GENETICS. Supporting Information
|
|
|
- Dwight Spencer Long
- 6 years ago
- Views:
Transcription
1 GENETI upporting Information Natural Diversity in lowering Responses of rabidopsis thaliana ausedbyvariationinatandemgenerray arah Marie Rosloski, athya heela Jali, ureshkumar Balasubramanian, Detlef Weigel and Vojislava Grbic opyright Ó 2010 by the Genetics ociety of merica DOI: /genetics
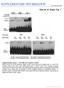
2 2 I. M. Rosloski et al. M2-ol var1 var2 B neg M2- UWO M2- ol M2 TUB IGURE 1. The banding profile of the M2-UWO variant is heritable and co-dominant. () Primer 44 and 45 sites and RT-PR products produced in M2-ol. (B) RT-PR banding profile of the M2-UWO variant, M2-ol and three independent 1 progeny of these parents.
3 . M. Rosloski et al. 3 I EG72 TTED2 0.5 cm M Locus 6.5 cm Expected 3.3 cm 7.3 cm Observed EG72 TTED2 RT-PR banding pattern hromosome 5 IGURE 2..The RT-PR banding pattern of the M2-UWO variant maps in the vicinity of the M cluster. Expected distances were obtained from the Lister and Dean map (
4 4 I. M. Rosloski et al. M3-ol M3-UWO M2-UWO M2-ha and Kas-1 2 last exon 3 last exon 3i 2i 3i * 2i 3i M4 2 last exon 2i 3i * 2i 3i 2 last exon 2i 3i * 2i M4 M3 100 bp M4 deleted region 2i M3 3i 1 Kb in deleted region B 466 bp deletion M2-Kas-1 IGURE 3..() ha and Kas-1 DN sequence downstream of the M2 coding region shows that M3 has been deleted in these two accessions. 2i and 3i, are sequences found downstream of the M2 or M3 loci in ol, respectively; *, refers to a region that cannot be definitively assigned to M2 or M3 because of the shared sequence identity. (B) M2-Kas-1 allele showing the location of the 466 bp deletion.
5 . M. Rosloski et al. 5 I M2-Ler var1 var2 M2-UWO B var1 var2 var3 var4 var5 M2-Tu-1 var1 B var2 var3 L var4 L var5 M2-ha E var1 var2 L var3 var4 M2 exon M3 exon Intron Translational stop-site 1 Kb IGURE 4..cDN sequence comparison between M2-ol, M2-Ler, M2-UWO, M2-ha and M2-Tu-1. Both M2-ha and M2-Tu-1 cdn sequences were aligned to the genomic region of M2-UWO to facilitate comparison of splice-site selection amongst the alleles.
6 6 I. M. Rosloski et al.
7 . M. Rosloski et al. 7 I IGURE 5..Predicted protein products of M2-ol, M2-UWO, M2-ha, M2 Tu-1, M3-ol and M3-UWO. Yellow box, MDs-box domain; Orange box, K-box domain.
8 8 I. M. Rosloski et al. TBLE 1 List of rabidopsis accessions used in the M2 Expression creen, n=147. Group 1: 23 accessions ng-1, BG1, BG4, BG6, EN-0, IB1 IB10, HL1, M10, M11, Gre-0, H1, Kin-0, N1, N10, NE1, REN1, Ri-0, RP1, RP10, f-2, f-2e, Zu-0 Group 2: 72 accessions Bch-3, Bd-0, Bla-4, Bla-12, Br-0, Bs-1, Bsch-2, Bu-11, Bur-0, 24, al-0, o-1, ol, vi-0, Db-0, Ei-6, EDI-0, El-0, Er-0, Est-1, Et-0, i-0, r-2, r-6, Ga-0, Gd-1, Ge-1, Got1, Got10, Gr-1, Gr-6, Gu-0, Hh-0, Hl-0, Hl-3,, Hs-0, Is-0, Kil-0, Kl-0, La-1, Ler, Li-2, Li-2:1, Ll-11, M3385, M7943s, Mc-0, Mc-1, Mh-1, Mz-0, Nc-1, Nd-1, Nok-0, Nok-1, Nok-2, Nok-3, Np-0, Nw-1, Ob-1, Old-2, Ove-0, Pa-2, Pi-0, Q4, te-0, Tsu-1, Ty-0, UK3, UK4, Wc-1 Wl-0, Wu-0 Group 3: 13 accessions Bla-1, Bla-2, Bla-5, Bla-6, Bla-11, Pla-0, Pla-2, Pog-0, Ra-0, e-0, Ts-1, Ts-5, Ts-6 Group 4: 36 accessions k-1, Blh-1, Blh-2, Bs-5, ha-0, hi-0, hi-2, Dr-0, Dra-1, e-1, Ge-2, Gr-3, Hodja-Obi, In-0, Jl-1, Jm-1, Kn-0, Kondara, KZ10, Lip-0, Lo-1, M73235, Mir-0, Nw-3, Ost-0, Per-2, Per-3, RLD1, Rsch-0, tw-0, n(5)-1, Ta-0, Te-0, UK2, UWO, Wei-0, Wil-1 Group 5: 3 accessions r-4, Ll-2, Mv-0 Representative accessions from igure 1 are in bold text.
9 . M. Rosloski et al. 9 I TBLE 2 List of rabidopsis accessions and their geographical coordinates used in the genomic DN screen Table 2 is available for download as an Excel file at
10 10 I. M. Rosloski et al. M2 Insertion llele ubclass TBLE 3 haracteristics of M2 insertion allele subclasses Number of ccessions in Genomic creen Length of M3 Insertion, bp g Gr UWO Tu KZ ha Length of M2 deletion a, bp all insertion allele subclasses, except M2-UWO, have both an insertion of the M3 gene sequence into the last exon of M2 and a deletion of M2 genomic sequence adjacent to the M3 insertion.
11 . M. Rosloski et al. 11 I TBLE 4 egregation distortion observed in the B 5 2 progeny of crosses between UWO, Tu-1, ha and Kas-1 accessions and Ler Observed Expected n x 2 p-value UWO-1 22:18:8 12:24: ** UWO-2 43:53:17 28:57: ** UWO-3 25:34:10 17:35: * UWO-4 25:34:8 17:34: * UWO total 115:139:43 74:149: *** Tu :24:15 16:31: ns Tu :36:14 16:33: ns Tu :27:13 14:28: ns Tu-1 total 54:87:42 46:92: ns ha-1 25:29:11 16:33: * ha-2 23:33:9 16:33: * ha-3 14:30:11 14:18: ns ha-total 62:92:31 46:93: ** Kas :35:10 16:32: ns Kas :28:10 14:28: ns Kas :23:10 13:25: ns Kas-1 total 53:86:30 42:85: *
12 12 I. M. Rosloski et al. TBLE 5 egregation distortion observed in the B 4 2 progeny of a cross between M2-UWO in Ler x L-ol in Ler egregating marker M2-UWO/M2-Ler n progeny ratio UWO/UWO UWO/ Ler Ler / Ler x 2 p-value 142 obs * exp obs ** exp obs * exp total 660 obs sum exp ** flc-ler/l-ol Ler / Ler Ler /ol ol/ol 142 obs n.s. exp obs n.s. exp obs n.s. exp total 660 obs sum exp n.s
13 . M. Rosloski et al. 13 I TBLE 6 Primers used in this study Primer # Primer sequence 5 > 3 Purpose PR specifications* 1 TGGTT cdn screen 1 and 2, 296 bp in ol, 55 o, 2μM, TTTTT 1.4% 2 TTGGT TTTT cdn screen 3 TTTTTT TTTTT 4 TTG TG TG GTG TG GG TT 5 TTGTGGGGG GTG 6 TTTGGTTTTT GTTTTG 7 TTGGGG TTTG 8 GTTTTGGT GT 9 GGGTTTGTTG TGTTGG 10 GGT TTTTTTT 11 TGGTTTTGTT GG 12 GTTGGGTTTG TTGTG 13 GGTTGGGGTTT TGGTG 14 TTTTTTTG T 15 GTTTTTGTTTTT GTTGTTTGTTG 16 TTGGTTGGGG TTTTGGTG 17 TTGTTGTTG TGT 18 TTTTGGTG TTT 19 TTTTTT TTTTTGT 20 GGTTTTTGTTTGG TTGTG 21 TTTTTG TT 20 GGTGGTG TG 21 TGTTTGGGT TGGT 24 TTTTGTTGGTGGG GTG 25 TTGGTGTTT 26 TGGTGTTTGT TTGG 27 GTGGGG TTG 28 GTTTT TTG 29 GTTGGTGTTT G 30 GTGGGG TT Ubiquitin, cdn screen Ubiquitin, cdn screen equencing M2 alleles equencing M2 alleles Genomic DN screen, 3 end of insertion Genomic DN screen, 3 end of insertion Genomic DN screen, 3 end of insertion equencing M2-ha and M2-Kas, M3 deletion equencing M2-ha and M2-Kas, M3 deletion Genotyping M2-UWO B5 2 populations Genotyping M2-UWO B5 2 populations Genotyping M2-UWO B5 2 populations Genomic DN screen, 5 end of insertion, Genotyping M2-ha B5 2 populations Genomic DN screen, 5 end of insertion Genotyping M2-ha B5 2 populations Genotyping M2-ha B5 2 populations Genotyping M2-Tu-1 B5 2 populations Genotyping M2-Tu-1 B5 2 populations Genotyping M2-Tu-1 B5 2 populations equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M2 alleles equencing M3-UWO allele 3 and 4, 415 bp, 55 o, 2μM, 32x, 1% 5 and 6, 1167 bp in UWO, 56 o, 1.2μM, 7, 8 and 9, 7 and 9 amplify 739 bp in ol, 7 and 8 amplify 638 in UWO, 56 o, 2μM, 10 and 11, 405 bp in ha, 54 o, 2μM, 12, 13 and 14, 12 and 14 amplify a 212 bp fragment in Ler, 12 and 13 amplify a 279 bp fragment in UWO, 54 o, 3μM, 15 and 16, 749 bp in ha, 58 o, 2μM, 15, 17 and 18, 15 and 18 amplify a 319 bp fragment in ol, 15 and 17 amplify a 299 bp fragment in ha and Kas-1, 56 o, 2μM, 32x, 2% 19, 20 and 21, 19 and 21 amplify a 215 bp fragment in Ler, 19 and 20 amplify a 244 bp fragment in Tu-1, 56 o, 2μM, 32x, 2% 20 and 21, 850 bp, annealing, 2μM, 32x, 1 % 24 and 25, 1111 bp, 58 o, 2μM, 32 x, 1% 26 and 27, 1174 bp, 58 o, 2μM, 32 x, 1% 28 and 29, 993 bp, 58 o, 2μM, 32x, 1% 30 and 31, 1005 bp, 58 o, 2μM,
14 14 I. M. Rosloski et al. 31 TTTTGGT G 32 GGTTG TTG 33 GGGTTGTGG TTGG 34 TTGGTGGGTG 35 TGGTGGT TT 36 TTTTTTG GTG 37 TTTGGGT GGT 38 GTTTGTGGGT TGTG 39 TGGGTGGTT GG 40 TTTTTTGTG T 41 TGTGGTT T 42 TGTTGG GTTGGTT 43 TTTTTTG GTT 44 TTGTGGGTTG GTGTTGGT- 45 TGGTGTGTT TGGTGG equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M3-UWO allele equencing M2 alleles equencing M2 alleles TUB2, Loading control for expression analysis of M2- insertion alleles TUB2, Loading control for expression analysis of M2- insertion alleles Expression analysis of M2- insertion alleles Expression analysis of M2- insertion alleles 32 and 33, 1063 bp, 58 o, 2μM, 34 and 35, 1155 bp, 58 o, 2μM, 36 and 39, 1746 bp, 58 o, 2μM, 37 and 38, band size, 58 o, 2μM, 36 and 39, 1746 bp, 58 o, 2μM, 40 and 41, 2354 bp in UWO, 56 o, 2μM, 30x, 0.8% 42 and 43, 920 bp, 60 o, 1μM, 22x, 0.8% 44 and 45, band sizes ranging from 600 to 1100 bp, 60 o, 1.5μM, 27x, % 47 GTGGGTTG T 48 GTTTGTT GTGT 49 GTGTTTGG TGG 50 GTTGTGTGGT GG 51 GTTTTTT TGTGTT 52 GGGTGG TTTT 53 GTTTTTGTGTGTT TGTTTTTTG 54 GGTTTTTTGT GGTT 55 TGTTTT 56 GTTGGTT TGTG 57 GTTGTTTGGTG GGTTTTTT TGTTT 58 GGGGTTTT TTTGG TTG 59 GGGTTTTT TTTTGGT GTGTTG 60 GTGTTGGTG GGTTTTT TTTT Mapping M2-UWO Mapping M2-UWO Mapping M2-UWO Mapping M2-UWO equencing M2-UWO 3 intergenic region equencing M2-ha and M2-Kas, M3 deletion equencing M2-UWO 3 intergenic region equencing M2-UWO 3 intergenic region equencing 1128 bp region of M3 equencing 1128 bp region of M3 I mir-s II mir-a III mir*s IV mir*a 47 and 48 + EcoRV digest, 3 fragments of 0.35, 0.23, 0.08 kb in ol and 2 fragments of kb in UWO 53 o, 1.5μM, 49 and 50 + XbaI digest, a single 1.2 Kb fragment is seen in ol and a 0.7 and 0.5 Kb fragment is seen in UWO 53 o, 1.5μM, 51 and 52, 947 bp in ha and Kas- 1, 56 o, 2μM, 53 and 54, 1181 bp, 56 o, 3μM, 55 and 56, 1128 bp, 55 o, 2 μm, 57 and 62, 300 bp, 55 o, 2.5μM, 58 and 59, 176 bp, 55 o, 2.5μM, 58 and 59, 176 bp, 55 o, 2.5μM, 60 and 61, 274 bp, 55 o, 2.5μM,
15 61 TGGGGTTG TTGGGT 62 GGGTTTT GGG Primer details from Materials and Methods. M. Rosloski et al. 15 I mir319a backbone specific primers mir319a backbone specific primers 61 and 62, 699 bp, 55 o, 2.5μM, 61 and 62, 699 bp, 55 o, 2.5μM, * PR information includes: primer combination, expected band size (bp), annealing temperature, Mgl 2 concentration, cycle number, % agarose for band resolution.
Supplementary Information for. Autoimmunity as a mechanism for hybrid necrosis, a genetic incompatibility syndrome in plants
 for Autoimmunity as a mechanism for hybrid necrosis, a genetic incompatibility syndrome in plants Kirsten Bomblies, Janne Lempe, Petra Epple, Norman Warthmann, Christa Lanz, Jeffery L. Dangl, and Detlef
for Autoimmunity as a mechanism for hybrid necrosis, a genetic incompatibility syndrome in plants Kirsten Bomblies, Janne Lempe, Petra Epple, Norman Warthmann, Christa Lanz, Jeffery L. Dangl, and Detlef
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
 1 2 APPENDIX Legends to figures 3 4 5 Figure A1: Distribution of perfect SSR along chromosome 1 of V. cholerae (El-Tor N191). a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
1 2 APPENDIX Legends to figures 3 4 5 Figure A1: Distribution of perfect SSR along chromosome 1 of V. cholerae (El-Tor N191). a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Natural variation in Arabidopsis, a tool to identify genetic bases of nitrogen use efficiency
 Natural variation in rabidopsis, a tool to identify genetic bases of nitrogen use efficiency. Chaillou, F. Chardon, J. Laurette,. Ikram, O. Loudet and F. Daniel-Vedele IN Versailles Plants have different
Natural variation in rabidopsis, a tool to identify genetic bases of nitrogen use efficiency. Chaillou, F. Chardon, J. Laurette,. Ikram, O. Loudet and F. Daniel-Vedele IN Versailles Plants have different
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
SUPPLEMENTAL FIGURE 1
 SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
The natural genetic variation of the fatty-acyl composition of seed oils in di erent ecotypes of Arabidopsis thaliana
 Phytochemistry 52 (1999) 1029±1033 The natural genetic variation of the fatty-acyl composition of seed oils in di erent ecotypes of Arabidopsis thaliana Anthony A. Millar, Ljerka Kunst* Department of Botany,
Phytochemistry 52 (1999) 1029±1033 The natural genetic variation of the fatty-acyl composition of seed oils in di erent ecotypes of Arabidopsis thaliana Anthony A. Millar, Ljerka Kunst* Department of Botany,
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplemental Figures Legends and Supplemental Figures. for. pirna-guided slicing of transposon transcripts enforces their transcriptional
 Supplemental Figures Legends and Supplemental Figures for pirn-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear pirn repertoire Kirsten-ndré
Supplemental Figures Legends and Supplemental Figures for pirn-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear pirn repertoire Kirsten-ndré
Supplemental Data. Di Giorgio et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Synteny analysis of NIP4;1 and NIP4;2. Examination of the NIP4;1-NIP4;2 region in rabidopsis thaliana and equivalent evolutionary regions in other dicot genomes are shown on the
Supplemental Figure 1. Synteny analysis of NIP4;1 and NIP4;2. Examination of the NIP4;1-NIP4;2 region in rabidopsis thaliana and equivalent evolutionary regions in other dicot genomes are shown on the
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Figure S1. Analysis of genomic and cdna sequences of the targeted regions in WT-KI and
 Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at
 Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
McAlpine PERK-GSK3 regulates foam cell formation. Supplemental Material. Supplementary Table I. Sequences of real time PCR primers.
 Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
EGFR shrna A: CCGGCGCAAGTGTAAGAAGTGCGAACTCGAGTTCGCACTTCTTACACTTGCG TTTTTG. EGFR shrna B: CCGGAGAATGTGGAATACCTAAGGCTCGAGCCTTAGGTATTCCACATTCTCTT TTTG
 Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Prospective study of MTHFR genetic polymorphisms as a possible etiology of male infertility
 Prospective study of MTHFR genetic polymorphisms as a possible etiology of male infertility S.-S. Li 1, J. Li 1, Z. Xiao 2, A.-G. Ren 3 and L. Jin 3 1 Beijing Obstetrics and Gynecology Hospital, Capital
Prospective study of MTHFR genetic polymorphisms as a possible etiology of male infertility S.-S. Li 1, J. Li 1, Z. Xiao 2, A.-G. Ren 3 and L. Jin 3 1 Beijing Obstetrics and Gynecology Hospital, Capital
Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice
 Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
HIV-1 resistance testing from proviral DNA
 HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
Supplementary Information
 Supplementary Information Overexpression of Fto leads to increased food intake and results in obesity Chris Church, Lee Moir, Fiona McMurray, Christophe Girard, Gareth T Banks, Lydia Teboul, Sara Wells,
Supplementary Information Overexpression of Fto leads to increased food intake and results in obesity Chris Church, Lee Moir, Fiona McMurray, Christophe Girard, Gareth T Banks, Lydia Teboul, Sara Wells,
CD31 5'-AGA GAC GGT CTT GTC GCA GT-3' 5 ' -TAC TGG GCT TCG AGA GCA GT-3'
 Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Mendelian & Complex Traits. Quantitative Imaging Genomics. Genetics Terminology 2. Genetics Terminology 1. Human Genome. Genetics Terminology 3
 Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
Relative activity (%) SC35M
 a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
*To whom correspondence should be addressed. This PDF file includes:
 www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
SUPPLEMENTARY INFORMATION
 SPPLEMENTY INFOMTION 2 =.75 SHPE + 1 (eplicate 2) 2 1 1 2 SHPE + 1 (eplicate 1) Figure S1. eproducibility of SHPE measurements between biological replicates. eactivities corresponding to extension reactions
SPPLEMENTY INFOMTION 2 =.75 SHPE + 1 (eplicate 2) 2 1 1 2 SHPE + 1 (eplicate 1) Figure S1. eproducibility of SHPE measurements between biological replicates. eactivities corresponding to extension reactions
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
A Sound Track to Reading
 A Sound Track to Reading Blending Flashcards Prepared by Donald L Potter June 1, 2018 Mr. Potter prepared these cards to be used with Sister Monica Foltzer s advanced intensive phonics program and reader,
A Sound Track to Reading Blending Flashcards Prepared by Donald L Potter June 1, 2018 Mr. Potter prepared these cards to be used with Sister Monica Foltzer s advanced intensive phonics program and reader,
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Figure S1. Molecular confirmation of the precise insertion of the AsMCRkh2 cargo into the kh w locus.
 Supporting Information Appendix Table S1. Larval and adult phenotypes of G 2 progeny of lines 10.1 and 10.2 G 1 outcrosses to wild-type mosquitoes. Table S2. List of oligonucleotide primers. Table S3.
Supporting Information Appendix Table S1. Larval and adult phenotypes of G 2 progeny of lines 10.1 and 10.2 G 1 outcrosses to wild-type mosquitoes. Table S2. List of oligonucleotide primers. Table S3.
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards
 Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
Supporting Information
 Supporting Information hatnagar et al. 1.173/pnas.141362111 SI Materials and Methods Single-Nucleotide Primer Extension ssay. single-nucleotide primer extension (SNuPE) assay for was carried out using
Supporting Information hatnagar et al. 1.173/pnas.141362111 SI Materials and Methods Single-Nucleotide Primer Extension ssay. single-nucleotide primer extension (SNuPE) assay for was carried out using
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Supplementary Figure 1 a
 Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE.
 ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE. Batalla A, Coto E*, González-Lara L, González- Fernández D, Maldonado-Seral C, García-García
ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE. Batalla A, Coto E*, González-Lara L, González- Fernández D, Maldonado-Seral C, García-García
Ambient temperature regulated flowering time
 Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
Supplemental Figure 1. Small RNA size distribution from different soybean tissues.
 Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
Cancer Genetics 204 (2011) 45e52
 Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
Problem set questions from Final Exam Human Genetics, Nondisjunction, and Cancer
 Problem set questions from Final Exam Human Genetics, Nondisjunction, and ancer Mapping in humans using SSRs and LOD scores 1. You set out to genetically map the locus for color blindness with respect
Problem set questions from Final Exam Human Genetics, Nondisjunction, and ancer Mapping in humans using SSRs and LOD scores 1. You set out to genetically map the locus for color blindness with respect
Suppl. Figure 1. NF-κB. gastrocnemius. WT mdx dko. mdx. pelk1. pets2. Ras. Ets2. praf. pp38. pmek-1. p38 SOS. p90 Rsk
 Suppl. Figure 1 NF-κ p65 gastrocnemius phase merge D pets2 dko E pelk1 dko Ets2 Ras pp38 p38 praf pmek-1 SOS p90 Rsk Suppl. Figure 2 pair 1 pair 2 NF-# pair1 pair 2 p65 p50 Oct-1 ;p50+ / - D dys.!-sg "-DG
Suppl. Figure 1 NF-κ p65 gastrocnemius phase merge D pets2 dko E pelk1 dko Ets2 Ras pp38 p38 praf pmek-1 SOS p90 Rsk Suppl. Figure 2 pair 1 pair 2 NF-# pair1 pair 2 p65 p50 Oct-1 ;p50+ / - D dys.!-sg "-DG
(43) Publication date: 04 September 2014 ( ) (22) Filing Date: 27 February 2014 ( )
 (54) Title (EN): INCOHERENT TYPE-III MATERIALS FOR CHARGE CARRIERS CONTROL DEVICES (54) Title (FR): MATÉRIAUX DE TYPE III INCOHÉRENTS POUR DISPOSITIFS DE RÉGULATION DE PORTEURS DE CHARGES (72) Inventor(s):
(54) Title (EN): INCOHERENT TYPE-III MATERIALS FOR CHARGE CARRIERS CONTROL DEVICES (54) Title (FR): MATÉRIAUX DE TYPE III INCOHÉRENTS POUR DISPOSITIFS DE RÉGULATION DE PORTEURS DE CHARGES (72) Inventor(s):
Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women
 www.bioinformation.net Volume 13(5) Hypothesis Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women Maniraja Jesintha
www.bioinformation.net Volume 13(5) Hypothesis Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women Maniraja Jesintha
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Germline mutations in shelterin complex genes are associated with familial chronic lymphocytic leukemia
 UEMEN NOMON ermline mutations in shelterin complex genes are associated with familial chronic lymphocytic leukemia Helen E. peedy 1, Ben Kinnersley 1, Daniel Chubb 1, eter Broderick 1, hilip J. aw 1, Kevin
UEMEN NOMON ermline mutations in shelterin complex genes are associated with familial chronic lymphocytic leukemia Helen E. peedy 1, Ben Kinnersley 1, Daniel Chubb 1, eter Broderick 1, hilip J. aw 1, Kevin
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients
 Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
IVF Michigan, Rochester Hills, Michigan, and Reproductive Genetics Institute, Chicago, Illinois
 FERTILITY AND STERILITY VOL. 80, NO. 4, OCTOBER 2003 Copyright 2003 American Society for Reproductive Medicine Published by Elsevier Inc. Printed on acid-free paper in U.S.A. CASE REPORTS Preimplantation
FERTILITY AND STERILITY VOL. 80, NO. 4, OCTOBER 2003 Copyright 2003 American Society for Reproductive Medicine Published by Elsevier Inc. Printed on acid-free paper in U.S.A. CASE REPORTS Preimplantation
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Supplementary Table 2. Conserved regulatory elements in the promoters of CD36.
 Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer
 AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
Spherical Bearings Heavy Duty Equipments
 Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table.
 Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tale. Type of file: VI Title of file for HTML: Supplementary Movie 1 Description:
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tale. Type of file: VI Title of file for HTML: Supplementary Movie 1 Description:
gliomas. Fetal brain expected who each low-
 Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Drug Metabolism Disposition
 Drug Metabolism Disposition The CYP2C19 intron 2 branch point SNP is the ancestral polymorphism contributing to the poor metabolizer phenotype in livers with CYP2C19*35 and CYP2C19*2 alleles Amarjit S.
Drug Metabolism Disposition The CYP2C19 intron 2 branch point SNP is the ancestral polymorphism contributing to the poor metabolizer phenotype in livers with CYP2C19*35 and CYP2C19*2 alleles Amarjit S.
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
SALSA MLPA KIT P050-B2 CAH
 SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
What do you think of when you here the word genome?
 What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
Validation of the MIA FORA NGS FLEX Assay Using Buccal Swabs as the Sample Source
 S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W
 Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
Annotation of Drosophila mojavensis fosmid 8 Priya Srikanth Bio 434W 5.1.2007 Overview High-quality finished sequence is much more useful for research once it is annotated. Annotation is a fundamental
The Alternative Choice of Constitutive Exons throughout Evolution
 The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
Supplementary Material
 Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
Supplemental Information For: The genetics of splicing in neuroblastoma
 Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
Supplemental Information For: The genetics of splicing in neuroblastoma Justin Chen, Christopher S. Hackett, Shile Zhang, Young K. Song, Robert J.A. Bell, Annette M. Molinaro, David A. Quigley, Allan Balmain,
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
PREVENTION OF HAEMOGLOBINOPATHIES: New methodologies and procedures Non-invasive Prenatal Diagnosis
 PREVENTION OF HAEMOGLOBINOPATHIES: New methodologies and procedures Non-invasive Prenatal Diagnosis Marina Kleanthous Cyprus School of Molecular Medicine The Cyprus Institute of Neurology and Genetics
PREVENTION OF HAEMOGLOBINOPATHIES: New methodologies and procedures Non-invasive Prenatal Diagnosis Marina Kleanthous Cyprus School of Molecular Medicine The Cyprus Institute of Neurology and Genetics
Prevalence of HPV-16 genomic variant carrying a 63-bp duplicated sequence within the E1 gene in Slovenian women
 Prevalence of HPV-16 genomic variant carrying a 63-bp duplicated sequence within the E1 gene in Slovenian women K E Y WORDS HPV-16, cervical cancer, E6-T350G variant A BSTRACT High-risk HPV, particularly
Prevalence of HPV-16 genomic variant carrying a 63-bp duplicated sequence within the E1 gene in Slovenian women K E Y WORDS HPV-16, cervical cancer, E6-T350G variant A BSTRACT High-risk HPV, particularly
SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A
 SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A1-1011. This SALSA MLPA probemix is for basic research and intended for experienced MLPA users only! This probemix enables you to quantify genes
SALSA MLPA probemix P360-A1 Y-Chromosome Microdeletions Lot A1-1011. This SALSA MLPA probemix is for basic research and intended for experienced MLPA users only! This probemix enables you to quantify genes
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
SpliceDB: database of canonical and non-canonical mammalian splice sites
 2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
MRC-Holland MLPA. Description version 13;
 SALSA MLPA probemix P027-C1 Uveal Melanoma Lot C1-0211: A large number of probes have been replaced by other probes in the same chromosomal regions as compared to previous lots, and several reference probes
SALSA MLPA probemix P027-C1 Uveal Melanoma Lot C1-0211: A large number of probes have been replaced by other probes in the same chromosomal regions as compared to previous lots, and several reference probes
Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report. a a*
 2009 63 6 379 384 Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report a b a a a* a b 380 63 6 Chest x ray and computed tomography (CT). A, Chest x ray on admission
2009 63 6 379 384 Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report a b a a a* a b 380 63 6 Chest x ray and computed tomography (CT). A, Chest x ray on admission
Supplementary Figure 1
 Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described
 Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
DCP Networks
 www.bps.org.uk/dcp DCP Networks To support and promote clinical psychology, working collaboratively with people who use our services and wider communities to improve wellbeing and to promote the unique
www.bps.org.uk/dcp DCP Networks To support and promote clinical psychology, working collaboratively with people who use our services and wider communities to improve wellbeing and to promote the unique
MRC-Holland MLPA. Description version 08; 30 March 2015
 SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany
 HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
Astaxanthin prevents and reverses diet-induced insulin resistance and. steatohepatitis in mice: A comparison with vitamin E
 Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
Probe. Hind III Q,!?R'!! /0!!!!D1"?R'! vector. Homologous recombination
 Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri
 Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri By Bria Collette Kettle Submitted to the graduate degree program in
Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri By Bria Collette Kettle Submitted to the graduate degree program in
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
WHAT S IN THIS LECTURE?
 What is meant by the term monogenic? WHAT S IN THIS LECTURE? WHAT S MENDEL S PRINCIPLE OF SEGREGATION? What s probability got to do with this? WHAT S MENDEL S PRINCIPLE OF INDEPENDENT ASSORTMENT? 1 FROM
What is meant by the term monogenic? WHAT S IN THIS LECTURE? WHAT S MENDEL S PRINCIPLE OF SEGREGATION? What s probability got to do with this? WHAT S MENDEL S PRINCIPLE OF INDEPENDENT ASSORTMENT? 1 FROM
MRC-Holland MLPA. Description version 19;
 SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
