Introduction to epigenetics: chromatin modifications, DNA methylation and the CpG Island landscape. Héctor Corrada Bravo CMSC702 Spring 2014
|
|
|
- Brook Lester
- 5 years ago
- Views:
Transcription
1 Introduction to epigenetics: chromatin modifications, DN methylation and the p Island landscape Héctor orrada Bravo MS702 Spring 2014
2 enetics: the alphabet of life Letters of DN sequence carry information How is this information read and parsed? We need grammar! 2
3 Differentiation Different genes are expressed during different stages and in different tissues 3
4 (3.4x10-10 meters/bp) x (6x10 9 bp/genome) = ~2 meters/genome Klug and ummings, 1997 Radius of the nucleus is ~ 10 µm!!!
5 [(6 x 10 9 bp/genome) / (195 bp/nucleosome)] = ~ 30.8 x 10 6 nucleosomes/genome ~ 5 % of nuclear volume
6 onformation is dynamic! (we ll discuss methods to assay this conformation later on )
7 We ll study methods to assay a number of mechanisms of epigenetic regulation DN methylation Nucleosome positioning and histone modifications In eukaryotes, DN methylation usually occur at p dinucleotides
8 DN Methylation is sometimes repressive Unmethylated p dinucleotides Methylated p dinucleotides ranscription repressors bound to methyl- group Robertson and Wolffe, Nat Rev enet, 2000
9 DN methylation in cancer
10 DN methylation, histone modifications and nucleosome positioning are coordinated!! New technologies are allowing us to now assay this coordination! [Brinkman, et al., enome Research 2012]
11 Epigenetics: the grammar of life Epigenetics literally means above the genome
12 One more thing.! How does a cell retain epigenetic state?
13 Methylation H 3 H 3!!
14 H 3 H 3!! What happens during cell replication?
15 !!!! H 3 H 3 What happens during cell replication?
16 !!!! H 3 H 3 H 3 H 3 What happens during cell replication? DN methylation is replicated!
17 Liver Brain!!
18 !! H 3 H 3 H 3 H 3 Liver Brain
19 !! H 3 H 3 H 3 H 3 Liver Brain!! H 3 H 3 H 3 H 3
20 How do we measure DN methylation?
21 !! H 3 H 3!! H 3 H 3 Liver Brain
22 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 85% MethylaWon chr3:44,031,616-44,031,626
23 Bisulfite reatment
24 Bisulfite reatment!
25 BS-Seq overage: 12 MethylaWon Evidence: 8 MethylaWon Percentage: 67% NN NN -
26 BS- seq lignment is much trickier: Naïve strategy: do nothing, hope not many p in a single read Smarter strategy: bisulfite convert reference: turn all s to s Smartest strategy: be unbiased and try all combinations of methylated/un- methylated ps in each read
27 BS- seq here are similarities to SNP calling EXEP: we want to measure percentages Use a binomial model to estimate p, percentage of methylation llow for sequencing errors, coverage differences, etc.
28 Smoothing
29 Proportion of neighboring p also methylated/ not methylated
30 rue signal (simulated)
31 Observed data
32 Observed data and true signal
33 What is methylated (above 50%)?
34 Naïve approach
35 Many false positives (FP)
36 Smooth
37 No FP, but one false negative
38 Smooth less? No FN, lots of FP
39 We prefer this!
40 Measuring DN Methylation -./ / / !"#$%&'()'*+,#"%!"#$%&'/)'*+,#"%!"#$%&'2)'*+,#"% EsWmaWng percentages Use local- likelihood smoothing method Based on loess High- frequency smoothing detects local methylawon levels Low- frequency smoothing detects long- range methylawon levels
41 Large regions of methylation loss in cancer (a) Methylation PMD LOKS LD Islands enes 100kb normals cancers! Found large blocks of hypo-methylation (sometimes Mbps long) in colon cancer hese regions coincide with hyper-variable regions across cancer types ene expression analysis [Hansen, et al., Nature enetics] Hector orrada Bravo 41
42 Hector orrada Bravo enes with hyper-variable gene expression in colon cancer are enriched in these blocks expression SLO1B3 XL11 LDN18 ME6 INHB MMP10 IL24 S10012 IL6 PH ME12 R3 ME6 XL5 NFIP6 Normal ancer ) False posi rue positive rate yo Sab Sabates, et al. norm yorffy, et al. normal range upper bound B) ) ene expression hyper-variability [Hansen, et al., Nature enetics]
43 [Feinberg & Irizarry, 2009] 1. Variability, perhaps epigenetically mediated, increases disease susceptibility
44 [Feinberg & Irizarry, 2009] 1. Variability, perhaps epigenetically mediated, increases disease susceptibility! 2.Increased gene expression variability of specific genes/regions is a defining characteristic of cancer
45 gene expression anti-profiles expression SLO1B3 XL11 LDN18 ME6 INHB MMP10 IL24 S10012 IL6 PH ME12 R3 ME6 XL5 NFIP6 Normal ancer RESERH RILE Open ccess ene expression anti-profiles as a basis for accurate universal cancer signatures Héctor orrada Bravo 1*, Vasyl Pihur 2, Matthew Mcall 3, Rafael Irizarry 2 and Jeffrey Leek 2 bstract orrada Bravo et al. BM Bioinformatics 2012, 13:272
46 Universal anti-profile signatures rue positive rate adrenal_cortex: 0.99 U colon: 0.98 U endometrium: 0.99 U kidney: 0.87 U skin: 0.98 U stomach: 0.88 U vulva: 1.00 U False positive rate orrada Bravo et al. BM Bioinformatics 2012, 13:272 RESERH RILE ene expression anti-profiles as a basis for accurate universal cancer signatures Héctor orrada Bravo 1*, Vasyl Pihur 2, Matthew Mcall 3, Rafael Irizarry 2 and Jeffrey Leek 2 Open ccess 46
47 Universal anti-profile signatures Increased variability increases through cancer progression () (B) density all probesets anti profile probesets density all probesets anti profile probesets tumor variance/normal variance cancer variance/adenoma variance [Dinalankara & HB, under submission] 47
48 Universal anti-profile signatures Relapse in lung cancer patients patient relapse Stratification by anti-profile score Score based on departure from lung normal expression on universal anti-profile genes log-rank statistic 15.6 probability of survival [Dinalankara & HB, under submission] Lower 50% op 50% time(years) 48
49 nomaly lassification (anti-profilesvm) Extend the anti-profile idea to nonlinear classification methods ore idea: measure similarity between anomalies indirectly through similarity to normal class arcinoma Normal denoma [Dinalankara, et al., under submission]
50 Finding hypo-methylation blocks w/ beadarrays chr1: Methylation kb Normal ancer Methylation difference Minfi/ 176.0Mb 176.5Mb 177.0Mb 177.5Mb 178.0Mb Hansen et al. hypo hyper Smoothing method for large region detection for Illumina 450k array Implemented in minfi Bioconductor package [ryee, et al., Bioinformatics] 50
51 Hypo-methylation blocks in multiple solid tumors verage Methylation ancer-normal Methylation Standard Deviation Methylation ) B) ) colon lung pan thyroid D) E) F) 450k profiling across five tumor types colon, lung, breast, pancreas and thyroid Hypo-methylation blocks occur in all tumor types some universal, many cancer-specific [imp et al., in preparation] 51
52 Hypo-methylation blocks in multiple solid tumors in block odds ratio in block odds ratio colon hyper variability log ratio pancreas hyper variability log ratio in block odds ratio in block odds ratio breast hyper variability log ratio thyroid hyper variability log ratio in block odds ratio lung hyper variability log ratio ene expression hyper-variability enriched in hypo-methylation blocks [imp et al., in preparation] 52
53 Hypo-methylation blocks in multiple solid tumors olon Progression hyroid Progression verage in blocks verage in Island ) D) E) F) verage in blocks verage in Island Normal denoma ancer Metastasis Normal Benign denoma MinInv ancer apsinv ancer VascInv ancer Metastasis Degree of hypo-methylation in blocks increases through cancer progression 53
54 Summary Large regions of hypo-methylation seems to be consistent in cancer occur in pre-cancerous lesions hypo-methylation increases with cancer progression ene expression hyper-variability enriched within these regions tissue-specific genes also enriched within these regions can use degree of deviation from normality as stable diagnosis and prognosis mark in 54
55 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 H 3 85% MethylaWon chr3:44,031,616-44,031,626
56 )Methylation his is a heterogenous cell population PMD LOKS LD Islands enes 100kb normals cancers Normal umor 56
57 )Methylation his is a heterogenous cell population PMD LOKS LD Islands enes 100kb normals cancers Normal umor 57
58 Epipolymorphism mos anay s Weizmann n n n n Figu met but 58
59 )Methylation his is a heterogenous cell population PMD LOKS LD Islands enes 100kb normals cancers How are cell populations changing in normal and tumor? Same set of methylation patterns, with different abundances? re patterns lost or gained? 59
60 Heterogeneous elltype omposition elltype-specific Methylation Patterns Methylated p Unmethylated p
61 ligned Reads their overlap allows reconstruction of long methylation patterns
62 he pattern reconstruction problem iven a set of mapped reads Determine composition of cell-specific methylation patterns omposition: Pattern identification: which cell-specific methylation patterns are present Pattern abundances: abundance of each cell-specific pattern present Derive statistics: community richness: how many unique patterns community diversity: distribution of pattern abundance fragment uniformity: long-scale spatial methylation correlation ompare statistics across groups: re richness and/or diversity increasing or decreasing in cancer? Is spatial correlation lost in cancer? 62
63 elltype-specific Methylation Patterns Heterogeneous elltype omposition Methylated p Unmethylated p Region raph ligned Reads no. reads assigned
64 Methylation pattern reconstruction by network flows Region raph Penalized method of moments estimation procedure: # parameters: # paths through region graph region coverage penalty induces small number of patterns with non-zero abundance min p X X y v v u:(v,u)2e `vu X p:(v,u)2p p + X p p 64
65 Bisulfite capture data Frequency P 4 Frequency P_N_5 Frequency P assembled fragment size P_N_ assembled fragment size P assembled fragment size P_N_6 Frequency Frequency Frequency assembled fragment size assembled fragment size assembled fragment size 65
66 Bisulfite capture data 66
67 Bisulfite apture Data normal num. patterns tumor num. patterns normal entropy tumor entropy normal entropy (normalized) tumor entropy (normalized Similar entropy distribution in normal and tumor, but many regions of differential entropy
68 Bisulfite apture Data 68 Not associated with methylation difference (with some caveats) Subject 1 methylation difference entropy difference Subject 2 methylation difference entropy difference Subject 3 methylation difference entropy difference
69 elltype-specific methylation pattern reconstruction Unprecedented view into molecular profiling omplex relationship between cell-to-cell differences and average methylation differences Efficient formulation as network flow problem Exploring relationship to stochastic optimization In general: properties of population inferences from inferred features Here: patterns are inferred from overlaps metagenomics: OUs inferred from sequence similarity clustering RNseq: transcripts inferred from junction spanning reads (or junction spanning k-mers) 69
Analysis of shotgun bisulfite sequencing of cancer samples
 Analysis of shotgun bisulfite sequencing of cancer samples Kasper Daniel Hansen Postdoc with Rafael Irizarry Johns Hopkins Bloomberg School of Public Health Brixen, July 1st, 2011 The
Analysis of shotgun bisulfite sequencing of cancer samples Kasper Daniel Hansen Postdoc with Rafael Irizarry Johns Hopkins Bloomberg School of Public Health Brixen, July 1st, 2011 The
ABSTRACT. follows from prognosis and diagnosis applications. Anomaly classification is also a
 ABSTRACT Title of dissertation: ANTI-PROFILES FOR ANOMALY CLASSIFICATION AND REGRESSION Wikum Dinalankara, Doctor of Philosophy, 2015 Dissertation directed by: Professor Hèctor Corrada-Bravo Department
ABSTRACT Title of dissertation: ANTI-PROFILES FOR ANOMALY CLASSIFICATION AND REGRESSION Wikum Dinalankara, Doctor of Philosophy, 2015 Dissertation directed by: Professor Hèctor Corrada-Bravo Department
Introduction to antiprofiles
 Introduction to antiprofiles Héctor Corrada Bravo hcorrada@gmail.com Modified: March 13, 2013. Compiled: April 30, 2018 Introduction This package implements the gene expression anti-profiles method in
Introduction to antiprofiles Héctor Corrada Bravo hcorrada@gmail.com Modified: March 13, 2013. Compiled: April 30, 2018 Introduction This package implements the gene expression anti-profiles method in
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
ChIP-seq data analysis
 ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response
 Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION. Jude Fitzgibbon
 FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
NIH Public Access Author Manuscript Nat Genet. Author manuscript; available in PMC 2012 February 1.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(8): 768 775. doi:10.1038/ng.865. Increased methylation variation in epigenetic domains across cancer types Kasper
NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(8): 768 775. doi:10.1038/ng.865. Increased methylation variation in epigenetic domains across cancer types Kasper
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Measuring DNA Methylation with the MinION. Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Measuring DNA Methylation with the MinION
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
Epigenetics. Jenny van Dongen Vrije Universiteit (VU) Amsterdam Boulder, Friday march 10, 2017
 Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Heritability enrichment of differentially expressed genes. Hilary Finucane PGC Statistical Analysis Call January 26, 2016
 Heritability enrichment of differentially expressed genes Hilary Finucane PGC Statistical Analysis Call January 26, 2016 1 Functional genomics + GWAS gives insight into disease relevant tissues Trynka
Heritability enrichment of differentially expressed genes Hilary Finucane PGC Statistical Analysis Call January 26, 2016 1 Functional genomics + GWAS gives insight into disease relevant tissues Trynka
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data
 Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics. Mike West Duke University
 Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
Increased methylation variation in epigenetic domains across cancer types
 Increased methylation variation in epigenetic domains across cancer types Kasper Daniel Hansen 1,2,1, Winston Timp 2 4,1, Héctor Corrada Bravo 2,5,1, Sarven Sabunciyan 2,6,1, Benjamin Langmead 1,2,1, Oliver
Increased methylation variation in epigenetic domains across cancer types Kasper Daniel Hansen 1,2,1, Winston Timp 2 4,1, Héctor Corrada Bravo 2,5,1, Sarven Sabunciyan 2,6,1, Benjamin Langmead 1,2,1, Oliver
Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors
 Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors The Harvard community has made this article openly available. Please share how this access benefits you.
Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors The Harvard community has made this article openly available. Please share how this access benefits you.
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Back to the Basics: Methyl-Seq 101
 Back to the Basics: Methyl-Seq 101 Presented By: Alex Siebold, Ph.D. October 9, 2013 Field Applications Scientist Agilent Technologies Life Sciences & Diagnostics Group Life Sciences & Diagnostics Group
Back to the Basics: Methyl-Seq 101 Presented By: Alex Siebold, Ph.D. October 9, 2013 Field Applications Scientist Agilent Technologies Life Sciences & Diagnostics Group Life Sciences & Diagnostics Group
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Integrative DNA methylome analysis of pan-cancer biomarkers in cancer discordant monozygotic twin-pairs
 Roos et al. Clinical Epigenetics (2016) 8:7 DOI 10.1186/s13148-016-0172-y RESEARCH Integrative DNA methylome analysis of pan-cancer biomarkers in cancer discordant monozygotic twin-pairs Open Access Leonie
Roos et al. Clinical Epigenetics (2016) 8:7 DOI 10.1186/s13148-016-0172-y RESEARCH Integrative DNA methylome analysis of pan-cancer biomarkers in cancer discordant monozygotic twin-pairs Open Access Leonie
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data
 A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data Leonardo Bottolo 1,2,3 1 Department of Medical Genetics, University of Cambridge, UK 2 The Alan Turing Institute, London,
A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data Leonardo Bottolo 1,2,3 1 Department of Medical Genetics, University of Cambridge, UK 2 The Alan Turing Institute, London,
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery
 CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
Exercises: Differential Methylation
 Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome
 The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi
 Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Supplemental Information. A Highly Sensitive and Robust Method. for Genome-wide 5hmC Profiling. of Rare Cell Populations
 Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Computational aspects of ChIP-seq. John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory
 Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Selection and Combination of Markers for Prediction
 Selection and Combination of Markers for Prediction NACC Data and Methods Meeting September, 2010 Baojiang Chen, PhD Sarah Monsell, MS Xiao-Hua Andrew Zhou, PhD Overview 1. Research motivation 2. Describe
Selection and Combination of Markers for Prediction NACC Data and Methods Meeting September, 2010 Baojiang Chen, PhD Sarah Monsell, MS Xiao-Hua Andrew Zhou, PhD Overview 1. Research motivation 2. Describe
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Transcript reconstruction
 Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Note. Nature Genetics: doi: /ng.2928
 Supplementary Note Loss of heterozygosity analysis (LOH). We used VCFtools v0.1.11 to extract only singlenucleotide variants with minimum depth of 15X and minimum mapping quality of 20 to create a ped
Supplementary Note Loss of heterozygosity analysis (LOH). We used VCFtools v0.1.11 to extract only singlenucleotide variants with minimum depth of 15X and minimum mapping quality of 20 to create a ped
Lecture 8 Understanding Transcription RNA-seq analysis. Foundations of Computational Systems Biology David K. Gifford
 Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Transcript-indexed ATAC-seq for immune profiling
 Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data
 DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Sudin Bhattacharya Institute for Integrative Toxicology
 Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
DNA methylation & demethylation
 DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
DNA methylation & demethylation Lars Schomacher (Group Christof Niehrs) What is Epigenetics? Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not
DiffVar: a new method for detecting differential variability with application to methylation in cancer and aging
 Genome Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. DiffVar: a new method for detecting
Genome Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. DiffVar: a new method for detecting
Pre-mRNA Secondary Structure Prediction Aids Splice Site Recognition
 Pre-mRNA Secondary Structure Prediction Aids Splice Site Recognition Donald J. Patterson, Ken Yasuhara, Walter L. Ruzzo January 3-7, 2002 Pacific Symposium on Biocomputing University of Washington Computational
Pre-mRNA Secondary Structure Prediction Aids Splice Site Recognition Donald J. Patterson, Ken Yasuhara, Walter L. Ruzzo January 3-7, 2002 Pacific Symposium on Biocomputing University of Washington Computational
Mapping by recurrence and modelling the mutation rate
 Mapping by recurrence and modelling the mutation rate Shamil Sunyaev Broad Institute of M.I.T. and Harvard Current knowledge is from Comparative genomics Experimental systems: yeast reporter assays Potential
Mapping by recurrence and modelling the mutation rate Shamil Sunyaev Broad Institute of M.I.T. and Harvard Current knowledge is from Comparative genomics Experimental systems: yeast reporter assays Potential
Introduction to Genetics and Genomics
 2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
Title: Pinpointing resilience in Bipolar Disorder
 Title: Pinpointing resilience in Bipolar Disorder 1. AIM OF THE RESEARCH AND BRIEF BACKGROUND Bipolar disorder (BD) is a mood disorder characterised by episodes of depression and mania. It ranks as one
Title: Pinpointing resilience in Bipolar Disorder 1. AIM OF THE RESEARCH AND BRIEF BACKGROUND Bipolar disorder (BD) is a mood disorder characterised by episodes of depression and mania. It ranks as one
Epigenetics and Chromatin Remodeling
 Epigenetics and Chromatin Remodeling Bradford Coffee, PhD, FACMG Emory University Atlanta, GA Speaker Disclosure Information Grant/Research Support: none Salary/Consultant Fees: none Board/Committee/Advisory
Epigenetics and Chromatin Remodeling Bradford Coffee, PhD, FACMG Emory University Atlanta, GA Speaker Disclosure Information Grant/Research Support: none Salary/Consultant Fees: none Board/Committee/Advisory
Corporate Medical Policy
 Corporate Medical Policy Microarray-based Gene Expression Testing for Cancers of Unknown File Name: Origination: Last CAP Review: Next CAP Review: Last Review: microarray-based_gene_expression_testing_for_cancers_of_unknown_primary
Corporate Medical Policy Microarray-based Gene Expression Testing for Cancers of Unknown File Name: Origination: Last CAP Review: Next CAP Review: Last Review: microarray-based_gene_expression_testing_for_cancers_of_unknown_primary
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Canadian Bioinforma1cs Workshops
 5/12/16 Canadian Bioinforma1cs Workshops www.bioinforma1cs.ca Module #: Title of Module 2 1 Module 3 Introduc1on to WGBS and analysis Guillaume Bourque Learning Objec/ves of Module Know the different technologies
5/12/16 Canadian Bioinforma1cs Workshops www.bioinforma1cs.ca Module #: Title of Module 2 1 Module 3 Introduc1on to WGBS and analysis Guillaume Bourque Learning Objec/ves of Module Know the different technologies
Ambient temperature regulated flowering time
 Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
What can lead to aneuploidy?
 11.06.17 What can lead to aneuploidy? A Mitotic checkpoint defects B Cohesion defect C Centrosome amplification D Hyperstabilized kinetochore microtubule interactions IB SL Biology Exam. Deadlines and
11.06.17 What can lead to aneuploidy? A Mitotic checkpoint defects B Cohesion defect C Centrosome amplification D Hyperstabilized kinetochore microtubule interactions IB SL Biology Exam. Deadlines and
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Large-scale identity-by-descent mapping discovers rare haplotypes of large effect. Suyash Shringarpure 23andMe, Inc. ASHG 2017
 Large-scale identity-by-descent mapping discovers rare haplotypes of large effect Suyash Shringarpure 23andMe, Inc. ASHG 2017 1 Why care about rare variants of large effect? Months from randomization 2
Large-scale identity-by-descent mapping discovers rare haplotypes of large effect Suyash Shringarpure 23andMe, Inc. ASHG 2017 1 Why care about rare variants of large effect? Months from randomization 2
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
CS 6824: Tissue-Based Map of the Human Proteome
 CS 6824: Tissue-Based Map of the Human Proteome T. M. Murali November 17, 2016 Human Protein Atlas Measure protein and gene expression using tissue microarrays and deep sequencing, respectively. Alternative
CS 6824: Tissue-Based Map of the Human Proteome T. M. Murali November 17, 2016 Human Protein Atlas Measure protein and gene expression using tissue microarrays and deep sequencing, respectively. Alternative
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
R2 Training Courses. Release The R2 support team
 R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Figure S1, Beyer et al.
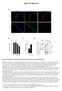 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Assignment 5: Integrative epigenomics analysis
 Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
Barrett s Esophagus and Esophageal Adenocarcinoma Epigenetic Biomarker Discovery Using Infinium Methylation
 February 2008 Application Note arrett s Esophagus and Esophageal Adenocarcinoma Epigenetic iomarker Discovery Using Infinium Methylation Contributed by Patricia C. Galipeau, Dennis L. Chao, Xiaohong Li,
February 2008 Application Note arrett s Esophagus and Esophageal Adenocarcinoma Epigenetic iomarker Discovery Using Infinium Methylation Contributed by Patricia C. Galipeau, Dennis L. Chao, Xiaohong Li,
Methylation patterns in ejaculated sperm from Nellore-Angus crossbred bulls
 Methylation patterns in ejaculated sperm from Nellore-Angus crossbred bulls M. R. Prause 1, G. Wang 1, & C. A. Gill 1 1 Texas A&M University, Department of Animal Science, College Station, TX 77843, USA
Methylation patterns in ejaculated sperm from Nellore-Angus crossbred bulls M. R. Prause 1, G. Wang 1, & C. A. Gill 1 1 Texas A&M University, Department of Animal Science, College Station, TX 77843, USA
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks
![MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks](/thumbs/80/82116384.jpg) MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1 Lecture 27: Systems Biology and Bayesian Networks Systems Biology and Regulatory Networks o Definitions o Network motifs o Examples
MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1 Lecture 27: Systems Biology and Bayesian Networks Systems Biology and Regulatory Networks o Definitions o Network motifs o Examples
The silence of the genes: clinical applications of (colorectal) cancer epigenetics
 The silence of the genes: clinical applications of (colorectal) cancer epigenetics Manon van Engeland, PhD Dept. of Pathology GROW - School for Oncology & Developmental Biology Maastricht University Medical
The silence of the genes: clinical applications of (colorectal) cancer epigenetics Manon van Engeland, PhD Dept. of Pathology GROW - School for Oncology & Developmental Biology Maastricht University Medical
