Sperm tsrnas contribute to intergenerational inheritance of an acquired metabolic disorder
|
|
|
- Jasper McKenzie
- 5 years ago
- Views:
Transcription
1 SMALL RNAS Sperm tsrnas contribute to intergenerational inheritance of an acquired metabolic disorder Qi Chen, 1,3 * Menghong Yan, 2 Zhonghong Cao, 1,5 Xin Li, 1 Yunfang Zhang, 1,5 Junchao Shi, 1,5 Gui-hai Feng, 1 Hongying Peng, 1,4 Xudong Zhang, 1,5 Ying Zhang, 1 Jingjing Qian, 1,5 Enkui Duan, 1 * Qiwei Zhai, 2 * Qi Zhou 1 * Increasing evidence indicates that metabolic disorders in offspring can result from the father s diet, but the mechanism remains unclear. In a paternal mouse model given a high-fat diet (HFD), we showed that a subset of sperm transfer RNA derived small RNAs (tsrnas), mainly from 5 transfer RNA halves and ranging in size from 30 to 34 nucleotides, exhibited changes in expression profiles and RNA modifications. Injection of sperm tsrna fractions from HFD males into normal zygotes generated metabolic disorders in the F 1 offspring and altered gene expression of metabolic pathways in early embryos and islets of F 1 offspring, which was unrelated to DNA methylation at CpG-enriched regions. Hence, sperm tsrnas represent a paternal epigenetic factor that may mediate intergenerational inheritance of diet-induced metabolic disorders. Increasing lines of evidence from worms to mammals suggest that parental environmental exposure can affect the germ line and influence future generations through epigenetic mechanisms (1, 2). Specifically, diet-induced metabolic changes in mammals are transmitted from father to offspring (3, 4), suggesting sperm-mediated epigenetic inheritance (5). DNA methylation is affected (6, 7), yet a causal relationship in transgenerational inheritance has not been established. Small noncoding RNAs (sncrnas) regulate DNA methylation, histone modifications, and mrna transcription (8) and can induce non-mendelian transgenerational inheritance in mammals (9 13). Altered sperm microrna (mirna) profiles have been observed after paternal exposure to dietary changes or trauma (14, 15); however, because mammalian sperm harbors a diversity of sncrnas(16), the specific population of RNAs that mediate intergenerational epigenetic memory remains unknown (17, 18). Here we report that a subset of sperm trna derived small RNAs (tsrnas), mainly from 5 trna halves and about 30 to 34 nucleotides (nt) in size (19), showed alterations in expression profiles and RNA modifications after paternal exposure to a high-fat diet and transmitted certain metabolic disorders from father to offspring. To establish a model of intergenerational transmission of paternal diet induced metabolic disorder (4, 7, 14), we continuously fed F 0 male mice with a high-fat diet (HFD, 60% fat) or a normal diet (ND, 10% fat) for 6 months beginning at 5 weeks of age. As expected, males fed a HFD became obese, glucose intolerant, and insulinresistant,whereasmalesinthendgroup did not (fig. S1). The sperm heads of ND and HFD mice were injected into normal mouse oocytes, and the embryos were transferred into surrogate mothers. Male offspring resulting from the HFDand ND-group sperm were fed a ND and exhibited no obvious differences in body weight over 16 weeks (fig. S2A). However, offspring produced by the HFD-group sperm exhibited the onset of impaired glucose tolerance and insulin resistance as early as 7 weeks of age (fig. S2, B and C), which became more severe at 15 weeks, as revealed by a glucose tolerance test (GTT) and an insulin tolerance test (ITT) (fig. S2, D and F). Although embryo manipulation procedures may induce epigenetic alterations, our parallel sperm-head injection experiments have eliminated the potential influence of male-female contact and semen factors during natural mating (20), suggesting that the sperm itself contains sufficient information 1 State Key Laboratory of Stem Cell and Reproductive Biology, Institute of Zoology, Chinese Academy of Sciences, Beijing , China. 2 Key Laboratory of Nutrition and Metabolism, Chinese Academy of Sciences Center for Excellence in Molecular Cell Science, Institute for Nutritional Sciences, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai , China. 3 Department of Physiology and Cell Biology, School of Medicine, University of Nevada, Reno, NV USA. 4 Beijing Royal Integrative Medicine Hospital, Beijing University of Chinese Medicine, Beijing, China. 5 University of Chinese Academy of Sciences, Beijing , China. *Corresponding author. cqi@medicine.nevada.edu (Q.C.); duane@ioz.ac.cn (E.D.); qwzhai@sibs.ac.cn (Q.Zha.); qzhou@ioz. ac.cn (Q.Zho.) These authors contributed equally to this work. Downloaded from on February 10, 2016 Fig. 1. Inheritance by F 1 offspring of paternally acquired metabolic disorder via sperm RNAs. (A to D) F 1 male offspring obtained by injecting sperm total RNAs from the HFD or ND groups into normal zygotes on a CD-1 background were examined for (A) growth curves (n =12NDand45HFDmice per group), (B) blood glucose during the GTT, (C) serum insulin during the GTT, and (D) blood glucose during the ITT. (GTT and ITT, n = 8 mice per group). Results are show as means ± SEM. AUC, area under curve. *P <0.05;**P <0.01; ***P < 0.001; NS, not significant (P >0.05).(E) Comparison of sperm tsrnas and mirnas from the HFD and ND males. SCIENCE sciencemag.org 22 JANUARY 2016 VOL 351 ISSUE
2 Fig. 2. Sperm tsrnas conferred paternally acquired metabolic disorder to F 1 offspring. (A) PAGE image of sperm total RNAs and isolated RNA fragments. (B) Growth curves for F 1 male offspring generated by injecting sperm tsrnas (30- to 40-nt RNAs) from the HFD or ND group (n = 12 ND and 32 HFD mice per group). (C to E) F 1 male offspring generated by injecting HFDgroup sperm tsrnas, ND-group sperm tsrnas, or synthetic (syn) tsrnas into CD-1 zygotes were examined, relative to F 1 control mice, for (C) blood glucose during the GTT, (D) serum insulin during the GTT, and (E) blood glucose during the ITT (GTT and ITT, n = 8 to 14 mice per group). Results are show as means ± SEM. *P <0.05;**P <0.01 (HFD versus ND tsrnas); # P <0.05(NDversus syn tsrnas). to transmit an acquired metabolic disorder to offspring. To assess whether sperm RNAs can induce intergenerational phenotypes (15), we purified total RNAs from the sperm of both HFD and ND mice and injected them into normal zygotes (RNA injection was normalized to about the amount of 10 sperm). Again, although the male offspring from both groups exhibited similar body-weight growth (Fig. 1A), the offspring from the HFD group developed impaired glucose tolerance, showing significantly higher blood glucose and serum insulin levels during the GTT than offspring from the ND group at both 7 and 15 weeks of age (Fig. 1, B and C, and fig. S3). However, the insulin sensitivity of the HFD group s offspring, as determined by the ITT test, was comparable to that of the ND group s offspring (Fig. 1D and fig S3), which differs from the results produced by sperm-head injection (fig. S2, D and F). These data demonstrate that total RNAs from the sperm of HFD males contain the information to induce glucose intolerance, but not insulin resistance, in the F 1 offspring, suggesting the involvement of other layers of regulations such as DNA methylation and histone modifications (7, 21). To identify the subpopulation of sperm RNAs that mediates paternally acquired metabolic disorder, we examined the sperm sncrna profiles of the HFD and ND F 0 males by small RNA seq. (18 to 40 nt) (fig. S4). Analysis of the sequencing data reinforced our recent discovery (19)that,in addition to the well-known mirna population, mature mouse sperm carry tsrnas (Fig. 1E). Comparative analysis of sperm small RNAs from HFD and ND mice showed that a larger proportion of tsrnas (11.53%) exhibited significant differences compared with mirnas (3.23%) (fig. S5 and tables S1 and S2), suggesting that the population of sperm tsrnas is more sensitive to HFD exposure. To further investigate whether sperm tsrnas or other sperm RNA fragments are able to induce intergenerational transmission of acquired traits, we ran sperm RNAs on a 15% PAGE (polyacrylamide gel electrophoresis) gel and separately collected RNA fragments at sizes of 30 to 40 nt (predominantly tsrnas), 15 to 25 nt (predominantly mirnas), and >40 nt from both HFD and ND males; this was followed by RNA extraction and confirmation by polymerase chain reactionandrna-seq(fig.2aandfigs.s6and S7). The three types of RNA fragments were injected separately into normal zygotes, with the same concentration as that of the injected sperm total RNAs. We found that injecting 15- to 25-nt RNAs at this concentration resulted in embryo lethality, whereas injecting 30- to 40-nt or >40-nt RNAs did not (table S3). The embryonic lethal effect in the former case was probably due to the knockdown effects of mirnas on multiple mrnas, which would interfere with normal embryonic development. Injection of a 20 dilution of the 15- to 25-nt RNAs did not cause embryo lethality, nor did it cause metabolic disorder in F 1 offspring (fig. S8, A and B). Injection of >40-nt RNAs also did not cause a metabolic phenotype in F 1 offspring (fig. S8, C and D). However, injection of 30- to 40-nt RNAs from HFD and ND groups led to phenotypes that mimicked those in the F 1 offspring that were produced by injecting sperm total RNAs: There was no significant difference in body-weight growth between HFD and ND offspring (Fig. 2B), but male offspring of the HFD group developed glucose intolerance (evident in the GTT; Fig. 2, C and D, and figs. S9 and S10) compared with those of of ND group, though with mild or no insulin resistance evident in the ITT (Fig. 2E and figs. S9 and S10). These results demonstrate that sperm 30- to 40-nt RNAs (predominantly tsrnas) are necessarytoreproducetheeffectofspermtotal RNAs in inducing acquired metabolic disorder in offspring. We next synthesized a combination of the most highly expressed tsrnas in the sperm Fig. 3. HFD alters RNA modifications in the sperm tsrna fraction. (A) ComparisonofRNA stability between sperm tsrnas and chemically synthesized tsrnas without RNA modifications (gly, glycine; glu, glutamic acid, val, valine). (B and C) HFD-group sperm shows significantly increased m 5 C and m 2 G in (B) the tsrna fraction (30- to 40-nt) but not in (C) the mirna fraction (15- to 25-nt) of RNAs, as compared with ND-group sperm (n = 3 independent samples per group). Results are show as means ± SEM. (accounting for ~70% of sperm tsrnas; table S4) to investigate whether they could resemble the function of endogenous sperm tsrnas. However, injecting synthetic tsrnas into normal zygotes with the same protocol that was JANUARY 2016 VOL 351 ISSUE 6271 sciencemag.org SCIENCE
3 Fig. 4. tsrnas from HFD-group sperm dysregulated gene expression in early embryos and islets of F 1 offspring. (A) Heat map of differentially expressed genes in islets of F 1 offspring. (B) Scatterplot analysis for differential gene expression in eight-cell embryos and blastocysts after injecting NDor HFD-group sperm tsrnas (FPKM, fragments per kilobase of transcript per million mapped reads). (C) Venn diagrams showing number of genes with significant changes at the eight-cell and blastocyst stages (upward- and downward-pointing arrows indicate up- and down-regulation, respectively). (D) Sperm tsrna sequences preferentially match to gene promoter regions rather than to mrna coding regions. (E) Venn diagrams of tsrna-matched genes that are expressed in eight-cell embryos (blue) and significantly changed genes (red) in eight-cell embryos after injecting sperm tsrnas. used for sperm tsrnas did not induce metabolic disorder in the offspring (Fig. 2, C to E). One explanation for this result could be that the chemically synthesized tsrnas are not as stable as physiologically derived sperm tsrnas (22). The synthetic tsrnas showed faster degradation rates in zygote lysates than did the spermderived tsrnas (Fig. 3A). In contrast to the unmodified synthetic tsrnas, sperm tsrnas might harbor various RNA modifications (inherited from their trna precursors) that influence RNA stability. To systematically analyze the RNA modification profiles of HFD and ND sperm tsrnas, we appliedourrecentlydevelopedhigh-throughput quantitative approach, which is based on liquid chromatography tandem mass spectrometry (fig. S11) (23) and which simultaneously identifies and quantifies multiple types of RNA modifications (table S5) in one RNA sample. With this approach, we stably detected and quantified 10 types of RNA modifications in 30- to 40-nt sperm RNAs (predominantly tsrnas) and found that two RNA modifications, 5-methylcytidine (m 5 C) and N 2 -methylguanosine (m 2 G), were significantly up-regulated in HFD-group sperm compared with ND-group sperm (Fig. 3B), whereas no significant differences in 15- to 25-nt sperm RNAs (predominantly mirnas) were found using the same protocol (Fig. 3C). Although the function of m 2 G on RNAs in mammals remains unknown, the presence of mammalian m 5 C has been reported to contribute to trna stability (24)andto be related to RNA-mediated transgenerational epigenetic inheritance (25). Thus, the elevated levels of m 5 C and m 2 G modifications in the tsrnas of HFD-group sperm provide another clue in explaining the function of tsrnas in mediating paternally acquired traits, pointing to a new direction for future investigations. To uncover the potential causes of glucose intolerance in F 1 offspring produced by injecting HFD-group sperm tsrnas, we isolated the F 1 offspring islet and performed RNA-seq and RRBS (reduced representation bisulfite sequencing) analyses for a genome-wide comparison of both transcriptome and DNA methylation. RNA-seq analysis revealed that differentially expressed genes are dominantly enriched in metabolic pathways (including ketone, carbohydrate, and monosaccharide metabolisms) for down- but not up-regulated genes in the HFD group s offspring, as indicated by gene ontology analysis (Fig. 4A and tables S6 and S7). These changed transcription profiles in the F 1 islet could explain the observed metabolic disorder in F 1 offspring. On the other hand, genome-wide RRBS analysis revealed differentially methylated regions (DMRs) between the ND and HFD offspring in 28 genes (table S8). There were no overlaps between the DMR-associated genes and those showing transcriptional changes, suggesting that differential DNA methylation is not directly responsible for the changed transcriptional activity of the F 1 islet. To test the alternate possibility that injecting sperm tsrnas into the zygote might cause a transcriptional cascade change in the early embryo that ultimately guides altered gene expression in the islet of F 1 offspring, we collected eight-cell embryos and blastocysts after injection of ND- or HFD-group sperm tsrnas for comparative RNA-seq analysis (Fig. 4B). Both eight-cell embryos and blastocysts showed that more genes were down-regulated (869 and 486, respectively, with 75 overlaps) than up-regulated (280 and 285, respectively, with nine overlaps) in the group injected with HFD sperm tsrnas, compared with the group injected with ND sperm tsrnas. (Fig. 4C and table S9). Embryonic genes showing down- but not up-regulation in the HFD group were enriched in metabolic regulation pathways, in addition to other essential cellular processes (e.g., protein transport and localization) (tables S10 and S11). These early-embryo transcriptional changes might cause profound downstream effects that result in reprogramed gene expression in the islet of F 1 offspring, hence leading to metabolic disorder. To explore how the injection of tsrnas could potentially affect embryonic gene expression, we analyzed sequence matches throughout the genome and found that sperm tsrnas (differentially expressed between ND and HFD males) preferentially match to gene promoter regions ( 2 kb from the transcription start site) rather than coding regions (Fig. 4D and table S12). Among the 1134 genes with a tsrna-matching promoter, 922 of them are expressed in the eightcell embryos, and 62 of them showed differential expression in the eight-cell embryos (Fig. 4E). Biological pathway analysis showed that these deregulated genes have regulatory potential for diverse cellular and molecular events, including apoptosis, autophagy, oxidative stress, glucose input, and others (fig. S12). The genes Maea, Ccnc, and Deptor have been reported to be involved in pancreatic b-cell function or to be associated with diabetic conditions (26 28). This correlation-based evidence suggests that sperm tsrnas might affect metabolic gene expression through embryo to adulthood via a transcriptional cascade effect and that the deregulation of this process can affect F 1 offspring phenotypes (fig. S13). REFERENCES AND NOTES 1. L. Daxinger, E. Whitelaw, Nat. Rev. Genet. 13, (2012). 2. O. Rechavi et al., Cell 158, (2014). 3. B. R. Carone et al., Cell 143, (2010). 4. S. F. Ng et al., Nature 467, (2010). 5. O. J. Rando, Cell 151, (2012). 6. E. J. Radford et al., Science 345, (2014). 7. Y. Wei et al., Proc. Natl. Acad. Sci. U.S.A. 111, (2014). 8. D.Holoch,D.Moazed,Nat. Rev. Genet. 16, (2015). 9. V. Grandjean et al., Development 136, (2009). 10. K. D. Wagner et al., Dev. Cell 14, (2008). 11. M. Rassoulzadegan et al., Nature 441, (2006). 12. S. Yuan, D. Oliver, A. Schuster, H. Zheng, W. Yan, Sci. Rep. 5, 9266 (2015). 13. A. B. Rodgers, C. P. Morgan, N. A. Leu, T. L. Bale, Proc. Natl. Acad. Sci. U.S.A. 122, (2015). 14. T. Fullston et al., FASEB J. 27, (2013). 15. K. Gapp et al., Nat. Neurosci. 17, (2014). 16. M. Jodar, S. Selvaraju, E. Sendler, M. P. Diamond, S. A. Krawetz, Hum. Reprod. Update 19, (2013). 17. W. Yan, Mol. Cell. Endocrinol. 398, (2014). 18. R. Liebers, M. Rassoulzadegan, F. Lyko, PLOS Genet. 10, e (2014). 19. H. Peng et al., Cell Res. 22, (2012). SCIENCE sciencemag.org 22 JANUARY 2016 VOL 351 ISSUE
4 20. M. Lane, R. L. Robker, S. A. Robertson, Science 345, (2014). 21. K. Siklenka et al., Science 350, aab2006 (2015). 22. Y. Zhang et al., J. Mol. Cell Biol. 6, (2014). 23. M. Yan et al., Anal. Chem. 85, (2013). 24. F. Tuorto et al., Nat.Struct.Mol.Biol.19, (2012). 25. J. Kiani et al., PLOS Genet. 9, e (2013). 26. Y. S. Cho et al., Nat. Genet. 44, (2012). 27. M. Jiménez-Palomares et al., Am. J. Physiol. Endocrinol. Metab. 308, E450 E459 (2015). 28. C. C. Dibble, L. C. Cantley, Trends Cell Biol. 25, (2015). ACKNOWLEDGMENTS Raw data are archived in the Gene Expression Omnibus under accession number GSE This research was supported by the National Basic Research Program of China (grants 2012CBA01300 to Q.Zho., 2015CB to Q.C., and 2014CB to Q.Zha.), the Strategic Priority Research Program of the Chinese Academy of Sciences (grant XDA to Q.Zho. and E.D), and the National Natural Science Foundation of China (grants to E.D., to Q.C., to H.P., to Y.Z., to M.Y., and and to Q.Zha.). SUPPLEMENTARY MATERIALS Materials and Methods Figs. S1 to S13 Tables S1 to S12 References (29 38) 4 November 2015; accepted 11 December 2015 Published online 31 December /science.aad7977 GENE EDITING Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy Chengzu Long, 1,2,3 * Leonela Amoasii, 1,2,3 * Alex A. Mireault, 1,2,3 John R. McAnally, 1,2,3 Hui Li, 1,2,3 Efrain Sanchez-Ortiz, 1,2,3 Samadrita Bhattacharyya, 1,2,3 John M. Shelton, 4 Rhonda Bassel-Duby, 1,2,3 Eric N. Olson 1,2,3 CRISPR/Cas9-mediated genome editing holds clinical potential for treating genetic diseases, such as Duchenne muscular dystrophy (DMD), which is caused by mutations in the dystrophin gene.to correct DMD by skipping mutant dystrophin exons in postnatal muscle tissue in vivo, we used adeno-associated virus 9 (AAV9) to deliver gene-editing components to postnatal mdx mice, a model of DMD. Different modes of AAV9 delivery were systematically tested, including intraperitoneal at postnatal day 1 (P1), intramuscular at P12, and retro-orbital at P18. Each of these methods restored dystrophin protein expression in cardiac and skeletal muscle to varying degrees, and expression increased from 3 to 12 weeks after injection. Postnatal gene editing also enhanced skeletal muscle function, as measured by grip strength tests 4 weeks after injection. This method provides a potential means of correcting mutations responsible for DMD and other monogenic disorders after birth. Duchenne muscular dystrophy (DMD) is a fatal muscle disease affecting 1 in 3500 to 5000 boys. Cardiomyopathy and heart failure are common, incurable, and lethal consequences of DMD. The disease is caused by mutations in the gene encoding dystrophin, a large intracellular protein that links the dystroglycan complex at the cell surface with the underlying cytoskeleton, thereby maintaining integrity of muscle cell membranes during contraction (1, 2). In the absence of dystrophin, muscles degenerate, causing weakness and myopathy (3). Many therapeutic approaches for DMD have failed, at least in part because of the size of the dystrophin protein and the necessity for lifelong restoration of dystrophin expression in the myriad skeletal muscles of the body as well as the heart. The CRISPR (clustered regularly interspaced short palindromic repeats)/cas9 (CRISPR-associated 1 Department of Molecular Biology, University of Texas 2 Hamon Center for Regenerative Science and Medicine, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA. 3 Sen. Paul D. Wellstone Muscular Dystrophy Cooperative Research Center, University of Texas 4 Department of Internal Medicine, University of Texas *These authors contributed equally to this work. Corresponding author. eric.olson@utsouthwestern.edu protein 9) system allows precise modification of the genome and represents a potential means of correcting disease-causing mutations (4, 5). In the presence of single guide RNAs (sgrnas), Cas9 is directed to specific sites in the genome adjacent to a protospacer adjacent motif (PAM), causing a double-strand break (DSB). When provided with an additional DNA template, a precise genomic modification is generated by homology-directed repair (HDR), whereas in the absence of an exogenous template, variable indel mutations are created at the target site via nonhomologous end joining (NHEJ) (6). Previously, we used CRISPR/ Cas9 to correct a single nonsense mutation in Dmd by HDR in the germ line of mdx mice, which allowed the restoration of dystrophin protein expression (7). However, germline genomic editing is not feasible in humans (8) and HDR does not occur in postmitotic adult tissues, such as heart and skeletal muscle (9), necessitating alternative strategies of gene correction in postnatal tissues. Here, we devised a method to correct Dmd mutations by CRISPR/Cas9-mediated NHEJ (termed Myoediting ) in postnatal muscle tissues after delivery of gene-editing components by means of adeno-associated virus 9 (AAV9), which displays high tropism for muscle (10, 11). The dystrophin protein contains several domains (fig. S1), including an actin-binding domain at the N terminus, a central rod domain with a series of spectrin-like and actin-binding repeats, and WW and cysteine-rich domains at the C terminus that mediate binding to dystroglycan, dystrobrevin, and syntrophin (12). The actinbinding and cysteine-rich domains are essential for function, but many regions of the protein are dispensable (3). It has been estimated that as many as 80% of DMD patients could benefit from exon-skipping strategies that bypass mutations in nonessential regions of the gene and partially restore dystrophin expression (13). This approach has been validated in vitro by CRISPR/Cas9- mediated correction of Dmd mutations in patients induced pluripotent stem cells (14) and immortalized myoblasts (15). Similarly, adenovirus-mediated gene editing was shown to restore dystrophin expression in specific muscles of mdx mice after intramuscular injection (16), but adenoviral delivery is not therapeutically favorable (17). Shown in Fig. 1A is the strategy whereby CRISPR/Cas9-mediated NHEJ can create internal genomic deletions to bypass the premature termination codon in exon 23 responsible for the dystrophic phenotype of mdx mice, potentially allowing reconstitution of the Dmd open reading frame. In principle, this approach could be applied to many mutations within the gene, including large deletions, duplications, and pseudoexons. An advantage of this approach is that it does not require precise correction of the disease-causing mutation. Instead, imprecise deletions that prevent splicing of mutant exons are sufficient to restore dystrophin protein expression. To test whether Myoediting could be adapted to skip the Dmd mutation in exon 23 in mdx mice, wefirstevaluatedapoolofsgrnasthatpotentially target the 5 and 3 ends of exon 23 (supplementary materials, fig. S2, and table S1). We co-injected Cas9 mrna with sgrna-mdx (directed toward the mutant sequence in exon 23) and either sgrna-r3 or sgrna-l8 (targeting the 3 and 5 end of exon 23, respectively) into mdx zygotes without a HDR template (fig. S3A). Strikingly, ~80% of progeny mice lacked exon 23 (termed mdx-dex23) (fig. S3, B and C, and table S2), representing an increase in the efficiency of mdx editing relative to HDR (7). Genomic polymerase chainreaction(pcr)productsfromthetarget sites of exon 23 and reverse transcription PCR (RT-PCR) products of mdx-dex23 mice were cloned and sequenced, confirming the skipping of exon 23 (fig. S3, D to F). As a result of skipping exon 23, the open reading frame of Dmd was restored, allowing dystrophin protein expression JANUARY 2016 VOL 351 ISSUE 6271 sciencemag.org SCIENCE
5 Sperm tsrnas contribute to intergenerational inheritance of an acquired metabolic disorder Qi Chen et al. Science 351, 397 (2016); DOI: /science.aad7977 This copy is for your personal, non-commercial use only. If you wish to distribute this article to others, you can order high-quality copies for your colleagues, clients, or customers by clicking here. Permission to republish or repurpose articles or portions of articles can be obtained by following the guidelines here. The following resources related to this article are available online at (this information is current as of February 10, 2016 ): Updated information and services, including high-resolution figures, can be found in the online version of this article at: /content/351/6271/397.full.html Supporting Online Material can be found at: /content/suppl/2015/12/29/science.aad7977.dc1.html A list of selected additional articles on the Science Web sites related to this article can be found at: /content/351/6271/397.full.html#related This article cites 38 articles, 14 of which can be accessed free: /content/351/6271/397.full.html#ref-list-1 This article appears in the following subject collections: Development /cgi/collection/development Downloaded from on February 10, 2016 Science (print ISSN ; online ISSN ) is published weekly, except the last week in December, by the American Association for the Advancement of Science, 1200 New York Avenue NW, Washington, DC Copyright 2016 by the American Association for the Advancement of Science; all rights reserved. The title Science is a registered trademark of AAAS.
Transferring Fragments of Paternal Metabolism to the Offspring. Erica D. Watson 1,* UK, CB2 3EG. *Correspondence:
 Transferring Fragments of Paternal Metabolism to the Offspring Erica D. Watson 1,* 1 Department of Physiology, Development and Neuroscience, University of Cambridge, Cambridge UK, CB2 3EG. *Correspondence:
Transferring Fragments of Paternal Metabolism to the Offspring Erica D. Watson 1,* 1 Department of Physiology, Development and Neuroscience, University of Cambridge, Cambridge UK, CB2 3EG. *Correspondence:
Muscular Dystrophy. Biol 405 Molecular Medicine
 Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Gene therapy and genome editing technologies for the study and potential treatment of :
 WORKSHOP ON GENOME EDITING Gene therapy and genome editing technologies for the study and potential treatment of : Duchenne Muscular Dystrophy by Dr France Piétri-Rouxel, Institut de Myologie Centre de
WORKSHOP ON GENOME EDITING Gene therapy and genome editing technologies for the study and potential treatment of : Duchenne Muscular Dystrophy by Dr France Piétri-Rouxel, Institut de Myologie Centre de
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
The Father s Early Contribution to the Birth of the Child: the Role of Paternal RNAs. DNA Cell Biol. 1995;14:155-61
 The Father s Early Contribution to the Birth of the Child: the Role of Paternal RNAs DNA Cell Biol. 1995;14:155-61 SPERMATOGENESIS Mol. Human Reprod. (1997) 3:15-19 www.advancedfertility.com/abnormal-ivf-egg-pictures.htm
The Father s Early Contribution to the Birth of the Child: the Role of Paternal RNAs DNA Cell Biol. 1995;14:155-61 SPERMATOGENESIS Mol. Human Reprod. (1997) 3:15-19 www.advancedfertility.com/abnormal-ivf-egg-pictures.htm
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery
 Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Results. Abstract. Introduc4on. Conclusions. Methods. Funding
 . expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
. expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Name: Xueming Zhao. Professional Title: Professor. Animal embryo biotechnology, mainly including in vitro maturation (IVM), in vitro fertilization
 Name: Xueming Zhao Professional Title: Professor Telephone:86-010-62815892 Fax:86-010-62895971 E-mail: zhaoxueming@caas.cn Website: http://www.iascaas.net.cn/yjspy/dsjj/sssds/dwyzyzypz1/62040.htm Research
Name: Xueming Zhao Professional Title: Professor Telephone:86-010-62815892 Fax:86-010-62895971 E-mail: zhaoxueming@caas.cn Website: http://www.iascaas.net.cn/yjspy/dsjj/sssds/dwyzyzypz1/62040.htm Research
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice
 An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice Mei-Yi Wu 1 *, Ming Jiang 1, Xiaodong Zhai 2, Arthur L. Beaudet 2, Ray-Chang Wu 1 * 1 Department of
An Unexpected Function of the Prader-Willi Syndrome Imprinting Center in Maternal Imprinting in Mice Mei-Yi Wu 1 *, Ming Jiang 1, Xiaodong Zhai 2, Arthur L. Beaudet 2, Ray-Chang Wu 1 * 1 Department of
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Dissecting gene regulation network in human early embryos. at single-cell and single-base resolution
 Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
Dissecting gene regulation network in human early embryos at single-cell and single-base resolution Fuchou Tang BIOPIC, College of Life Sciences Peking University 07/10/2015 Cockburn and Rossant, 2010
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Paternal exposure and effects on microrna and mrna expression in developing embryo. Department of Chemical and Radiation Nur Duale
 Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation
 Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation Robert FEIL Director of Research CNRS & University of Montpellier, Montpellier, France. E-mail:
Epigenetic Regulation of Health and Disease Nutritional and environmental effects on epigenetic regulation Robert FEIL Director of Research CNRS & University of Montpellier, Montpellier, France. E-mail:
Current Research Strategies and Therapeutic Approaches in Duchenne Muscular Dystrophy
 Current Research Strategies and Therapeutic Approaches in Duchenne Muscular Dystrophy H. Lee Sweeney, Ph.D. Department of Physiology University of Pennsylvania Perelman School of Medicine Current Research
Current Research Strategies and Therapeutic Approaches in Duchenne Muscular Dystrophy H. Lee Sweeney, Ph.D. Department of Physiology University of Pennsylvania Perelman School of Medicine Current Research
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Epigenetics 101. Kevin Sweet, MS, CGC Division of Human Genetics
 Epigenetics 101 Kevin Sweet, MS, CGC Division of Human Genetics Learning Objectives 1. Evaluate the genetic code and the role epigenetic modification plays in common complex disease 2. Evaluate the effects
Epigenetics 101 Kevin Sweet, MS, CGC Division of Human Genetics Learning Objectives 1. Evaluate the genetic code and the role epigenetic modification plays in common complex disease 2. Evaluate the effects
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy
 Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
EFFECTS OF STRESS ACROSS GENERATIONS: WHY SEX MATTERS
 Commentary submitted to Biological Psychiatry EFFECTS OF STRESS ACROSS GENERATIONS: WHY SEX MATTERS Invited commentary on: Saavedra-Rodriguez L, Feig LA (2012): Chronic Social Instability Induces Anxiety
Commentary submitted to Biological Psychiatry EFFECTS OF STRESS ACROSS GENERATIONS: WHY SEX MATTERS Invited commentary on: Saavedra-Rodriguez L, Feig LA (2012): Chronic Social Instability Induces Anxiety
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Strategic delivery: Setting standards Increasing and. Details: Output: Demonstrating efficiency. informing choice.
 Strategic delivery: Setting standards Increasing and informing choice Demonstrating efficiency economy and value Details: Meeting Scientific and Clinical Advances Advisory Committee Agenda item 6 Paper
Strategic delivery: Setting standards Increasing and informing choice Demonstrating efficiency economy and value Details: Meeting Scientific and Clinical Advances Advisory Committee Agenda item 6 Paper
Understanding genetics, mutation and other details. Stanley F. Nelson, MD 6/29/18
 Understanding genetics, mutation and other details Stanley F. Nelson, MD 6/29/18 1 6 11 16 21 Duchenne muscular dystrophy 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 600 500 400 300 200 100 0 Duchenne/Becker
Understanding genetics, mutation and other details Stanley F. Nelson, MD 6/29/18 1 6 11 16 21 Duchenne muscular dystrophy 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 600 500 400 300 200 100 0 Duchenne/Becker
Mutations. A2 Biology For WJEC
 12. Mutation is a change in the amount, arrangement or structure in the DNA of an organism. 13. There are two types of mutations, chromosome mutations and gene mutations. Mutations A2 Biology For WJEC
12. Mutation is a change in the amount, arrangement or structure in the DNA of an organism. 13. There are two types of mutations, chromosome mutations and gene mutations. Mutations A2 Biology For WJEC
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Mutation specific therapies
 Taken from www.dmd.nl/gt. Used with permission Mutation specific therapies Introduction Two therapies for Duchenne patients are currently being tested in clinical trials, which are applicable only to patients
Taken from www.dmd.nl/gt. Used with permission Mutation specific therapies Introduction Two therapies for Duchenne patients are currently being tested in clinical trials, which are applicable only to patients
The Meaning of Genetic Variation
 Activity 2 The Meaning of Genetic Variation Focus: Students investigate variation in the beta globin gene by identifying base changes that do and do not alter function, and by using several CD-ROM-based
Activity 2 The Meaning of Genetic Variation Focus: Students investigate variation in the beta globin gene by identifying base changes that do and do not alter function, and by using several CD-ROM-based
Transgenerational Effects of Diet: Implications for Cancer Prevention Overview and Conclusions
 Transgenerational Effects of Diet: Implications for Cancer Prevention Overview and Conclusions John Milner, Ph.D. Director, USDA Beltsville Human Nutrition Research Center Beltsville, MD 20705 john.milner@ars.usda.gov
Transgenerational Effects of Diet: Implications for Cancer Prevention Overview and Conclusions John Milner, Ph.D. Director, USDA Beltsville Human Nutrition Research Center Beltsville, MD 20705 john.milner@ars.usda.gov
Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing
 Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing PacBio Americas User Group Meeting Sample Prep Workshop June.27.2017 Tyson Clark, Ph.D. For Research Use Only. Not
Iso-Seq Method Updates and Target Enrichment Without Amplification for SMRT Sequencing PacBio Americas User Group Meeting Sample Prep Workshop June.27.2017 Tyson Clark, Ph.D. For Research Use Only. Not
Human inherited diseases
 Human inherited diseases A genetic disorder that is caused by abnormality in an individual's DNA. Abnormalities can range from small mutation in a single gene to the addition or subtraction of a whole
Human inherited diseases A genetic disorder that is caused by abnormality in an individual's DNA. Abnormalities can range from small mutation in a single gene to the addition or subtraction of a whole
Insulin Resistance. Biol 405 Molecular Medicine
 Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
TRANSLATION: 3 Stages to translation, can you guess what they are?
 TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
Clinical Genomics. Ina E. Amarillo, PhD FACMGG
 Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Single Gene (Monogenic) Disorders. Mendelian Inheritance: Definitions. Mendelian Inheritance: Definitions
 Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Supplemental Data. Integrating omics and alternative splicing i reveals insights i into grape response to high temperature
 Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
SUPPLEMENTARY INFORMATION
 Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Epigenetic processes are fundamental to development because they permit a
 Early Life Nutrition and Epigenetic Markers Mark Hanson, PhD Epigenetic processes are fundamental to development because they permit a range of phenotypes to be formed from a genotype. Across many phyla
Early Life Nutrition and Epigenetic Markers Mark Hanson, PhD Epigenetic processes are fundamental to development because they permit a range of phenotypes to be formed from a genotype. Across many phyla
he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003)
 MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
DMD Genetics: complicated, complex and critical to understand
 DMD Genetics: complicated, complex and critical to understand Stanley Nelson, MD Professor of Human Genetics, Pathology and Laboratory Medicine, and Psychiatry Co Director, Center for Duchenne Muscular
DMD Genetics: complicated, complex and critical to understand Stanley Nelson, MD Professor of Human Genetics, Pathology and Laboratory Medicine, and Psychiatry Co Director, Center for Duchenne Muscular
The Chromosomal Basis Of Inheritance
 The Chromosomal Basis Of Inheritance Chapter 15 Objectives Explain the chromosomal theory of inheritance and its discovery. Explain why sex-linked diseases are more common in human males than females.
The Chromosomal Basis Of Inheritance Chapter 15 Objectives Explain the chromosomal theory of inheritance and its discovery. Explain why sex-linked diseases are more common in human males than females.
Gene Regulation Part 2
 Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Basic Definitions. Dr. Mohammed Hussein Assi MBChB MSc DCH (UK) MRCPCH
 Basic Definitions Chromosomes There are two types of chromosomes: autosomes (1-22) and sex chromosomes (X & Y). Humans are composed of two groups of cells: Gametes. Ova and sperm cells, which are haploid,
Basic Definitions Chromosomes There are two types of chromosomes: autosomes (1-22) and sex chromosomes (X & Y). Humans are composed of two groups of cells: Gametes. Ova and sperm cells, which are haploid,
Human Genetics 542 Winter 2018 Syllabus
 Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
Human Genetics 542 Winter 2018 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Jan 3 rd Wed Mapping disease genes I: inheritance patterns and linkage analysis
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Epigenetics: Basic Principals and role in health and disease
 Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
Epigenetics: Basic Principals and role in health and disease Cambridge Masterclass Workshop on Epigenetics in GI Health and Disease 3 rd September 2013 Matt Zilbauer Overview Basic principals of Epigenetics
The Chromosomal Basis of Inheritance
 Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways
 Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways Suppressor mutation: a second mutation results in a less severe phenotype than the original mutation Suppressor mutations
Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways Suppressor mutation: a second mutation results in a less severe phenotype than the original mutation Suppressor mutations
Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice
 Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
Supplementary Information Genome-editing via Oviductal Nucleic Acids Delivery (GONAD) system: a novel microinjection-independent genome engineering method in mice Gou Takahashi, Channabasavaiah B Gurumurthy,
Human Genetics 542 Winter 2017 Syllabus
 Human Genetics 542 Winter 2017 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Module I: Mapping and characterizing simple genetic diseases Jan 4 th Wed Mapping
Human Genetics 542 Winter 2017 Syllabus Monday, Wednesday, and Friday 9 10 a.m. 5915 Buhl Course Director: Tony Antonellis Module I: Mapping and characterizing simple genetic diseases Jan 4 th Wed Mapping
Chapter 15 Notes 15.1: Mendelian inheritance chromosome theory of inheritance wild type 15.2: Sex-linked genes
 Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Islets of Langerhans consist mostly of insulin-producing β cells. They will appear to be densely labeled
 Are adult pancreatic beta cells formed by self-duplication or stem cell differentiation? Introduction Researchers have long been interested in how tissues produce and maintain the correct number of cells
Are adult pancreatic beta cells formed by self-duplication or stem cell differentiation? Introduction Researchers have long been interested in how tissues produce and maintain the correct number of cells
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Reanalyzing variable directionality of gene expression in transgenerational epigenetic
 Reanalyzing variable directionality of gene expression in transgenerational epigenetic inheritance Abhay Sharma CSIR-Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research
Reanalyzing variable directionality of gene expression in transgenerational epigenetic inheritance Abhay Sharma CSIR-Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research
Ch. 15 The Chromosomal Basis of Inheritance
 Ch. 15 The Chromosomal Basis of Inheritance Nov 12 12:58 PM 1 Essential Question: Are chromosomes the basis of inheritance? Nov 12 1:00 PM 2 1902 Walter S. Sutton, Theodor Boveri, et al Chromosome Theory
Ch. 15 The Chromosomal Basis of Inheritance Nov 12 12:58 PM 1 Essential Question: Are chromosomes the basis of inheritance? Nov 12 1:00 PM 2 1902 Walter S. Sutton, Theodor Boveri, et al Chromosome Theory
Figure S1, Beyer et al.
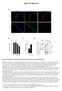 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Emerging Treatment Strategies for FSHD
 Department of Pharmacology Emerging Treatment Strategies for FSHD Peter L. Jones, Ph.D. and Takako I. Jones, Ph.D. Co-Principal Investigators Department of Pharmacology Disclosures: Peter Jones and Takako
Department of Pharmacology Emerging Treatment Strategies for FSHD Peter L. Jones, Ph.D. and Takako I. Jones, Ph.D. Co-Principal Investigators Department of Pharmacology Disclosures: Peter Jones and Takako
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Page 1. Name:
 Name: 4734-1 - Page 1 Warts result when certain viruses cause skin cells to reproduce at a high rate. This rapid reproduction of skin cells is due to the viruses stimulating cellular digestion mitotic
Name: 4734-1 - Page 1 Warts result when certain viruses cause skin cells to reproduce at a high rate. This rapid reproduction of skin cells is due to the viruses stimulating cellular digestion mitotic
Untitled Document. 1. Which of the following is the template for the production of RNA within a cell? A. DNA B. ATP C. protein D.
 Name: Date: 1. Which of the following is the template for the production of RNA within a cell? A. DNA B. ATP C. protein D. carbohydrate 2. Which sequence of DNA bases would pair with the ones shown in
Name: Date: 1. Which of the following is the template for the production of RNA within a cell? A. DNA B. ATP C. protein D. carbohydrate 2. Which sequence of DNA bases would pair with the ones shown in
Proteins. Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids
 Proteins Protein carbon, hydrogen, oxygen, nitrogen and often sulphur Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids During protein synthesis, amino acids
Proteins Protein carbon, hydrogen, oxygen, nitrogen and often sulphur Length of protein varies from thousands of amino acids to only a few insulin only 51 amino acids During protein synthesis, amino acids
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
MicroRNA-mediated incoherent feedforward motifs are robust
 The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.
 Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology
 Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Session 2: Biomarkers of epigenetic changes and their applicability to genetic toxicology Bhaskar Gollapudi, Ph.D The Dow Chemical Company Workshop: Genetic Toxicology: Opportunities to Integrate New Approaches
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
The Chromosomal Basis of Inheritance
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 15 The Chromosomal Basis of Inheritance
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 15 The Chromosomal Basis of Inheritance
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
The subcortical maternal complex controls symmetric division of mouse zygotes by
 The subcortical maternal complex controls symmetric division of mouse zygotes by regulating F-actin dynamics Xing-Jiang Yu 1,2, Zhaohong Yi 1, Zheng Gao 1,2, Dan-dan Qin 1,2, Yanhua Zhai 1, Xue Chen 1,
The subcortical maternal complex controls symmetric division of mouse zygotes by regulating F-actin dynamics Xing-Jiang Yu 1,2, Zhaohong Yi 1, Zheng Gao 1,2, Dan-dan Qin 1,2, Yanhua Zhai 1, Xue Chen 1,
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
