LUMC: LOPAC screen analysis
|
|
|
- Gordon Ferguson
- 5 years ago
- Views:
Transcription
1 LUMC: LOPAC screen analysis Preliminary results November 9, 2012 This document briefly presents the results from the LOPAC compounds screen performed at LUMC. It contains the following information: The set of all protein targets that were targeted by at least one compound - AllProteinTargets uniprot.csv (Section 1) List of compounds and the proteins that they target - Lopac2Target uniprot.csv and Lopac2Target bin uniprot.csv (Section 1) Pruned list of protein targets. For both Mtb and Stm - LopacFiltered joint.csv. For Mtb - LopacFiltered Mtb.csv. For Stm - LopacFiltered Stm.csv (Section 2) Gene to gene interactions (provided in a separate file) - g2g.txt. Gene annotations with terms from Gene Ontology (provided in a separate file) - g2ont.txt. Results from predictive clustering trees. Results from feature ranking. Results from predictive clustering trees including the info about cell viability. 1 Protein targets We describe each compound from the LOPAC library with the protein targets for which was showed the given compound is active. From the 1260 compounds in the library, 967 compounds were found to be active on human protein targets. In the further analysis, we include only those. Furthermore, we identified 760 (human) protein targets for which at least one LOPAC compound was found active. The proteins are given by their Uniprot ID. The list of protein targets is given in a separate file (AllProteinTargets uniprot.csv). The association of each LOPAC compound to the targets is given first in the file Lopac2Target.csv, where each row represents a compound. Each compound is identifeid by its LOPAC catnum (the first field in the row), and then the protein targets (i.e., the respective accession number ids) are listed. Furthermore, we provide a matrix with dimensions in file Lopac2Target bin.csv. Here, each row presents a compound and each column is a protein target. Note that, the first row contains the protein accession numbers, while the first column contains the LOPAC catnum identifeirs. 1
2 2 Prunning of the protein targets The number of 760 target proteins is large and it includes protein that are targeted by a large variety of compounds. Typically, these proteins are not specific and closely relevant for the Mtb and Stm measurements. Furthermore, more interesting are the protein targets that were found active in studies involving hit compounds (compounds with z-score larger than 2 or smaller than -2) and not active in the remainder of the compounds. To this end, we prune the set of the proteins using the following rule: A protein target is removed from the set of protein targets if at least 90% of the compounds that are targeted by are not identified as hits. Considering this, we provide three separate lists of compounds and the respective protein targets. The first list (given in the file LopacFiltered joint.csv) gives the protein targets that are obtained when the hit compounds were defined using both Mtb and Stm z-score values. The second list (given in the file LopacFiltered Mtb.csv) gives the protein targets that are relevent for the hit compounds for the Mtb study. The third list (given in the file LopacFiltered Stm.csv) gives the protein targets relevant for the hit compounds for the Stm study. We further briefly show the gene networks obtained from the pruned protein targets using STRING. First, we give the network for both Mtb and Stm (the 44 proteins given in the file LopacFiltered joint.csv). Then, we show the network for Mtb (the 41 proteins from file LopacFiltered Mtb.csv). Finally, we present the network for Stm (the 11 proteins from file LopacFiltered Stm.csv) 3 Interactions and annotations We describe each of the targets with the Gene Ontology (GO) and KEGG terms that are annotated with. Afterwards, for each compound, we take the union of the GO terms from each protein target that the given protein was described with. The file that contains the GO and KEGG annotations is g2ont.txt. Next, we include the interactions of the annotated proteins. Namely, using the file g2g.txt, each compound is described with the union of the GO and KEGG terms for each of the proteins and additionally, the union of the GO and KEGG terms that the interacting proteins are annotated with. 4 Resuls from Feature Ranking Here, we present the results obtained by ReliefF feature ranking algorithm (from WEKA toolbox). Instead of given the complete list, we present the proteins networks for Mtb and Stm for both numeric and discrete values of the z-score. 2
3 Figure 1: The protein network for both Mtb and Stm. 3
4 Figure 2: The protein network for Mtb. Figure 3: The protein network for Stm. 4
5 Figure 4: The protein network for the top 50 proteins for Mtb numeric. 5
6 Figure 5: The protein network for the top 50 proteins for Mtb discrete. 6
7 Figure 6: The protein network for the top 50 proteins for Stm numeric. 7
8 Figure 7: The protein network for the top 50 proteins for Stm discrete. 8
9 5 Results from PCTs with cell viability info This section includes the results from the analysis of the data using predictive clustering trees (PCTs). We present the results in three sub-sections, each dealing with the specific representation of the protein targets and therefore the compounds. In other words, the three scenarios differ in the descriptive attributes, while the target attributes are always the same: the z-scores for each compound. For each of the scenarios we present the result for the real values of the z-scores and the discretized counterparts (positive and negative hit ). The results shown here, besides the bacterial load, also have the z-score for cell viability as target variables. The first value in the tree leafs is for bacterial load and the second value is for cell viability. In this scenario, we are more focused on two outcomes of the application of the compounds: kill bacteria - do nothing to host and promote bacteria - do nothing to host. We are however not interested in the outcomes where nothing happens to the bacteria or both bacteria and the host are killed. Interesting outcomes are also: kill bacteria - promote host and promote bacteria - promote host. However, in this analysis we do not consider them since they do not appear in any of the PCTs, probably because there is no specific GO term or protein target that offers a good differentiation between these conditions and the rest of the conditions. In the respective section, we give a short overview of each PCT in the context of the three outcomes of compounds application. 5.1 Uniprot protein identifiers Mtb - numeric A2TJX0 +--yes: [-0.12, ]: 4 compound: [C9911,C9754,P4405,O3125] +--no: F1D8R8 +--yes: [-0.866, ]: no: Q9BYT3 +--yes: [-1.47, ]: no: P yes: [0.435,-7.01]: 2 compound: [C6645,M6383] +--no: P yes: [-5.305,-1.23]: 2 compound: [S8442,S9692] +--no: [ , ]: 912 compound: [...] A2TJX0: Krueppel-like factor 5 F1D8R8: Steroidogenic factor 1 nuclear receptor Q9BYT3: Serine/threonine-protein kinase P05067: Amyloid beta A4 protein P07949: Proto-oncogene tyrosine-protein kinase recept In this tree, most interesting is the group of compounds that consists of S8442 and S9692, which when applied kill the bacteria and do nothing to the host. Namely, this happens when the protein P07949 is targeted. 9
10 5.1.2 Mtb - discrete F1D8R8 +--yes: [0,-1] [9.0,10.0]: no: A2TJX0 +--yes: [1,-1] [2.0,3.0]: 3 compound: [C9911,C9754,O3125] +--no: Q9BYT3 +--yes: [0,0] [24.0,23.0]: no: P yes: [-1,-1] [1.0,2.0]: 2 compound: [C6645,M6383] +--no: P yes: [-1,-1] [2.0,1.0]: 2 compound: [S8442,S9692] +--no: P yes: [0,0] [36.0,35.0]: no: Q96DE8 +--yes: [-1,0] [2.0,2.0]: 3 compound: [T4318,B8433,C3930] +--no: [0,0] [797.0,839.0]: 865 compound: [...] F1D8R8: Steroidogenic factor 1 nuclear receptor A2TJX0: Krueppel-like factor 5 Q9BYT3: Serine/threonine-protein kinase 33 P05067: Amyloid beta A4 protein P07949: Proto-oncogene tyrosine-protein kinase receptor P43116: Prostaglandin E2 receptor EP2 subtype Q96DE8: PSMD14 protein In this tree, a group of compounds kill the bacteria and do nothing to the host. Thia group consists of the following compounds T4318, B8433 and C3930 and it targets the Q96DE8 protein. 10
11 5.1.3 Stm - numeric Q yes: [10.795,-6.075]: 2 compound: [C9754,M1404] +--no: Q yes: [6.625,-4.105]: 4 compound: [I3766,V1377,P4405,V8879] +--no: O yes: [3.71,-5.18]: 2 compound: [C9911,E1383] +--no: A0PJU7 +--yes: [2.754,-2.488]: 5 compound: [N3510,E1779,N1786,T182,C7522] +--no: P yes: [2.53,-1.916]: 5 compound: [C6645,R5010,T1132,O3125,M6383] +--no: Q yes: [2.46,-2.45]: 2 compound: [A2385,A9809] +--no: [ , ]: 947 compound: [...] Q92793: CREB-binding protein Q04206: Transcription factor p65 O14757: Serine/threonine-protein kinase Chk1 A0PJU7: TSHR protein P05067: Amyloid beta A4 protein Q15788: Nuclear receptor coactivator 1 This tree presents several outcomes where the bacterial load is increased and cell viability is decreased Stm - discrete A2TJX0 +--yes: [1,-1] [3.0,3.0]: 4 compound: [C9911,C9754,P4405,O3125] +--no: F1D8R8 +--yes: [0,0] [10.0,10.0]: no: P yes: O yes: [1,-1] [2.0,2.0]: 2 compound: [C6645,M6383] +--no: [0,0] [2.0,2.0]: 2 compound: [R5010,T1132] +--no: A0PJU7 +--yes: [0,-1] [2.0,2.0]: 3 compound: [N3510,E1779,C7522] +--no: P
12 +--yes: [0,0] [1.0,1.0]: 2 compound: [G0668,C8221] +--no: P yes: [0,-1] [1.0,1.0]: 2 compound: [P108,C5134] +--no: Q9BYT3 +--yes: [0,0] [25.0,24.0]: no: O yes: [-1,0] [1.0,2.0]: 2 compound: [H89,S5567] +--no: P yes: [-1,0] [1.0,2.0]: 2 compound: [P8386,O8757] +--no: Q yes: [0,0] [1.0,2.0]: 2 compound: [N2255,D9766] +--no: F1D8R9 +--yes: [0,0] [2.0,1.0]: 2 compound: [O2139,D8399] +--no: P yes: [0,0] [2.0,1.0]: 2 compound: [A8723,I8250] +--no: Q yes: [0,-1] [2.0,1.0]: 2 compound: [L2167,G6793] +--no: [0,0] [884.0,882.0]: 897 compound: [...] A2TJX0: Krueppel-like factor 5 F1D8R8: Steroidogenic factor 1 nuclear receptor P05067: Amyloid beta A4 protein O60674: Tyrosine-protein kinase JAK2 A0PJU7: TSHR protein P04626: Receptor tyrosine-protein kinase erbb-2 P47901: Vasopressin V1b receptor Q9BYT3: Serine/threonine-protein kinase 33 O75582: Ribosomal protein S6 kinase alpha-5 P02768: Serum albumin Q99808: Equilibrative nucleoside transporter 1 F1D8R9: Liver nuclear receptor homolog-1 variant 2 P21462: fmet-leu-phe receptor Q03181: Peroxisome proliferator-activated receptor In this tree, the most interesting outcome is produced when the protein is targeted by the H89 or S5567. Similarly, when P02768 is targeted the bacterial load is decreased and no harm to the cell viability: P8386 and O8757 target this protein. Furthermore, this tree reveals a connection between the proteins P05067 and O Namely, when both of the proteins are targeted then the bacteria are promoted while the cell viability is reduced. However, if only P05067 is targeted and O60674 is not targets then nothing happens to the bacteria and the host. 12
13 5.2 GO and KEGG terms Mtb - numeric GO yes: [-0.12, ]: 4 compound: [C9911,C9754,P4405,O3125] +--no: GO yes: GO yes: [-4.02, ]: 3 compound: [V1377,V8879,N1786] +--no: [ , ]: no: GO yes: [ , ]: no: GO yes: [-4.33, ]: 3 compound: [A0966,Q3251,S9692] +--no: GO yes: GO yes: GO yes: [-0.995,-0.775]: no: [2.33,-3.4]: 2 compound: [B168,A8598] +--no: GO yes: GO yes: GO yes: [ ,0.75]: 3 compound: [L2167,G6793,F8175] +--no: [ , ]: no: [ , ]: no: GO yes: [-1.235,-3.01]: 2 compound: [A5006,N5751] +--no: [ , ]: no: [1.2525, ]: 4 compound: [T7692,C3270,A6770,C0330] GO : microvillus assembly GO : luteinization GO : immune response-activating cell surface receptor signaling pathway GO : protein autophosphorylation GO : oxygen transporter activity GO : molecular function GO : positive regulation of protein ubiquitination GO : membrane part GO : DNA repair 13
14 GO : response to organic substance GO : response to cold GO : regulation of leukocyte mediated cytotoxicity This tree elucidates two interesting outcomes where the bacterial load is reduced while nothing happens to the host. The first outcome occurs when the oxygen transporter activity is targeted (GO ) by some of the compounds S9692, A0966 and Q3251. The second outcome occurs when DNA repair (GO ), the response to organic substance (GO ) and the response to cold (GO ) are targeted simultaneously by some of the compounds L2167, G6793 and F Mtb - discrete GO yes: GO yes: [0,0] [7.0,7.0]: 7 compound: [A2385,R5010,C7632,C9511,T9652,D7910,P9375] +--no: [-1,-1] [7.0,11.0]: no: GO yes: [1,-1] [2.0,3.0]: 3 compound: [C9911,C9754,O3125] +--no: GO yes: [0,0] [71.0,70.0]: no: GO yes: [-1,-1] [1.0,2.0]: 2 compound: [C6645,M6383] +--no: GO yes: [-1,0] [4.0,5.0]: 6 compound: [M6760,L8789,Q0125,E7881,H89,M5250] +--no: GO yes: [-1,0] [2.0,2.0]: 3 compound: [M2537,H1512,K1003] +--no: GO yes: GO yes: [-1,0] [3.0,3.0]: 3 compound: [A0966,Q3251,S9692] +--no: [0,0] [2.0,2.0]: 2 compound: [A4638,F6627] +--no: GO yes: [0,0] [91.0,93.0]: no: GO yes: [0,0] [20.0,21.0]: no: GO yes: [-1,0] [2.0,3.0]: 3 compound: [L2167,L3791,S3065] +--no: [0,0] [668.0,698.0]:
15 compound: [...] GO : melanocyte differentiation GO : endosomal transport GO : microvillus assembly GO : regulation of protein ubiquitination GO : serine-type endopeptidase inhibitor activity GO : protein kinase B signaling cascade GO : cholesterol biosynthetic process GO : oxygen transporter activity GO : two-component sensor activity GO : negative regulation of DNA replication GO : response to prostaglandin stimulus GO : ensheathment of neurons This tree illustrates four interesting outcomes where the bacterial load is reduced, while nothing happens to the host. The first outcome occurs when the protein kinase B signaling cascade (GO ) is targeted by some of the following compounds: M6760, L8789, Q0125, E7881, H89 and M5250. The second outcome occurs when the cholesterol biosynthetic process (GO ) is targeted by some of the following compounds: M2537,H1512 and K1003. The third outcome occurs when both oxygen transporter activity (GO ) and two-component sensor activity (GO ) are targeted simultaneously by some of the following compounds A0966,Q3251 and S9692. The fourth outcome occurs when the ensheathment of neurons (GO ) is targeted by some of the following compounds L2167, L3791 and S Stm - numeric GO yes: [10.795,-6.075]: 2 compound: [C9754,M1404] +--no: GO yes: [6.625,-4.105]: 4 compound: [I3766,V1377,P4405,V8879] +--no: GO yes: [3.71,-5.18]: 2 compound: [C9911,E1383] +--no: GO yes: [ , ]: no: GO yes: [2.46,-2.45]: 2 compound: [A2385,A9809] +--no: GO yes: [-3.185,0.215]: 2 compound: [H1512,F1293] +--no: [ , ]: 929 compound: [...] GO : RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in negative regulation of transcription 15
16 GO : phosphate ion binding GO : regulation of double-strand break repair via homologous recombination GO : hormone-mediated signaling pathway GO : regulation of transcription by galactose GO : steroid delta-isomerase activity This tree points-out to an outcome where the bacterial load is reduced, while the host cell remains unharmed. This occurs when the steroid delta-isomerase activity (GO ) is targeted by some of the following compounds: H1512 and F Stm - discrete GO yes: GO yes: [1,-1] [4.0,4.0]: 4 compound: [C9911,V1377,P4405,V8879] +--no: GO yes: GO yes: [0,0] [2.0,2.0]: 2 compound: [A2385,R5010] +--no: [1,-1] [4.0,4.0]: 4 compound: [C9754,M1404,A9809,E1383] +--no: GO yes: [0,-1] [1.0,2.0]: 2 compound: [N3510,C7522] +--no: GO yes: [0,0] [26.0,26.0]: no: [0,-1] [2.0,2.0]: 3 compound: [B168,A8598,T7402] +--no: GO yes: GO yes: [0,0] [2.0,2.0]: 2 compound: [T1132,O3125] +--no: [1,-1] [2.0,2.0]: 2 compound: [C6645,M6383] +--no: GO yes: [0,0] [19.0,22.0]: no: GO yes: [0,-1] [1.0,1.0]: 2 compound: [P108,C5134] +--no: GO yes: [-1,0] [1.0,2.0]: 2 compound: [P8386,O8757] +--no: GO yes: [0,0] [1.0,2.0]: 2 compound: [N2255,D9766] +--no: GO yes: [0,-1] [2.0,1.0]: 2 16
17 compound: [H2380,S7690] +--no: [0,0] [878.0,872.0]: 889 compound: [...] GO : regulation of histone H3-K9 acetylation GO : regulation of chemokine biosynthetic process GO : protein acetylation GO : urea cycle GO : aggressive behavior GO : membrane part GO : serine-type endopeptidase inhibitor activity GO : MAPK cascade GO : positive regulation of translation GO : hyperosmotic salinity response GO : cytolysis by symbiont of host cells GO : uridine transport GO : protein phosphatase inhibitor activity This tree points-out to an outcome where the bacterial load is reduced, while the host cell remains unharmed. This occurs when the cytolysis by symbiont of host cells (GO ) is targeted by some of the following compounds: P8386 and O GO and KEGG terms plus interaction information Mtb - numeric GO yes: [-0.12, ]: 4 compound: [C9911,C9754,P4405,O3125] +--no: GO yes: [-0.866, ]: no: IAGO yes: [ , ]: 3 compound: [G6416,S9692,M1404] +--no: GO yes: [-2.585,-1.238]: 10 compound: [M6760,R5010,Q0125,N3510,F9397,P0453,P8139,T5575,H89,S3567] +--no: GO yes: [0.24,-4.9]: 3 compound: [C6645,T1132,M6383] +--no: GO yes: [ , ]: no: IAGO yes: [ , ]: no: [ , ]: 777 compound: [...] 17
18 GO : microvillus assembly GO : luteinization IAGO : regulation of interferon-beta biosynthetic process GO : regulation of glucose transport GO : serine-type endopeptidase inhibitor activity GO : cellular macromolecular complex subunit organization IAGO : cleavage furrow This tree presents a scenario where the bacterial load is decreased while the host is unharmed. This scenario occurs when the regulation of glucose transport (GO ) is targeted by M6760, R5010, Q0125, N3510, F9397, P0453, P8139, T5575, H89 and S Mtb - discrete IAGO yes: GO yes: [-1,-1] [5.0,7.0]: 7 compound: [C6645,V1377,P4405,P8139,V8879,C3930,M6383] +--no: [0,0] [32.0,31.0]: no: GO yes: [-1,0] [5.0,5.0]: 8 compound: [C9911,M6760,L8789,Q0125,E7881,W1628,H89,M5250] +--no: GO yes: [0,0] [23.0,20.0]: no: GO yes: [0,-1] [2.0,2.0]: 3 compound: [P0453,T9033,T182] +--no: GO yes: [0,0] [26.0,34.0]: no: GO yes: [0,-1] [3.0,2.0]: 4 compound: [Z0878,M2537,I1149,D8690] +--no: GO yes: GO yes: [-1,0] [3.0,3.0]: 3 compound: [A0966,Q3251,S9692] +--no: [0,0] [2.0,2.0]: 2 compound: [A4638,F6627] +--no: [0,0] [776.0,808.0]: 832 compound: [...] IAGO : carbohydrate derivative biosynthetic process GO : response to interferon-gamma GO : protein kinase B signaling cascade GO : bone resorption GO : melanocyte differentiation GO : DNA double-strand break processing 18
19 GO : positive regulation of striated muscle tissue development GO : oxygen transporter activity GO : two-component sensor activity This tree presents a scenario where the bacterial load is decreased while nothing happens to the host. There are two possibilities for this scenario to occur. To begin with, the protein kinase B signaling cascade (GO ) needs to be targeted by C9911, M6760, L8789, Q0125, E7881, W1628, H89 or M5250. Next, the two-component sensor activity (GO ) needs to be targeted by A0966, Q3251 or S Stm - numeric IAGO yes: [6.954,-3.874]: 5 compound: [I3766,V1377,N3510,P4405,V8879] +--no: GO yes: [10.795,-6.075]: 2 compound: [C9754,M1404] +--no: IAGO yes: [3.085,-3.815]: 4 compound: [C9911,A2385,A9809,E1383] +--no: GO yes: [2.53,-1.916]: 5 compound: [C6645,R5010,T1132,O3125,M6383] +--no: IAGO yes: GO yes: [2.07,-3.15]: 3 compound: [N1786,T182,C7522] +--no: GO yes: [-3.8,0.2]: 2 compound: [H1512,D9815] +--no: [ , ]: no: [ , ]: 899 compound: [...] IAGO : JUN kinase activity GO : RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in negative regulation of transcription IAGO : branching involved in prostate gland morphogenesis GO : serine-type endopeptidase inhibitor activity IAGO : superoxide metabolic process GO : orexin receptor activity GO : negative regulation of the force of heart contraction involved in baroreceptor response to increased systemic arterial blood pressure This tree presents an outcome where the bacterial load is decreased and nothing happens to the host. This outcome occurs when the negative regulation of the force of heart contraction involved in baroreceptor response to increased systemic arterial blood pressure (GO ) is targeted by H1512 or D9815, while the orexin receptor activity (GO ) is not targeted. 19
20 5.3.4 Stm - discrete IAGO yes: [0,-1] [8.0,7.0]: no: GO yes: [1,-1] [2.0,2.0]: 2 compound: [C9754,M1404] +--no: GO yes: [1,-1] [2.0,2.0]: 3 compound: [C9911,N3510,S5567] +--no: IAGO yes: [-1,0] [1.0,2.0]: 3 compound: [G0668,C8221,H89] +--no: GO yes: [0,-1] [1.0,1.0]: 2 compound: [P108,C5134] +--no: GO yes: [0,0] [29.0,28.0]: no: GO yes: [-1,0] [1.0,2.0]: 2 compound: [P8386,O8757] +--no: GO yes: [0,0] [1.0,2.0]: 2 compound: [N2255,D9766] +--no: GO yes: [0,0] [117.0,122.0]: no: GO yes: [0,-1] [2.0,1.0]: 2 compound: [C0993,S7690] +--no: [0,0] [774.0,765.0]: 777 compound: [...] IAGO : adrenal gland development GO : RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in negative regulation of transcription GO : positive regulation of Notch signaling pathway IAGO : negative regulation of muscle cell apoptotic process GO : hyperosmotic salinity response GO : regulation of fatty acid beta-oxidation GO : cytolysis by symbiont of host cells GO : uridine transport GO : double-strand break repair GO : protein phosphatase type 2A regulator activity This tree presents two scenarios for decrease of bacterial load while nothing happens to the cell. The first scenario occurs when we target a protein that is in interaction with a protein involved in negative regulation of muscle cell apoptotic process (IAGO ). This occurs 20
21 when some of the following compounds are applied: G0668,C8221 and H89. The second scenario occurs when the cytolysis by symbiont of host cells (GO ) is targeted by P8386 or O
NBCE Mock Board Questions Biochemistry
 1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1
 GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1 1. The endocrine system consists of glands that secrete chemical signals, called hormones, into the blood. In addition, other organs and cells
GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1 1. The endocrine system consists of glands that secrete chemical signals, called hormones, into the blood. In addition, other organs and cells
Point total. Page # Exam Total (out of 90) The number next to each intermediate represents the total # of C-C and C-H bonds in that molecule.
 This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler
 Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
Supplementary information for: Community detection for networks with. unipartite and bipartite structure. Chang Chang 1, 2, Chao Tang 2
 Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Identification of Tissue-Specific Protein-Coding and Noncoding. Transcripts across 14 Human Tissues Using RNA-seq
 Identification of Tissue-Specific Protein-Coding and Noncoding Transcripts across 14 Human Tissues Using RNA-seq Jinhang Zhu 1, Geng Chen 1, Sibo Zhu 1,2, Suqing Li 3, Zhuo Wen 3, Bin Li 1, Yuanting Zheng
Identification of Tissue-Specific Protein-Coding and Noncoding Transcripts across 14 Human Tissues Using RNA-seq Jinhang Zhu 1, Geng Chen 1, Sibo Zhu 1,2, Suqing Li 3, Zhuo Wen 3, Bin Li 1, Yuanting Zheng
levels of genes were separated by their expression levels; 2,000 high, medium, and low
 Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
LIX1 APOD RRM2 FAIM2 MGST1 UHRF1 PPIA BEX1
 C2orf80 1 C2orf80 A2M ANXA1 APOD ARL4A BEX1 C1S C3 CD44 CHI3L1 CHI3L2 CLU DPYD DSCAM EMP3 FN1 GBP1 GEM GLIPR1 GRIA2 IDH1 LIX1 MAP2 MDK MGST1 MIR3917 NAMPT PCMTD2 PPIA PTN PTPRZ1 S100B SCG3 SERPINA3 TCF12
C2orf80 1 C2orf80 A2M ANXA1 APOD ARL4A BEX1 C1S C3 CD44 CHI3L1 CHI3L2 CLU DPYD DSCAM EMP3 FN1 GBP1 GEM GLIPR1 GRIA2 IDH1 LIX1 MAP2 MDK MGST1 MIR3917 NAMPT PCMTD2 PPIA PTN PTPRZ1 S100B SCG3 SERPINA3 TCF12
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
MITOCW watch?v=wyt7efjlbak
 MITOCW watch?v=wyt7efjlbak The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
MITOCW watch?v=wyt7efjlbak The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
Inflammatory Bowel Disease - Target Initial Survey
 Date: / / This document has been created as input for the upcoming target evaluation meeting with where this initial candidate list will be discussed. After the evaluation meeting Euretos will undertake
Date: / / This document has been created as input for the upcoming target evaluation meeting with where this initial candidate list will be discussed. After the evaluation meeting Euretos will undertake
Activity: Biologically Important Molecules
 Activity: Biologically Important Molecules AP Biology Introduction We have already seen in our study of biochemistry that the molecules that comprise living things are carbon-based, and that they are thought
Activity: Biologically Important Molecules AP Biology Introduction We have already seen in our study of biochemistry that the molecules that comprise living things are carbon-based, and that they are thought
2.2 Cell Construction
 2.2 Cell Construction Elemental composition of typical bacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
2.2 Cell Construction Elemental composition of typical bacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Metabolomic Data Analysis with MetaboAnalystR
 Metabolomic Data Analysis with MetaboAnalystR Name: guest623839008349489450 February 7, 2018 1 Background Understanding the functional importance of metabolites in untargeted metabolomics is limited due
Metabolomic Data Analysis with MetaboAnalystR Name: guest623839008349489450 February 7, 2018 1 Background Understanding the functional importance of metabolites in untargeted metabolomics is limited due
HORMONES (Biomedical Importance)
 hormones HORMONES (Biomedical Importance) Hormones are the chemical messengers of the body. They are defined as organic substances secreted into blood stream to control the metabolic and biological activities.
hormones HORMONES (Biomedical Importance) Hormones are the chemical messengers of the body. They are defined as organic substances secreted into blood stream to control the metabolic and biological activities.
Cell Communication. Cell Communication. Communication between cells requires: ligand: the signaling molecule
 Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12864 Supplementary Table 1 1 2 3 4 5 6 7 Peak Gene code Screen Function or Read analysis AMP reads camp annotation reads minor Tb927.2.1810 AMP ISWI Confirmed
SUPPLEMENTARY INFORMATION doi:10.1038/nature12864 Supplementary Table 1 1 2 3 4 5 6 7 Peak Gene code Screen Function or Read analysis AMP reads camp annotation reads minor Tb927.2.1810 AMP ISWI Confirmed
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Endocrine System Hormones. AP Biology
 Endocrine System Hormones 2007-2008 Regulation Why are hormones needed? u chemical messages from one body part to another u communication needed to coordinate whole body u daily homeostasis & regulation
Endocrine System Hormones 2007-2008 Regulation Why are hormones needed? u chemical messages from one body part to another u communication needed to coordinate whole body u daily homeostasis & regulation
Org Biochem Final Test Student Section Ch Samples Page 1 of 5
 Ch 31-35 Samples Page 1 of 5 13. Which of the following is a purine? a) guanine b) cytosine c) thymine d) uracil 20. The three components of a nucleotide are a) glucose, a phosphate group, and choline.
Ch 31-35 Samples Page 1 of 5 13. Which of the following is a purine? a) guanine b) cytosine c) thymine d) uracil 20. The three components of a nucleotide are a) glucose, a phosphate group, and choline.
Supplementary Figure 1.
 Supplementary Figure 1. Increased expression of cell cycle pathway genes in insulin + Glut2 low cells of STZ-induced diabetic islets. A) random blood glucose measuers of STZ and vehicle treated MIP-GFP
Supplementary Figure 1. Increased expression of cell cycle pathway genes in insulin + Glut2 low cells of STZ-induced diabetic islets. A) random blood glucose measuers of STZ and vehicle treated MIP-GFP
Sub-Topic Classification of HIV related Opportunistic Infections. Miguel Anderson and Joseph Fonseca
 Sub-Topic Classification of HIV related Opportunistic Infections Miguel Anderson and Joseph Fonseca Introduction Image collected from the CDC https://www.cdc.gov/hiv/basics/statistics.html Background Info
Sub-Topic Classification of HIV related Opportunistic Infections Miguel Anderson and Joseph Fonseca Introduction Image collected from the CDC https://www.cdc.gov/hiv/basics/statistics.html Background Info
Cancer. Questions about cancer. What is cancer? What causes unregulated cell growth? What regulates cell growth? What causes DNA damage?
 Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
3.D- Cell Communication
 3.D- Cell Communication Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. EU 3.A: Heritable information provides for continuity of life. EU 3.B:
3.D- Cell Communication Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. EU 3.A: Heritable information provides for continuity of life. EU 3.B:
Supplementary Files. Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis
 Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
Supplementary Files Novel blood-based microrna biomarker panel for early diagnosis of chronic pancreatitis Lei Xin 1 *, M.D., Jun Gao 1 *, Ph.D., Dan Wang 1 *, M.D., Jin-Huan Lin 1, M.D., Zhuan Liao 1,
HORMONES AND CELL SIGNALLING
 HORMONES AND CELL SIGNALLING TYPES OF CELL JUNCTIONS CHEMICAL SIGNALS AND MODES OF ACTION Endocrine system produces chemical messages = hormones that are transported from endocrine gland to target cell
HORMONES AND CELL SIGNALLING TYPES OF CELL JUNCTIONS CHEMICAL SIGNALS AND MODES OF ACTION Endocrine system produces chemical messages = hormones that are transported from endocrine gland to target cell
Endocrine System Hormones (Ch. 45)
 Endocrine System Hormones (Ch. 45) Regulation Why are hormones needed? chemical messages from one body part to another communication needed to coordinate whole body daily homeostasis & regulation of large
Endocrine System Hormones (Ch. 45) Regulation Why are hormones needed? chemical messages from one body part to another communication needed to coordinate whole body daily homeostasis & regulation of large
Office number.
 The University of Jordan Faculty: Pharmacy Department: Biopharmaceutics and Clinical Pharmacy Program: Pharmacy Academic Year/ Fall Semester: 2014/15 BIOCHEMISTRY 2 [1203253] Credit hours 3 Level 2 nd
The University of Jordan Faculty: Pharmacy Department: Biopharmaceutics and Clinical Pharmacy Program: Pharmacy Academic Year/ Fall Semester: 2014/15 BIOCHEMISTRY 2 [1203253] Credit hours 3 Level 2 nd
Metabolism of pentoses, glycogen, fructose and galactose. Jana Novotna
 Metabolism of pentoses, glycogen, fructose and galactose Jana Novotna 1. The Pentose Phosphate Pathway The pentose phosphate pathway (PPP): (hexose monophosphate or 6-phosphogluconate patway) Process that
Metabolism of pentoses, glycogen, fructose and galactose Jana Novotna 1. The Pentose Phosphate Pathway The pentose phosphate pathway (PPP): (hexose monophosphate or 6-phosphogluconate patway) Process that
Formalin-fixed paraffin-embedded sections of liver from a recipient mouse sacrificed after two rounds
 Supplementary figure legends Supplementary Figure 1 Fah + hepatocytes in a Fah -/- mouse transplanted with sorted cells. Formalin-fixed paraffin-embedded sections of liver from a recipient mouse sacrificed
Supplementary figure legends Supplementary Figure 1 Fah + hepatocytes in a Fah -/- mouse transplanted with sorted cells. Formalin-fixed paraffin-embedded sections of liver from a recipient mouse sacrificed
Name: Date: Block: Biology 12
 Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Validated Mouse Quantitative RT-PCR Genes Gene Gene Bank Accession Number Name
 5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
TNFSF13B tumor necrosis factor (ligand) superfamily, member 13b NF-kB pathway cluster, Enrichment Score: 3.57
 Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
COMMON ASSESSMENT
 1. The diagram above is a model of a cellular process called transcription. What class of biological molecules is represented in the diagram? A. Carbohydrates B. Nucleic acids C. Proteins D. Lipids B.9.A.R
1. The diagram above is a model of a cellular process called transcription. What class of biological molecules is represented in the diagram? A. Carbohydrates B. Nucleic acids C. Proteins D. Lipids B.9.A.R
Chapter 11: Cell Communication
 Name Period Chapter 11: Cell Communication The special challenge in Chapter 11 is not that the material is so difficult, but that most of the material will be completely new to you. Cell communication
Name Period Chapter 11: Cell Communication The special challenge in Chapter 11 is not that the material is so difficult, but that most of the material will be completely new to you. Cell communication
Evidence for an Alternative Glycolytic Pathway in Rapidly Proliferating Cells. Matthew G. Vander Heiden, et al. Science 2010
 Evidence for an Alternative Glycolytic Pathway in Rapidly Proliferating Cells Matthew G. Vander Heiden, et al. Science 2010 Introduction The Warburg Effect Cancer cells metabolize glucose differently Primarily
Evidence for an Alternative Glycolytic Pathway in Rapidly Proliferating Cells Matthew G. Vander Heiden, et al. Science 2010 Introduction The Warburg Effect Cancer cells metabolize glucose differently Primarily
Chapter 11 Guided Reading: Cell Communication
 Name Chapter 11 Guided Reading: Cell Communication The special challenge in Chapter 11 is not that the material is so difficult, but that most of the material will be completely new to you. Cell communication
Name Chapter 11 Guided Reading: Cell Communication The special challenge in Chapter 11 is not that the material is so difficult, but that most of the material will be completely new to you. Cell communication
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Molecular building blocks
 2.22 Cell Construction Elemental l composition of ftypical lbacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
2.22 Cell Construction Elemental l composition of ftypical lbacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 3 Review Assignment
 Class: Date: Chapter 3 Review Assignment Multiple Choice 40 MC = 40 Marks Identify the choice that best completes the statement or answers the question. 1. Which of the following organelles produces transport
Class: Date: Chapter 3 Review Assignment Multiple Choice 40 MC = 40 Marks Identify the choice that best completes the statement or answers the question. 1. Which of the following organelles produces transport
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Biochemistry of Cancer and Tumor Markers
 Biochemistry of Cancer and Tumor Markers The term cancer applies to a group of diseases in which cells grow abnormally and form a malignant tumor. It is a long term multistage genetic process. The first
Biochemistry of Cancer and Tumor Markers The term cancer applies to a group of diseases in which cells grow abnormally and form a malignant tumor. It is a long term multistage genetic process. The first
Mechanistic Toxicology
 SECOND EDITION Mechanistic Toxicology The Molecular Basis of How Chemicals Disrupt Biological Targets URS A. BOELSTERLI CRC Press Tavlor & France Croup CRC Press is an imp^t o* :H Taylor H Francn C'r,,jpi
SECOND EDITION Mechanistic Toxicology The Molecular Basis of How Chemicals Disrupt Biological Targets URS A. BOELSTERLI CRC Press Tavlor & France Croup CRC Press is an imp^t o* :H Taylor H Francn C'r,,jpi
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine
 Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
About This Chapter. Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution
 About This Chapter Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution Hormones: Function Control Rates of enzymatic reactions
About This Chapter Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution Hormones: Function Control Rates of enzymatic reactions
Carbon Compounds. Lesson Overview. Lesson Overview. 2.3 Carbon Compounds
 Lesson Overview Carbon Compounds Lesson Overview 2.3 THINK ABOUT IT In the early 1800s, many chemists called the compounds created by organisms organic, believing they were fundamentally different from
Lesson Overview Carbon Compounds Lesson Overview 2.3 THINK ABOUT IT In the early 1800s, many chemists called the compounds created by organisms organic, believing they were fundamentally different from
2.3 Carbon-Based Molecules. KEY CONCEPT Carbon-based molecules are the foundation of life.
 KEY CONCEPT Carbon-based molecules are the foundation of life. Carbon atoms have unique bonding properties. Carbon forms covalent bonds with up to four other atoms, including other carbon atoms. Carbon-based
KEY CONCEPT Carbon-based molecules are the foundation of life. Carbon atoms have unique bonding properties. Carbon forms covalent bonds with up to four other atoms, including other carbon atoms. Carbon-based
Hand in the Test Sheets (with the checked multiple choice answers) and your Sheets with written answers.
 Page 1 of 13 IMPORTANT INFORMATION Hand in the Test Sheets (with the checked multiple choice answers) and your Sheets with written answers. THE exam has an 'A' and 'B' section SECTION A (based on Dykyy's
Page 1 of 13 IMPORTANT INFORMATION Hand in the Test Sheets (with the checked multiple choice answers) and your Sheets with written answers. THE exam has an 'A' and 'B' section SECTION A (based on Dykyy's
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis
 Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Chromosomes Chromosomes were first observed by the German embryologist Walther Fleming in 1882. Chromosome number varies among organisms most
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Chromosomes Chromosomes were first observed by the German embryologist Walther Fleming in 1882. Chromosome number varies among organisms most
An Introduction to Carbohydrates
 An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney
 BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
Hormones. Prof. Dr. Volker Haucke Institut für Chemie-Biochemie Takustrasse 6
 Hormones Prof. Dr. Volker Haucke Institut für Chemie-Biochemie Takustrasse 6 Tel. 030-8385-6920 (Sekret.) 030-8385-6922 (direkt) e-mail: vhaucke@chemie.fu-berlin.de http://userpage.chemie.fu-berlin.de/biochemie/aghaucke/teaching.html
Hormones Prof. Dr. Volker Haucke Institut für Chemie-Biochemie Takustrasse 6 Tel. 030-8385-6920 (Sekret.) 030-8385-6922 (direkt) e-mail: vhaucke@chemie.fu-berlin.de http://userpage.chemie.fu-berlin.de/biochemie/aghaucke/teaching.html
Reproductive Endocrinology
 Reproductive Endocrinology Reproductive Endocrinology Hypothalamic hormones Gonadotropin releasing hormone (GnRH) - stimulate release of FSH = follicle stimulating hormone LH = luteinizing hormone from
Reproductive Endocrinology Reproductive Endocrinology Hypothalamic hormones Gonadotropin releasing hormone (GnRH) - stimulate release of FSH = follicle stimulating hormone LH = luteinizing hormone from
Unit 9: The Cell Cycle
 Unit 9: The Cell Cycle Name: Period: Test Date: 1 Table of Contents Title of Page Page Number Teacher Stamp Unit 9 Warm-Ups 3-4 Cell Cycle/Interphase Notes 5-6 DNA Replication Notes 7-8 DNA replication
Unit 9: The Cell Cycle Name: Period: Test Date: 1 Table of Contents Title of Page Page Number Teacher Stamp Unit 9 Warm-Ups 3-4 Cell Cycle/Interphase Notes 5-6 DNA Replication Notes 7-8 DNA replication
Table S1 Differentially expressed genes showing > 2 fold changes and p <0.01 for 0.01 mm NS-398, 0.1mM ibuprofen and COX-2 RNAi.
 Table S1 Differentially expressed genes showing > 2 fold changes and p
Table S1 Differentially expressed genes showing > 2 fold changes and p
18. PANCREATIC FUNCTION AND METABOLISM. Pancreatic secretions ISLETS OF LANGERHANS. Insulin
 18. PANCREATIC FUNCTION AND METABOLISM ISLETS OF LANGERHANS Some pancreatic functions have already been discussed in the digestion section. In this one, the emphasis will be placed on the endocrine function
18. PANCREATIC FUNCTION AND METABOLISM ISLETS OF LANGERHANS Some pancreatic functions have already been discussed in the digestion section. In this one, the emphasis will be placed on the endocrine function
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Nonspecific Defenses of the Host. Chapter 16
 Nonspecific Defenses of the Host Chapter 16 I. Introduction: Overview of host defenses A. Resistance Ability to ward off disease through body defenses 1. Nonspecific All body defenses that protect one
Nonspecific Defenses of the Host Chapter 16 I. Introduction: Overview of host defenses A. Resistance Ability to ward off disease through body defenses 1. Nonspecific All body defenses that protect one
Supplementary figure legends
 Supplementary figure legends Fig. S1. Lineweaver-Burk plot of putrescine uptake by YeeF. An overnight culture of SK629 was inoculated in 100-mL LBG medium in 500-mL Erlenmeyer flasks. The medium was supplemented
Supplementary figure legends Fig. S1. Lineweaver-Burk plot of putrescine uptake by YeeF. An overnight culture of SK629 was inoculated in 100-mL LBG medium in 500-mL Erlenmeyer flasks. The medium was supplemented
Zool 3200: Cell Biology Exam 4 Part I 2/3/15
 Name: Key Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name: Key Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Campbell Biology in Focus (Urry) Chapter 9 The Cell Cycle. 9.1 Multiple-Choice Questions
 Campbell Biology in Focus (Urry) Chapter 9 The Cell Cycle 9.1 Multiple-Choice Questions 1) Starting with a fertilized egg (zygote), a series of five cell divisions would produce an early embryo with how
Campbell Biology in Focus (Urry) Chapter 9 The Cell Cycle 9.1 Multiple-Choice Questions 1) Starting with a fertilized egg (zygote), a series of five cell divisions would produce an early embryo with how
Comparative analysis revealed dosage sensitivity and regulatory patterns of lncrna in prostate cancer
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Comparative analysis revealed dosage sensitivity and regulatory patterns of lncrna
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Comparative analysis revealed dosage sensitivity and regulatory patterns of lncrna
* Kyoto Encyclopedia of Genes and Genomes.
 Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
BCM 226 LECTURE SALEMCITY, A.J
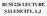 BCM 226 LECTURE SALEMCITY, A.J BIOLOGICAL MEMBRANE Biological membranes are composed of proteins associated with a lipid bilayer matrix. They are the molecular gateway to the cell. Viewed under electron
BCM 226 LECTURE SALEMCITY, A.J BIOLOGICAL MEMBRANE Biological membranes are composed of proteins associated with a lipid bilayer matrix. They are the molecular gateway to the cell. Viewed under electron
Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz )
 Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz ) Biochemistry Submitted by Marie Havlová on 8. February 2012-0:00 Syllabus of Biochemistry Mechanisms of enzyme catalysis.
Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz ) Biochemistry Submitted by Marie Havlová on 8. February 2012-0:00 Syllabus of Biochemistry Mechanisms of enzyme catalysis.
Cell Cell
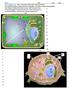 Go to cellsalive.com. Select Interactive Cell Models: Plant and Animal. Fill in the information on Plant and Animal Organelles, then Click on Start the Animation Select Plant or Animal Cell below the box.
Go to cellsalive.com. Select Interactive Cell Models: Plant and Animal. Fill in the information on Plant and Animal Organelles, then Click on Start the Animation Select Plant or Animal Cell below the box.
Cell are made up of organelles. An ORGANELLE is a specialized subunit within a cell that has a specific function.
 Plant and Animal Cells The Cell Theory All living things are made up of one or more cells. All cells come from other cells. Organization of Living Things Cell are made up of organelles. An ORGANELLE is
Plant and Animal Cells The Cell Theory All living things are made up of one or more cells. All cells come from other cells. Organization of Living Things Cell are made up of organelles. An ORGANELLE is
Homeostasis. Salt sucks! Review: What is distilled water? What would a salty solution be considered?
 You will get your practice tests back to correct. Take a few minutes to look over your answers and you can use resources like notes and book to help you correct the test. Salt sucks! Review: What is distilled
You will get your practice tests back to correct. Take a few minutes to look over your answers and you can use resources like notes and book to help you correct the test. Salt sucks! Review: What is distilled
Lesson Overview. Carbon Compounds. Lesson Overview. 2.3 Carbon Compounds
 Lesson Overview 2.3 The Chemistry of Carbon What elements does carbon bond with to make up life s molecules? Carbon can bond with many elements, including Hydrogen, Oxygen, Phosphorus, Sulfur, and Nitrogen
Lesson Overview 2.3 The Chemistry of Carbon What elements does carbon bond with to make up life s molecules? Carbon can bond with many elements, including Hydrogen, Oxygen, Phosphorus, Sulfur, and Nitrogen
Bachelor Nutrition Science Seminar Nutritional Biochemistry (Module BE2.3) Topics
 Nutritional Biochemistry (Module BE2.3) Prof. Dr. Stefan Lorkowski 1 Bachelor Nutrition Science Seminar Nutritional Biochemistry (Module BE2.3) Topics Biosynthesis of amino acids 1. Amino acids: general
Nutritional Biochemistry (Module BE2.3) Prof. Dr. Stefan Lorkowski 1 Bachelor Nutrition Science Seminar Nutritional Biochemistry (Module BE2.3) Topics Biosynthesis of amino acids 1. Amino acids: general
The Structure and Function of Biomolecules
 The Structure and Function of Biomolecules The student is expected to: 9A compare the structures and functions of different types of biomolecules, including carbohydrates, lipids, proteins, and nucleic
The Structure and Function of Biomolecules The student is expected to: 9A compare the structures and functions of different types of biomolecules, including carbohydrates, lipids, proteins, and nucleic
Sul Ross State University Syllabus for Biochemistry II: CHEM 4302 (Fall 2017) (Alpine and Midland)
 Sul Ross State University Syllabus for Biochemistry II: CHEM 4302 (Fall 2017) (Alpine and Midland) Class: Biochemistry II Instructor: Dr. David Leaver Room: WSB 321 (Alpine) Office: WSB 318 Time: MWF 11:00-11:50am
Sul Ross State University Syllabus for Biochemistry II: CHEM 4302 (Fall 2017) (Alpine and Midland) Class: Biochemistry II Instructor: Dr. David Leaver Room: WSB 321 (Alpine) Office: WSB 318 Time: MWF 11:00-11:50am
Most life processes are a series of chemical reactions influenced by environmental and genetic factors.
 Biochemistry II Most life processes are a series of chemical reactions influenced by environmental and genetic factors. Metabolism the sum of all biochemical processes 2 Metabolic Processes Anabolism-
Biochemistry II Most life processes are a series of chemical reactions influenced by environmental and genetic factors. Metabolism the sum of all biochemical processes 2 Metabolic Processes Anabolism-
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Ayman Mesleh & Leen Alnemrawi. Bayan Abusheikha. Faisal
 24 Ayman Mesleh & Leen Alnemrawi Bayan Abusheikha Faisal We were talking last time about receptors for lipid soluble hormones.the general mechanism of receptors for lipid soluble hormones: 1. Receptors
24 Ayman Mesleh & Leen Alnemrawi Bayan Abusheikha Faisal We were talking last time about receptors for lipid soluble hormones.the general mechanism of receptors for lipid soluble hormones: 1. Receptors
Supplemental Information:
 SupplementalInformation: Content: FigureS1.AccessibilityoftheL.interrogansproteomebyLC MSanalysis. FigureS2.Functionalannotationoftheidentifiedproteins. FigureS3.DistributionofMS1 featureintensities. FigureS4.ComparisonoftheDDA
SupplementalInformation: Content: FigureS1.AccessibilityoftheL.interrogansproteomebyLC MSanalysis. FigureS2.Functionalannotationoftheidentifiedproteins. FigureS3.DistributionofMS1 featureintensities. FigureS4.ComparisonoftheDDA
Cell Communication. Local and Long Distance Signaling
 Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
Hormones, Receptors and Receptor-Hormone Interactions
 Classification of Hormones Hormones, Receptors and Receptor-Hormone Interactions Synthesis of Protein Hormones and Amine Hormones Hormone Activity Locations of Receptors Mechanisms of Hormone Action Types
Classification of Hormones Hormones, Receptors and Receptor-Hormone Interactions Synthesis of Protein Hormones and Amine Hormones Hormone Activity Locations of Receptors Mechanisms of Hormone Action Types
BIOH111. o Cell Module o Tissue Module o Skeletal system o Muscle system o Nervous system o Endocrine system o Integumentary system
 BIOH111 o Cell Module o Tissue Module o Skeletal system o Muscle system o Nervous system o Endocrine system o Integumentary system Endeavour College of Natural Health endeavour.edu.au 1 Textbook and required/recommended
BIOH111 o Cell Module o Tissue Module o Skeletal system o Muscle system o Nervous system o Endocrine system o Integumentary system Endeavour College of Natural Health endeavour.edu.au 1 Textbook and required/recommended
The Star of The Show (Ch. 3)
 The Star of The Show (Ch. 3) Why study Carbon? All of life is built on carbon Cells ~72% 2 O ~25% carbon compounds carbohydrates lipids proteins nucleic acids ~3% salts Na, Cl, K Chemistry of Life Organic
The Star of The Show (Ch. 3) Why study Carbon? All of life is built on carbon Cells ~72% 2 O ~25% carbon compounds carbohydrates lipids proteins nucleic acids ~3% salts Na, Cl, K Chemistry of Life Organic
Introduction to Metabolism Cell Structure and Function
 Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
The Structure and Function of Large Biological Molecules
 NAME DATE Chapter 5 - The Structure and Function of Large Biological Molecules Guided Reading Concept 5.1: Macromolecules are polymers, built from monomers 1. The large molecules of all living things fall
NAME DATE Chapter 5 - The Structure and Function of Large Biological Molecules Guided Reading Concept 5.1: Macromolecules are polymers, built from monomers 1. The large molecules of all living things fall
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Chp 2 (cont.) Organic Molecules. Spider s web and close up of capture strand - spider silk protein
 Chp 2 (cont.) Organic Molecules Spider s web and close up of capture strand - spider silk protein 1! Molecular Diversity is Based on Carbon An organic molecule contains both carbon and hydrogen. Ex: Methane
Chp 2 (cont.) Organic Molecules Spider s web and close up of capture strand - spider silk protein 1! Molecular Diversity is Based on Carbon An organic molecule contains both carbon and hydrogen. Ex: Methane
