X Mutation and Breast Cancer DMEM supplemented with 10% FBS and 1% penicillin streptomycin and glutamate. The HMLE and HEK293T lines were kindly provided by Robert Benezra (MSKCC), and the XPCSBA-sv40 cell line was a generous gift from Jan Hoeijmakers (Erasmus MC, Rotterdam, the Netherlands). The cell lines were not further authenticated. Cell cultures were maintained in a humidified incubator at 37 C in 5% CO 2 and tested for Mycoplasma on a monthly basis. Transfections were carried out with the Amaxa Cell Line Nucleofector Kit V (Lonza) and Nucleofector IIb device according to the manufacturer s instructions. Viral vectors, cotransfected with pspax2 and pseudotyped with VSV-G, were produced in HEK293T cells using the Lipofectamin2000 transfection reagent. The virus supernatant was concentrated by centrifugation for 90 minutes at 20,000 RPM at 4 C, and pellets were dissolved in OptiMEM (Gibco). Transduction of cells with virus supernatant was carried out in the presence of 8 μg/ml polybrene. Stably transfected cell lines were generated by selection with μg/ml puromycin. Cells cotransfected with pmaxgfp (Lonza) and the px458-sgrna plasmids, containing guide RNAs targeting the ERCC3 locus in close proximity to the ERCC3 mutation site (chr2: ) with or without a repair template (ssodn) containing the specific mutation, were sorted as single cells into 96-well plates based on GFP fluorescence using a BD FACSAria cell sorter. Screening of CRISPR/Cas9-Edited Cell Lines Single-cell clones generated from the transfection mentioned above were expanded and genomic DNA extracted using the Quick- Extract DNA Extraction Solution (Epicentre). The genomic region surrounding the target site was amplified using the following primer sequences: ERCC3_SURV F, 5 -TGTGGTGTTGGGCAGCTTAT-3 ; ERCC3_SURV R, 5 -ACACTCACTTTGGGCTGCAT-3 The purified PCR products were subjected to Sanger sequencing. Real-Time PCR RNA was extracted 24 hours after transduction using the RNeasy Mini Kit (Qiagen) and reverse transcribed with the ReadyScript cdna Synthesis Mix (Sigma-Aldrich). Quantitative real-time PCR analyses were performed on an ABI PRISM 7900HT Sequence Detection System using the Power SYBR Green PCR Master Mix (Life Technologies) according to the manufacturer s instructions. Following initial incubation for 10 minutes at 95 C, amplification was performed for 40 cycles at 95 C for 15 seconds and 60 C for 1 minute. The RPL32 gene was used as the internal standard. Analysis was performed based on the comparative CT method. Values reported are mean of triplicate experiments. The following primer sequences were used: RPL32 F, 5 -CATCTCCTTCTCGGCATCA-3 ; RPL32 R, 5 -AACCCTGTTGTCAATGCCTC-3 ; ERCC3 F1, 5 -GTCCGCGAAGATGACAAAATTG-3 ; ERCC3 R1, 5 -AATTCAGGAGACATAGGGCAC-3 ; ERCC3 F3, 5 -ATGGGCAAAAGAGACCGAG-3 ; ERCC3 R3, 5 -CTGACTCATCCACCTGCTTC-3 Western Blotting Protein lysates were prepared in RIPA buffer (Pierce). Samples were run on 4% to 12% gradient Bis-Tris SDS-PAGE gels (Invitrogen), transferred onto PVDF membranes (Bio-Rad), and probed with antibodies against ERCC3 (ARP37963_P050; 1:2,500; Aviva Systems Biology), phospho-histone H2A.X (Ser139; 1:1,000; Cell Signaling Technology), phospho-chk1 (Ser345; 1:1,000; Cell Signaling Technology), and GAPDH (V-18; 1:400; Santa Cruz Biotechnology). HRPconjugated secondary antibodies were detected using ECL Prime Western Blotting Detection Reagent (GE Healthcare). Cell Viability Assays For viability assessment following treatment with IlludinS (Cayman Chemical), cells were seeded into 96-well plates 24 hours prior to treatment. For post-uv viability assays, cells were exposed to the appropriate doses of UV irradiation and subsequently seeded into 96-well plates. Cell viability was measured after 72 hours using the CellTiter-Glo Luminescent Cell Viability Assay (Promega). Host Cell Reactivation Assay The pis0 plasmid, derived from the pgl3-control vector containing the Firefly luciferase gene (Promega) and the pis2, a derivative from the prl-sv40 vector (Promega) containing the Renilla luciferase gene (as an internal transfection control), were used for transfection of cells. The pis0 vector was treated with increasing doses of UVC radiation. These vectors were cotransfected into the XPCS2BA and stable ERCC3 WT and R109X-overexpressing cell lines using the Fugene 6 Transfection Reagent (Promega). Forty-eight hours after DNA transfection, the luciferases activity was measured with the Dual-Glo Luciferase Assay System (Promega) and a GloMAX Luminometer (Promega). Flow Cytometry For flow cytometric analysis, 24 hours after seeding, cells were either left untreated or were treated with 8 ng/ml IlludinS for 1 hour. Treated cells were subsequently washed with PBS and supplemented with drug-free medium. The cells were harvested at different time points after treatment and fixed with 4% paraformaldehyde for 10 minutes at room temperature. Cells were quickly chilled on ice, and ice-cold MeOH was added to a final concentration of 90%. The cells were incubated on ice for 30 minutes and stored at 20 C. For immunostaining, the cells were washed twice with PBS/% BSA and incubated with anti phospho-histone H2A.X (Ser139) antibody (1:250; Cell Signaling Technology) for 1 hour at room temperature, washed again and incubated with Alexa-488 conjugated secondary antibody for 45 minutes at room temperature. For each condition, 20,000 cells were recorded, and data from two replicate experiments were analyzed using the FlowJo software (V10.1). Disclosure of Potential Conflicts of Interest M. Robson is a consultant/advisory board member for AstraZeneca. No potential conflicts of interest were disclosed by the other authors. Disclaimer The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH, the U.S. Army, or the Department of Defense. The funding institutions had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication. Authors Contributions Conception and design: J. Vijai, S. Topka, L. Norton, K. Offit Development of methodology: J. Vijai, S. Topka, S.N. Hart, F. Lejbkowicz Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): J. Vijai, S. Topka, D. Villano, V. Ravichandran, K.N. Maxwell, T. Thomas, P. Gaddam, B. Wenz, S.M. Lipkin, L. Zhang, Z.K. Stadler, J.N. Weitzel, S. Domchek, F.J. Couch, K.L. Nathanson, G. Rennert, K. Offit Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): J. Vijai, S. Topka, D. Villano, V. Ravichandran, T. Thomas, S. Carmi, S.N. Hart, S.M. Lipkin, NOVEMBER 2016 CANCER DISCOVERY 1273
8 Vijai et al. M.F. Walsh, F. Lejbkowicz, Z.K. Stadler, M.J. Daly, G. Rennert, K. Offit Writing, review, and/or revision of the manuscript: J. Vijai, S. Topka, D. Villano, K.N. Maxwell, T. Thomas, S. Kazzaz, B. Wenz, K.A. Schrader, S.M. Lipkin, S.L. Neuhausen, M.F. Walsh, F. Lejbkowicz, Z.K. Stadler, M. Robson, J.N. Weitzel, S. Domchek, F.J. Couch, K.L. Nathanson, L. Norton, G. Rennert, K. Offit Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): D. Villano, V. Ravichandran, K.N. Maxwell, A. Maria, P. Gaddam, A. Lincoln, S. Kazzaz, B. Wenz, H. Rennert, J.N. Weitzel, S. Domchek Study supervision: J. Vijai, S.M. Lipkin, J.N. Weitzel, K. Offit Other (team coordination): L. Norton Acknowledgments We thank the members of the Clalit National Israeli Cancer Control Center and Department of Community Medicine and Epidemiology, Carmel Medical Center and B Rappaport Faculty of Medicine, Haifa, Israel, and the New York Genome Center for the genotyping data. We also thank the members of the Ashkenazi Jewish Genome Project for the control genotypes, and Drs. Benezra and Mayr from MSKCC and Dr. Hoeijmakers from Erasmus University for cell lines and plasmids. Grant Support This work was supported by the Sharon Corzine Cancer Research Fund, The Robert and Kate Niehaus Center for Inherited Cancer Genomics, the Miele family initiative (K. Offit), and the Andrew Sabin Family Fund. This work was also supported by NIH R01CA and the Breast Cancer Research Foundation (F.J. Couch, K.L. Nathanson, M. Robson, and K. Offit), R21-CA (J. Vijai and Z.K. Stadler), and Department of Defense W81XWH (K.L. Nathanson). This work was also supported by funds from the City of Hope Clinical Cancer Genetics Community Research Network and the Hereditary Cancer Research Registry, supported in part by Award Number RC4CA (PI: J. Weitzel) from the National Cancer Institute and the Office of the Director, NIH. This work was also supported by the MSKCC Core grant NIH P30CA Received April 27, 2016 ; revised September 14, 2016 ; accepted September 16, 2016 ; published OnlineFirst September 21, REFERENCES 1. Couch FJ, Nathanson KL, Offit K. Two decades after BRCA: setting paradigms in personalized cancer care and prevention. Science 2014 ;343: Easton DF, Pharoah PD, Antoniou AC, Tischkowitz M, Tavtigian SV, Nathanson KL, et al. Gene-panel sequencing and the prediction of breast-cancer risk. N Engl J Med 2015 ;372: Michailidou K, Beesley J, Lindstrom S, Canisius S, Dennis J, Lush MJ, et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet 2015 ;47: Risinger MA, Groden J. Crosslinks and crosstalk: human cancer syndromes and DNA repair defects. Cancer Cell 2004 ;6: Brosh RM Jr. DNA helicases involved in DNA repair and their roles in cancer. Nat Rev Cancer 2013 ;13: Petrovski S, Wang Q, Heinzen EL, Allen AS, Goldstein DB. Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Genet 2013 ;9:e Vermeulen W, Scott RJ, Rodgers S, Muller HJ, Cole J, Arlett CF, et al. Clinical heterogeneity within xeroderma pigmentosum associated with mutations in the DNA repair and transcription gene ERCC3. Am J Hum Genet 1994 ;54: Mcmorris TC, Kelner MJ, Wang W, Estes LA, Montoya MA, Taetle R. Structure-Activity-Relationships of Illudins Analogs with Improved Therapeutic Index. J Org Chem 1992 ;57 : Marti TM, Hefner E, Feeney L, Natale V, Cleaver JE. H2AX phosphorylation within the G1 phase after UV irradiation depends on nucleotide excision repair and not DNA double-strand breaks. Proc Natl Acad Sci U S A 2006 ;103 : Carmi S, Hui KY, Kochav E, Liu X, Xue J, Grady F, et al. Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins. Nat Commun 2014 ;5 : Schaeffer L, Roy R, Humbert S, Moncollin V, Vermeulen W, Hoeijmakers JH, et al. DNA repair helicase: a component of BTF2 (TFIIH) basic transcription factor. Science 1993 ; 260 : Oh K-S, Khan SG, Jaspers NGJ, Raams A, Ueda T, Lehmann A, et al. Phenotypic heterogeneity in the XPB DNA helicase gene (ERCC3): xeroderma pigmentosum without and with Cockayne syndrome. Hum Mut 2006 ;27 : Singh A, Compe E, Le May N, Egly JM. TFIIH subunit alterations causing xeroderma pigmentosum and trichothiodystrophy specifically disturb several steps during transcription. Am J Hum Genet 2015 ;96 : Seemanova E. An increased risk for malignant neoplasms in heterozygotes for a syndrome of microcephaly, normal intelligence, growth retardation, remarkable facies, immunodeficiency and chromosomal instability. Mut Res 1990 ;238 : Digweed M, Reis A, Sperling K. Nijmegen breakage syndrome: consequences of defective DNA double strand break repair. BioEssays 1999 ;21 : Bassing CH, Suh H, Ferguson DO, Chua KF, Manis J, Eckersdorff M, et al. Histone H2AX: a dosage-dependent suppressor of oncogenic translocations and tumors. Cell 2003 ;114 : Celeste A, Difilippantonio S, Difilippantonio MJ, Fernandez-Capetillo O, Pilch DR, Sedelnikova OA, et al. H2AX haploinsufficiency modifies genomic stability and tumor susceptibility. Cell 2003 ;114 : Goss KH, Risinger MA, Kordich JJ, Sanz MM, Straughen JE, Slovek LE, et al. Enhanced tumor formation in mice heterozygous for Blm mutation. Science 2002 ;297 : Lam MH, Liu Q, Elledge SJ, Rosen JM. Chk1 is haploinsufficient for multiple functions critical to tumor suppression. Cancer Cell 2004 ;6 : Nikkila J, Parplys AC, Pylkas K, Bose M, Huo Y, Borgmann K, et al. Heterozygous mutations in PALB2 cause DNA replication and damage response defects. Nat Commun 2013 ; 4 : Andressoo JO, Weeda G, de Wit J, Mitchell JR, Beems RB, van Steeg H, et al. An Xpb mouse model for combined xeroderma pigmentosum and cockayne syndrome reveals progeroid features upon further attenuation of DNA repair. Mol Cell Biol 2009 ;29 : Diderich K, Alanazi M, Hoeijmakers JH. Premature aging and cancer in nucleotide excision repair-disorders. DNA Repair (Amst) 2011 ; 10 : Itoh T, Cado D, Kamide R, Linn S. DDB2 gene disruption leads to skin tumors and resistance to apoptosis after exposure to ultraviolet light but not a chemical carcinogen. Proc Natl Acad Sci U S A 2004 ;101 : Cheo DL, Meira LB, Burns DK, Reis AM, Issac T, Friedberg EC. Ultraviolet B radiation-induced skin cancer in mice defective in the Xpc, Trp53, and Apex (HAP1) genes: genotype-specific effects on cancer predisposition and pathology of tumors. Cancer Res 2000 ;60 : Wijnhoven SW, Kool HJ, Mullenders LH, van Zeeland AA, Friedberg EC, van der Horst GT, et al. Age-dependent spontaneous mutagenesis in Xpc mice defective in nucleotide excision repair. Oncogene 2000 ;19 : Andressoo JO, Mitchell JR, de Wit J, Hoogstraten D, Volker M, Toussaint W, et al. An Xpd mouse model for the combined xeroderma pigmentosum/cockayne syndrome exhibiting both cancer predisposition and segmental progeria. Cancer Cell 2006 ;10 : Shaag A, Walsh T, Renbaum P, Kirchhoff T, Nafa K, Shiovitz S, et al. Functional and genomic approaches reveal an ancient CHEK2 allele 1274 CANCER DISCOVERY NOVEMBER
9 ERCC3 R109X Mutation and Breast Cancer associated with breast cancer in the Ashkenazi Jewish population. Hum Mol Genet 2005 ;14 : Redston M, Nathanson KL, Yuan ZQ, Neuhausen SL, Satagopan J, Wong N, et al. The APCI1307K allele and breast cancer risk. Nat Genet 1998 ;20 : Weischer M, Bojesen SE, Ellervik C, Tybjaerg-Hansen A, Nordestgaard BG. CHEK2*1100delC genotyping for clinical assessment of breast cancer risk: meta-analyses of 26,000 patient cases and 27,000 controls. J Clin Oncol 2008 ;26 : Alanee S, Couch F, Offit K. Association of a HOXB13 variant with breast cancer. N Engl J Med 2012 ;367 : Erkko H, Xia B, Nikkila J, Schleutker J, Syrjakoski K, Mannermaa A, et al. A recurrent mutation in PALB2 in Finnish cancer families. Nature 2007 ;446 : Schnurbein G, Hauke J, Wappenschmidt B, Weber-Lassalle N, Engert S, Hellebrand H, et al. RAD51C deletion screening identifies a recurrent gross deletion in breast cancer and ovarian cancer families. Breast Cancer Res 2013 ;15 :R Goyal G, Fan T, Silberstein PT. Hereditary cancer syndromes: utilizing DNA repair deficiency as therapeutic target. Fam Cancer 2016 ;15 : Sonnenblick A, de Azambuja E, Azim HA Jr., Piccart M. An update on PARP inhibitors moving to the adjuvant setting. Nat Rev Clin Oncol 2015 ;12 : Chen MC, Zhou B, Zhang K, Yuan YC, Un F, Hu S, et al. The novel ribonucleotide reductase inhibitor COH29 inhibits DNA repair in vitro. Mol Pharmacol 2015 ;87 : Lok BH, Powell SN. Molecular pathways: understanding the role of Rad52 in homologous recombination for therapeutic advancement. Clin Cancer Res 2012 ;18 : Zhang F. Genome Engineering Resources Zhang MIT. N.p., Web. 01 Apr < /> NOVEMBER 2016 CANCER DISCOVERY 1275
10 A Recurrent ERCC3 Truncating Mutation Confers Moderate Risk for Breast Cancer Joseph Vijai, Sabine Topka, Danylo Villano, et al. Cancer Discov 2016;6: Published OnlineFirst September 21, Updated version Supplementary Material Access the most recent version of this article at: doi: / cd Access the most recent supplemental material at: Cited articles Citing articles This article cites 36 articles, 10 of which you can access for free at: This article has been cited by 1 HighWire-hosted articles. Access the articles at: alerts Sign up to receive free -alerts related to this article or journal. Reprints and Subscriptions Permissions To order reprints of this article or to subscribe to the journal, contact the AACR Publications Department at pubs@aacr.org. To request permission to re-use all or part of this article, use this link Click on "Request Permissions" which will take you to the Copyright Clearance Center's (CCC) Rightslink site.
Functional validation of cancer susceptibility genes using gene editing
 Functional validation of cancer susceptibility genes using gene editing 2-22-2017 Sabine Topka Research Fellow Niehaus Center for Inherited Cancer Genomics www.mskcc.org Inherited Predisposition to Cancer
Functional validation of cancer susceptibility genes using gene editing 2-22-2017 Sabine Topka Research Fellow Niehaus Center for Inherited Cancer Genomics www.mskcc.org Inherited Predisposition to Cancer
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Inherited Cancer Genomics and Prevention:
 Inherited Cancer Genomics and Prevention: How much cancer risk is inherited? Clinical utility of germline genetic testing in precision prevention and targeted therapy Kenneth Offit MD MPH Chief, Clinical
Inherited Cancer Genomics and Prevention: How much cancer risk is inherited? Clinical utility of germline genetic testing in precision prevention and targeted therapy Kenneth Offit MD MPH Chief, Clinical
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Medical Policy Manual. Topic: Genetic Testing for Hereditary Breast and/or Ovarian Cancer. Date of Origin: January 27, 2011
 Medical Policy Manual Topic: Genetic Testing for Hereditary Breast and/or Ovarian Cancer Date of Origin: January 27, 2011 Section: Genetic Testing Last Reviewed Date: July 2014 Policy No: 02 Effective
Medical Policy Manual Topic: Genetic Testing for Hereditary Breast and/or Ovarian Cancer Date of Origin: January 27, 2011 Section: Genetic Testing Last Reviewed Date: July 2014 Policy No: 02 Effective
Biochemistry 201: DNA repair January 24, 26, 2000 Gilbert Chu
 1) Why study DNA repair? Biochemistry 201: DNA repair January 24, 26, 2000 Gilbert Chu The genome is assaulted by a myriad of different agents that cause DNA damage. Sources of damage can be both exogenous
1) Why study DNA repair? Biochemistry 201: DNA repair January 24, 26, 2000 Gilbert Chu The genome is assaulted by a myriad of different agents that cause DNA damage. Sources of damage can be both exogenous
The Genetics of Breast and Ovarian Cancer Prof. Piri L. Welcsh
 The Genetics of Breast Piri L. Welcsh, PhD Research Assistant Professor University of Washington School of Medicine Division of Medical Genetics 1 Genetics of cancer All cancers arise from genetic and
The Genetics of Breast Piri L. Welcsh, PhD Research Assistant Professor University of Washington School of Medicine Division of Medical Genetics 1 Genetics of cancer All cancers arise from genetic and
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
CRISPR/Cas9 cleavage of viral DNA efficiently suppresses hepatitis B virus
 0 0 CRISPR/Cas cleavage of viral DNA efficiently suppresses hepatitis B virus Authors: Vyas Ramanan,, Amir Shlomai,, David B.T. Cox,,,, Robert E. Schwartz,,, Eleftherios Michailidis, Ankit Bhatta, David
0 0 CRISPR/Cas cleavage of viral DNA efficiently suppresses hepatitis B virus Authors: Vyas Ramanan,, Amir Shlomai,, David B.T. Cox,,,, Robert E. Schwartz,,, Eleftherios Michailidis, Ankit Bhatta, David
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: DAMD17-03-1-0392 TITLE: The Role of Notch Signaling Pathway in Breast Cancer Pathogenesis PRINCIPAL INVESTIGATOR: Annapoorni Rangarajan, Ph.D. CONTRACTING ORGANIZATION: Indian Institute
AD Award Number: DAMD17-03-1-0392 TITLE: The Role of Notch Signaling Pathway in Breast Cancer Pathogenesis PRINCIPAL INVESTIGATOR: Annapoorni Rangarajan, Ph.D. CONTRACTING ORGANIZATION: Indian Institute
Germline Testing for Hereditary Cancer with Multigene Panel
 Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy
 Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
The lymphoma-associated NPM-ALK oncogene elicits a p16ink4a/prb-dependent tumor-suppressive pathway. Blood Jun 16;117(24):
 DNA Sequencing Publications Standard Sequencing 1 Carro MS et al. DEK Expression is controlled by E2F and deregulated in diverse tumor types. Cell Cycle. 2006 Jun;5(11) 2 Lassandro L et al. The DNA sequence
DNA Sequencing Publications Standard Sequencing 1 Carro MS et al. DEK Expression is controlled by E2F and deregulated in diverse tumor types. Cell Cycle. 2006 Jun;5(11) 2 Lassandro L et al. The DNA sequence
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology
 Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology Epithelial Ovarian Cancer - Standard Current Treatment: Surgery with De-bulking + Platinum-Taxane based Chemotherapy - No
Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology Epithelial Ovarian Cancer - Standard Current Treatment: Surgery with De-bulking + Platinum-Taxane based Chemotherapy - No
CANCER. Inherited Cancer Syndromes. Affects 25% of US population. Kills 19% of US population (2nd largest killer after heart disease)
 CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
Cancer Treatment and Research
 Cancer Treatment and Research Volume 155 Series Editor Steven T. Rosen For further volumes: http://www.springer.com/series/5808 Boris Pasche Editor Cancer Genetics 123 Editor Boris Pasche, MD, PhD, FACP
Cancer Treatment and Research Volume 155 Series Editor Steven T. Rosen For further volumes: http://www.springer.com/series/5808 Boris Pasche Editor Cancer Genetics 123 Editor Boris Pasche, MD, PhD, FACP
Chapter 9. Cells Grow and Reproduce
 Chapter 9 Cells Grow and Reproduce DNA Replication DNA polymerase Addition of a nucleotide to the 3 end of a growing strand Use dntps as substrate Release of pyrophosphate Initiation of Replication Replication
Chapter 9 Cells Grow and Reproduce DNA Replication DNA polymerase Addition of a nucleotide to the 3 end of a growing strand Use dntps as substrate Release of pyrophosphate Initiation of Replication Replication
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Asingle inherited mutant gene may be enough to
 396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
Supplementary Figure S1A
 Supplementary Figure S1A-G. LocusZoom regional association plots for the seven new cross-cancer loci that were > 1 Mb from known index SNPs. Genes up to 500 kb on either side of each new index SNP are
Supplementary Figure S1A-G. LocusZoom regional association plots for the seven new cross-cancer loci that were > 1 Mb from known index SNPs. Genes up to 500 kb on either side of each new index SNP are
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Problem Set 8 Key 1 of 8
 7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
Example: Distance in M.U. % Crossing Over Why? Double crossovers
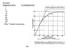 Example: Distance in M.U. % Crossing Over 1 5 10 15 50 80 100 107 Why? Double crossovers 232 .. A B. a b. 1. A fully heterozygous gray-bodied (b+), normal winged (vg+) female F 1 fruit fly crossed with
Example: Distance in M.U. % Crossing Over 1 5 10 15 50 80 100 107 Why? Double crossovers 232 .. A B. a b. 1. A fully heterozygous gray-bodied (b+), normal winged (vg+) female F 1 fruit fly crossed with
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
Leveraging Interaction between Genetic Variants and Mammographic Findings for Personalized Breast Cancer Diagnosis
 Leveraging Interaction between Genetic Variants and Mammographic Findings for Personalized Breast Cancer Diagnosis Jie Liu, PhD 1, Yirong Wu, PhD 1, Irene Ong, PhD 1, David Page, PhD 1, Peggy Peissig,
Leveraging Interaction between Genetic Variants and Mammographic Findings for Personalized Breast Cancer Diagnosis Jie Liu, PhD 1, Yirong Wu, PhD 1, Irene Ong, PhD 1, David Page, PhD 1, Peggy Peissig,
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Role of genetic testing in familial breast cancer outside of BRCA1 and BRCA2
 Role of genetic testing in familial breast cancer outside of BRCA1 and BRCA2 Introduction Most commonly diagnosed cancer in South African women and the second most commonly diagnosed cancer in Black women
Role of genetic testing in familial breast cancer outside of BRCA1 and BRCA2 Introduction Most commonly diagnosed cancer in South African women and the second most commonly diagnosed cancer in Black women
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-06-1-0524 TITLE: Elucidating and Modeling Irradiation Effects on Centrosomal and Chromosomal Stability within Breast Cancer PRINCIPAL INVESTIGATOR: Christopher A. Maxwell, Ph.D.
AD Award Number: W81XWH-06-1-0524 TITLE: Elucidating and Modeling Irradiation Effects on Centrosomal and Chromosomal Stability within Breast Cancer PRINCIPAL INVESTIGATOR: Christopher A. Maxwell, Ph.D.
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
RNA-Seq guided gene therapy for vision loss. Michael H. Farkas
 RNA-Seq guided gene therapy for vision loss Michael H. Farkas The retina is a complex tissue Many cell types Neural retina vs. RPE Each highly dependent on the other Graw, Nature Reviews Genetics, 2003
RNA-Seq guided gene therapy for vision loss Michael H. Farkas The retina is a complex tissue Many cell types Neural retina vs. RPE Each highly dependent on the other Graw, Nature Reviews Genetics, 2003
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
609G: Concepts of Cancer Genetics and Treatments (3 credits)
 Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands)
 Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
Cancer. Questions about cancer. What is cancer? What causes unregulated cell growth? What regulates cell growth? What causes DNA damage?
 Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
MRC-Holland MLPA. Description version 18; 09 September 2015
 SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Managing Moderate Penetrance
 Managing Moderate Penetrance Thomas Slavin, MD, FACMG Assistant Clinical Professor, Department of Medical Oncology, Division of Clinical Cancer Genetics Program Member, Cancer Control and Population Sciences
Managing Moderate Penetrance Thomas Slavin, MD, FACMG Assistant Clinical Professor, Department of Medical Oncology, Division of Clinical Cancer Genetics Program Member, Cancer Control and Population Sciences
Hereditary Prostate Cancer: From Gene Discovery to Clinical Implementation
 Hereditary Prostate Cancer: From Gene Discovery to Clinical Implementation Kathleen A. Cooney, MD MACP Duke University School of Medicine Duke Cancer Institute (No disclosures to report) Overview Prostate
Hereditary Prostate Cancer: From Gene Discovery to Clinical Implementation Kathleen A. Cooney, MD MACP Duke University School of Medicine Duke Cancer Institute (No disclosures to report) Overview Prostate
Early Embryonic Development
 Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Speakers. Text to be added
 Speakers SESSION 1 Clinical status and perspectives for hereditary breast and ovarian cancer (HBOC) risk prediction. Fergus Couch Mayo Clinic Cancer Center and the Center for Individualized Medicine, Diana
Speakers SESSION 1 Clinical status and perspectives for hereditary breast and ovarian cancer (HBOC) risk prediction. Fergus Couch Mayo Clinic Cancer Center and the Center for Individualized Medicine, Diana
Corporate Medical Policy Genetic Testing for Breast and Ovarian Cancer
 Corporate Medical Policy Genetic Testing for Breast and Ovarian Cancer File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_breast_and_ovarian_cancer 8/1997 8/2017
Corporate Medical Policy Genetic Testing for Breast and Ovarian Cancer File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_breast_and_ovarian_cancer 8/1997 8/2017
w ª wy xvwz A ª vw xvw P ª w} xvw w Æ w Æ V w,x Æ w Æ w Æ y,z Æ { Æ y,z, w w w~ w wy}æ zy Æ wyw{ xæ wz w xywæ xx Æ wv Æ } w x w x w Æ w Æ wy} zy Æ wz
 w ª wy xvwz A ª vw xvw P ª w} xvw w Æ w Æ V w,x Æ w Æ w Æ y,z Æ { Æ y,z, w w w~ w wy}æ zy Æ wyw{ xæ wz w xywæ xx Æ wv Æ } w x w x w Æ w Æ wy} zy Æ wz {w Æ Æ wyw{ x w Germ-line mutations in BRCA1 are associated
w ª wy xvwz A ª vw xvw P ª w} xvw w Æ w Æ V w,x Æ w Æ w Æ y,z Æ { Æ y,z, w w w~ w wy}æ zy Æ wyw{ xæ wz w xywæ xx Æ wv Æ } w x w x w Æ w Æ wy} zy Æ wz {w Æ Æ wyw{ x w Germ-line mutations in BRCA1 are associated
Section D: The Molecular Biology of Cancer
 CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Supplementary Text. Novel variants in NUDT15 and thiopurine intolerance in children with acute lymphoblastic leukemia from diverse ancestry
 Supplementary Text Novel variants in NUDT15 and thiopurine intolerance in children with acute lymphoblastic leukemia from diverse ancestry Takaya Moriyama a, Yung-Li Yang b, Rina Nishii a, c, Hany Ariffin
Supplementary Text Novel variants in NUDT15 and thiopurine intolerance in children with acute lymphoblastic leukemia from diverse ancestry Takaya Moriyama a, Yung-Li Yang b, Rina Nishii a, c, Hany Ariffin
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER"
 IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER" Edmond Marzbani, MD December 2, 2008 Early diagnosis of cancer Many solid tumors are potentially curable if diagnosed at an early stage
IDENTIFICATION OF BIOMARKERS FOR EARLY DIAGNOSIS OF BREAST CANCER" Edmond Marzbani, MD December 2, 2008 Early diagnosis of cancer Many solid tumors are potentially curable if diagnosed at an early stage
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
RESISTANCE RELATIONSHIPS BETWEEN PLATINUM AND PARP-INHIBITORS IN OVARIAN CANCER.
 RESISTANCE RELATIONSHIPS BETWEEN PLATINUM AND PARP-INHIBITORS IN OVARIAN CANCER. Britta Stordal 1, Yidong Chen 2, Angela Farrelly 3, Danielle Gallagher 3, Steven Busschots 1, Mattia Cremona 3, Mark Carey
RESISTANCE RELATIONSHIPS BETWEEN PLATINUM AND PARP-INHIBITORS IN OVARIAN CANCER. Britta Stordal 1, Yidong Chen 2, Angela Farrelly 3, Danielle Gallagher 3, Steven Busschots 1, Mattia Cremona 3, Mark Carey
Impact of hyper-o-glcnacylation on apoptosis and NF-κB activity SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
TOUR REPORT. Report on participation of the ICMR International Fellow (ICMR IF) in Training / Research abroad
 TOUR REPORT Report on participation of the ICMR International Fellow (ICMR IF) in Training / Research abroad 1. Name and Designation of ICMR IF : Dr. A. Venkateshwari, Assistant Professor 2. Address: Dr.
TOUR REPORT Report on participation of the ICMR International Fellow (ICMR IF) in Training / Research abroad 1. Name and Designation of ICMR IF : Dr. A. Venkateshwari, Assistant Professor 2. Address: Dr.
Summary... 2 TRANSLATIONAL RESEARCH Tumour gene expression used to direct clinical decision-making for patients with advanced cancers...
 ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
Pre-made Reporter Lentivirus for NF-κB Signal Pathway
 Pre-made Reporter for NF-κB Signal Pathway Cat# Product Name Amounts LVP965-P or: LVP965-P-PBS NFKB-GFP (Puro) LVP966-P or: LVP966-P-PBS NFKB-RFP (Puro) LVP967-P or: LVP967-P-PBS NFKB-Luc (Puro) LVP968-P
Pre-made Reporter for NF-κB Signal Pathway Cat# Product Name Amounts LVP965-P or: LVP965-P-PBS NFKB-GFP (Puro) LVP966-P or: LVP966-P-PBS NFKB-RFP (Puro) LVP967-P or: LVP967-P-PBS NFKB-Luc (Puro) LVP968-P
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Human Genetics 542 Winter 2018 Syllabus
 Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
5/2/18. After this class students should be able to: Stephanie Moon, Ph.D. - GWAS. How do we distinguish Mendelian from non-mendelian traits?
 corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
Disclosure. Summary. Circulating DNA and NGS technology 3/27/2017. Disclosure of Relevant Financial Relationships. JS Reis-Filho, MD, PhD, FRCPath
 Circulating DNA and NGS technology JS Reis-Filho, MD, PhD, FRCPath Director of Experimental Pathology, Department of Pathology Affiliate Member, Human Oncology and Pathogenesis Program Disclosure of Relevant
Circulating DNA and NGS technology JS Reis-Filho, MD, PhD, FRCPath Director of Experimental Pathology, Department of Pathology Affiliate Member, Human Oncology and Pathogenesis Program Disclosure of Relevant
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
BRCA Gene Family at Age 20. Considering the Impact
 BRCA Gene Family at Age 20 Considering the Impact What Have We Learned About Breast Cancer from BRCA Genes? Introduction What did we learn over the past 20 years? Learn about BRCA genes? Learn about breast
BRCA Gene Family at Age 20 Considering the Impact What Have We Learned About Breast Cancer from BRCA Genes? Introduction What did we learn over the past 20 years? Learn about BRCA genes? Learn about breast
Pre-made Reporter Lentivirus for MAPK/ERK Signal Pathway
 Pre-made Reporter for MAPK/ERK Signal Pathway Cat# Product Name Amounts LVP957-P or: LVP957-P-PBS SRE-GFP (Puro) LVP958-P or: LVP958-P-PBS SRE-RFP (Puro) LVP959-P or: LVP959-P-PBS SRE-Luc (Puro) LVP960-P
Pre-made Reporter for MAPK/ERK Signal Pathway Cat# Product Name Amounts LVP957-P or: LVP957-P-PBS SRE-GFP (Puro) LVP958-P or: LVP958-P-PBS SRE-RFP (Puro) LVP959-P or: LVP959-P-PBS SRE-Luc (Puro) LVP960-P
Cancer genetics
 Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer
 AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
Spring 2015 Module 2 System Engineering and Protein Founda<ons
 20.109 Spring 2015 Module 2 System Engineering and Protein Founda
20.109 Spring 2015 Module 2 System Engineering and Protein Founda
Supplemental Information. Inhibition of the Proteasome b2 Site Sensitizes. Triple-Negative Breast Cancer Cells
 Cell Chemical Biology, Volume 24 Supplemental Information Inhibition of the Proteasome b2 Site Sensitizes Triple-Negative Breast Cancer Cells to b5 Inhibitors and Suppresses Nrf1 Activation Emily S. Weyburne,
Cell Chemical Biology, Volume 24 Supplemental Information Inhibition of the Proteasome b2 Site Sensitizes Triple-Negative Breast Cancer Cells to b5 Inhibitors and Suppresses Nrf1 Activation Emily S. Weyburne,
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
