Deciphering molecular circuits from genetic variation underlying transcriptional responsiveness to stimuli
|
|
|
- Benjamin Holland
- 5 years ago
- Views:
Transcription
1 Deciphering molecular circuits from genetic variation underlying transcriptional responsiveness to stimuli Irit Gat-Viks,2,, Nicolas Chevrier,3,4, Roni Wilentzik 2, Thomas Eisenhaure,5, Raktima Raychowdhury, Yael Steuerman 2, Alex K. Shalek 6,7, Nir Hacohen,8, Ido Amit,9, and Aviv Regev,, Broad Institute of MIT and Harvard, 7 Cambridge Center, Cambridge, MA 242, USA. 2 Department of Cell Research and Immunology, George S. Wise Faculty of Life Sciences, Tel Aviv University, Tel Aviv 69978, Israel. 3 Graduate Program in Immunology, Division of Medical Sciences, Harvard Medical School, Boston, MA 25, USA. 4 FAS Center for Systems Biology, Harvard University, Cambridge, MA 238, USA. 5 Center for Immunology and Inflammatory Diseases, Massachusetts General Hospital, Charlestown, MA 229, USA. 6 Departments of Chemistry and Chemical Biology and 7 Department of Physics, Harvard University, Cambridge, MA 238, USA. 8 Center for Immunology and Inflammatory Diseases, Massachusetts General Hospital, Charlestown, MA 229, USA, and Department of Medicine, Harvard Medical School, Boston, MA 25, USA. 9 Department of Immunology, Weizmann Institute of Science, Rehovot, 76, Israel. Howard Hughes Medical Institute, Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 24, USA. To whom correspondence should be addressed: iritgv@post.tau.ac.il (IGV), ido.amit@weizmann.ac.il (IA), aregev@broad.mit.edu (AR). Supplementary Figures Supplementary Figures -3 (p.2-8) Supplementary Note Supplementary Notes,2,3 (p.9-26) Supplementary Tables Supplementary Tables -6 (p.27-42)
2 c fraction Fraction a.7 All genes Selected genes # genes in an InSignature class ** p= -4 b 2 Average Heritability (H ) Cis-associated genes All signature genes Biological relevnace Responsiveness Other I-b II I-c III Supp. Figure I-a I-d IV Biological relevnace Responsiveness Other Selection criteria InSignature d Median LR score I-b II I-c III InSignature p=.2 p=.9 * * I-b II I-c III I-a I-d IV * p=.4 p=.7 * I-a I-d IV Selection criteria Selection criteria InSignature Biological relevnace Responsiveness Other e InSignature cis InSignature non-cis Sum ncounter, cis ncounter, non-cis Sum = 424 Supplementary Figure. The variation signature genes. (a) Distribution of all and signature genes in InSignature classes. Shown is the fraction of classes (y axis) that include a certain number of genes (x axis) based on the number of all genes included in a class (blue) or the number of variation signature genes included in a class (red). (b-d) Dissection of results for the 424 signature genes based on the criterion by which they were selected (x axis, detailed in Methods and Supplementary Note 2): I - heritability, by InSignature; II biological relevance; III- positive responsiveness controls; IV- negative controls. Within the heritability (criterion I) group, genes are further partitioned by top InSignature score in class (I-a); biological relevance (I-b); responsiveness (I-c) and randomly selected from class (I-d). (b) Heritability in responsiveness to LPS (y axis). As expected, negative controls (group IV) have a significantly lower heritability compared to genes selected by InSignature (group I) (P-value < -9 ), and top scoring genes (group I-a) outperform those randomly selected from the class (group I-d) (P<.3). (c) cis-associations. For each group (x axis), the red-boundary bars show the fraction of cis-associated genes 2
3 Supp. Figure (cont.) in the class (in at least one of stimulus) out of all associated genes (y axis). The black-boundary bars show the fraction of genes in this class among all 424 signature genes. cis-associated genes are enriched in the top-scoring group (I-a; P-value < -4, hypergeometric test) and are depleted from negative control genes (group IV) (P < -3 ). (d) LR scores. Shown are the median LR scores for all traits in each group, except those that attain their best LR score in cis (up to Mbp from the gene). Selection criteria that are based on InSignature are marked with a + sign at the bottom. Comparing matching pairs of groups, the results highlight the utility of InSignature. For biologically relevant genes, selection based on InSignature (group I-b) outperforms selection based on biological relevance only (group II) (P-value <.2); for responding genes, selection based on InSignature (group I-c) is slightly better than based on responsiveness only (group III) (P-value <.9). (e) Sensitivity and specificity of InSignature in selecting cis-associated genes. Shown is the number of genes that are predicted as cis-associated (left column) and the remaining genes (non cis-associated, middle column) by InSignature (from array data), separated by whether they were subsequently found in the ncounter dataset as cis-associated (top row) or not (middle row) using a relatively permissive cutoff (LR score > ). Thus, columns are based on analysis of the microarray dataset (6 BXD strains) whereas rows are based on the ncounter dataset (44 BXD strains). Hence, there is a high specificity (9%), moderate sensitivity (45%) and a moderate false negative rate (54%). 3
4 a B6 D2 BXD2 BXD5 BXD28 BXD4 BXD4 BXD6 B6 D2 BXD2 BXD5 BXD28 BXD4 BXD4 BXD6 Supp. Figure 2 b Fraction.5.5 Microarray ncounter correlation LPS-activated DCs Reshuffled data set ncounter Microarray ncounter correlation c Inter-strain fold change B6 D2 BXD2 BXD5 BXD28 BXD4 BXD4 BXD6 Cxcl Lta Microarray Microarray ncounter d LPS at 6 h; (BXD89 experimental repeat 2) Lta Mertk Ccl7 Cxcl Ccl3 Irf7 Ccl4 Tnf Ila Acpp Trim2 92H23Rik Lhx2 Nmi Slco3a Egr2 r = LPS at 6 h (BXD89 experimental repeat ) Supplementary Figure 2. Consistency and reproducibility of ncounter assay. (a) Shown are the expression levels for a pilot set of 6 genes with significant scores for inter-individual variation (>5.5, Methods, rows) measured in the same samples with both technologies across individual mice (columns; two individuals per strain consecutively). Intensity corresponds to inter-strain fold changes relative to the average level in B6 (Red (blue) - higher (lower) than B6). The bar chart (left) shows the correlation between the microarray and ncounter profiles for each gene. The four genes with lower correlations (92H23Rik, Ila, Tnf and Ccl4) show higher heritability based on the ncounter measurements than with microarrays (data not shown), suggesting that the ncounter assay is more sensitive, consistent with previous studies (Amit et al. 29). (b) Shown is the distribution of ncounter-microarray correlation of gene profiles from (a) (red) compared to the distribution of correlations when reshuffling the measurements of each gene separately (black). (c) Comparison of inter-strain fold changes (y-axis) across the mice individuals (x-axis) between microarray (red) and nanostring ncounter (blue) measurements. Top: Cxcl, bottom: Lta. Between-strain variability is significantly higher than within-strain variability (d) A scatter plot of Nanostring ncounter measured responsiveness levels in two independently repeated experiments with DCs from a single mouse individual from strain BXD89 following LPS stimulation (6 hours). 4
5 a Fraction.8 Responsiveness (R) Reshuffled R b Absolute responsiveness Supp. Figure 3 2 Gene-stimuli interaction ( log p-value) LR score c d e Log2 abundance Log2 abundance Log2 abundance f LR score PAM poly IC LPS Time (hours) Time (hours) Time (hours) Mina Ube2m St3gal5 Inherited B6 Inherited D2 PAM poly IC LPS PAM poly IC LPS Stimulus-specific Stimulus non-specific g Fraction Responsiveness Responsiveness Responsiveness * * * * Mina * * Ube2m * * * * St3gal5 B6 D2 B6 D2 B6 D2 PAM PIC LPS trans-linked R traits cis-linked R traits 5 5 LR score Supplementary Figure 3. Association of responsiveness traits in cis. (a) Significant interactions effect between cis-acting reqtls and stimuli. Shown is the distribution of reqtlstimuli interactions (-log p-value, X axis, Methods) in the dataset (black) and in reshuffled data (gray). (b) High responsiveness is required but not sufficient for high LR scores. Shown is a 5
6 Supp. Figure 3 (cont.) scatter plot of LR scores (x axis) and absolute responsiveness (y axis) across all traits presented in Fig. 3a. Absolute responsiveness values are normalized by the maximum absolute log ratio attained for that gene under any of the stimulations and strains. (c-e) The relation of LR score and responsiveness in the Mina (c), Ube2m (d) and St3Gal5 (e) genes. Left: Shown are the time courses of gene expression levels (X axis) in three stimuli (PAM (left), poly IC (middle), LPS (right)) in strains with a B6 (black) or D2 (grey) genotype in cis. Right: Shown are the responsiveness levels for BXD strains that inherited the genotype in cis from B6 (columns,3,5) and D2 (columns 2,4,6) following stimulation of PAM (columns,2), Poly IC (columns 3,4) and LPS (columns 5,6). For example, the gene Ube2m (d) is highly responsive in LPS and PAM but cis-associated only in LPS. (f) Stimulus-specific cis-reqtls have smaller effects. Shown are the LR scores for stimulus-specific cis-reqtls (left) and stimulus non-specific cis-reqtls (right). In each box, the central mark is the median; edges are the 25th and 75th percentiles; whiskers extend to the most extreme data points not considered outliers; and outliers are plotted individually. (g) Higher LR scores for cis-associations than for trans. Shown is the distribution of LR scores in cis (black) and trans (red). 6
7 Supp. Figure 4 a Association score Association score t2 t A Genomic location t2 t A Genomic location c c2 c3 c c2 c3 A b Association score Association score t3 t4 C D Genomic location t3 C F D Genomic location t4 C D F c Association score t t6 A E A B E Genomic location Supplementary Fig. 4. Examples of merge steps in InVamod. Detailing the steps shown in Fig. 4a. In (a) trait t forms a single-trait seed module (exceeds c in position A), but trait t2 does not. The traits are merged to a legal module of size 2 that remains associated at A, the best association for both traits. The new association score of the module equals its optimum (2+5=7, loss fraction ε = maximum permitted loss of.), and both traits exceed the double-trait cutoff c2 in position A, thus forming a legal module. (b) Traits t3 and t4 each form a single-trait seed modules (each exceed c) at positions C and D, respectively. They are merged into a new legal module, associated at position F, which maximizes the association score when considering both traits. Both traits exceed c2 at position F, and the module s association score is ( ), the optimal score is 2 (6+6) and the loss fraction is ε = (2-)/2 <=.. (c) Each of traits t and t6 form single-trait seed modules (exceed c in positions A, E, respectively). They cannot form a legal module since when merged their loss fraction ε is higher than.. In particular, the maximal joint association score (in B) is (5+5) resulting in a loss fraction ε = (3-)/3 >.. 7
8 a Stimulation specificity Real modules Permutation-based modules Size > Module size b Heritability (H 2 ) # #2 #3 #4 #5 Module identifier Supp. Figure 5 c Explained heritability (%) #6 # #2 #3 #4 #5 #6 Module identifier d # Isg2 Oasa Oasl2 Iigp2 Slfn8 Ifi44 Ddx6 Ifit2 Ifit3 Irf7 Dhx58 Stat2 #2 Iigp2 Slfn8 Ifi44 Ddx6 Ifit2 Ifit3 Irf7 Dhx58 Cox8 Nmi Sp Daxx Iigp Arid5a Irf2 Trim2 Stat2 Ints4 Csf Timeless Rbm43 Tgif Myd88 Tlr2 Smpdl3b Vwf Tnf Clec5a Hdac5 Rusc2 PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS RP LPS poly IC leading RP LPS PAM poly IC leading RP LPS PAM RP #3 Apbb3 Acox Ccl7 Socs2 LPS poly IC Med2 Cdd leading RP Ypel3 Cdd2 Ifna2 BC372 Ifna4 LPS PAM poly IC RP Cxcl9 Hmgn3 Idi Plscr Ifnb Pmvk Il2rb2 Ptger4 Dusp6 Clcn7 Kctd4 Gpr9a RP #4 Tlr7 Trim25 Atad3a Ddx8 Ftsj3 Bysl Irf8 Prmt3 Pdk Ikbke Slc6a4 Exosc5 Egln3 Zmat3 PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS RP PAM RP LPS RP LPS PAM RP LPS poly IC RP LPS PAM poly ICRP RP LPS poly IC leading RP LPS PAM poly IC leading RP PAM poly IC RP LPS PAM RP #5 Nlrp3 Ehd Cxcr4 Gadd45a Spred Dhrs3 Six Eps8 Rasgrp Nfkb2 Tsc22d3 Lfng Tef Socs6 E2f5 Gtf2e2 Fam7b Net #6 Areg Ereg Ripk3 Ptgs Nfe2 Batf Rnh Ugcg Trem Spry C5ar Il5ra PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS PAM Poly IC LPS RP LPS PAM leading RP LPS PAM poly IC leading RP LPS poly IC RP RP Early divergence Late divergence Convergence LPS PAM leading RP LPS PAM poly IC leading RP LPS poly IC RP Supplementary Figure 5. Association of responsiveness traits in trans. (a) Module sizes. Shown is a scatter-plot of module sizes (x-axis) and the degree of stimulus specificity of the traits in the module (y-axis), defined as minus Shannon entropy measure applied on the distribution of module traits in each of the stimuli. The score is low for a non-specific module, (consisting of the same number of traits in each stimulus) and higher when the traits are unevenly distributed across stimuli. Shown are all InVamod modules generated from our dataset (orange) and on permuted data (black), generated only by independently reshuffling each gene s 8
9 Supp. Figure 5 (cont.) measurements. Module with at least 4 traits are highly unlikely to be generated at random (P<.; vertical grey line). The six modules that cross this threshold (modules #-#6) are stimulus-dependent. (b,c) High heritability and high % explained heritability in Modules #-#6. Shown are the average heritability (H 2, b) and average percentage of heritability explained by the module s reqtl (c) for each module. In each box, the central mark is the median; edges are the 25th and 75th percentiles; whiskers extend to the most extreme data points not considered outliers; and outliers are plotted individually. (d) Gene members of each module. Shown are the association profiles (left, orange: associated) of the member genes (rows) and their responsiveness profiles (right, grey: responsive, as in Fig. 3a). Distinct RPs are marked (right). Genes related to more than one module (in different conditions), or associated only under a stimulus in which the reqtl is predicted to be inactive, are excluded for simplicity. The full module membership is reported in Supplementary Table 5. 9
10 Supp. Figure 6 a Input: responsiveness profiles S S2 S3 RP Input: linked traits S S2 S3 Module genes RP2 Module genes Stimulations RP3 Stimulations Input: regulatory circuit model S S2 S3 Leading responsiveness profiles = {RP,RP2} S S2 S3 S reqtl activity profile = {S} S2 S3 Search a branch conditioned on {S} and upstream to {RP,RP2}? S S2 S3 RP RP2 RP3 RP RP2 RP3 RP RP2 RP3 RP RP2 RP3 b Leading responsiveness profiles reqtl activity profile Regulatory circuit model () RP RP2 S S2 S3 S S2 S3 S S2 S Consistent with model RP RP2 RP3 (2) RP S S2 S3 S S2 S3 S 3 2 S2 S Consistent with model RP RP2 RP3 (3) RP S S2 S3 S S2 S3 S 3 2 S2 S Model refinement RP RP2 RP3 (4) RP S S2 S3 S S2 S3 S 3 2 RP S2 S RP2 RP3 Consistent with model Model selection rule applied
11 Supp. Figure 6 (cont.) Supplementary Figure 6. The InCircuit procedure for positioning reqtls in a wiring diagram. (a) InCircuit takes as input the responsiveness profiles of module genes (RPs, top left), the linked traits of the module (top right) (depicted as in Fig. 4c, Supplementary Fig. 5d), and a regulatory wiring diagram (bottom left), consisting of three stimuli (S, S2, S3) and 3 RPs (RP, RP2, RP3). In step, leading responsiveness profiles (RPs) are marked on the diagram (RP and RP2, orange). In step 2, the stimuli under which the reqtl is active are marked (S, orange). In step 3, InCircuit searches a branch coupling the upstream active stimulations and the downstream leading RPs. If such a branch is found, an reqtl is positioned on it (colored orange circle). (b) Examples of reqtl positioning by InCircuit. Shown are combinations of leading responsiveness profiles (column ), reqtl activity profiles (column 2), and the reqtl predicted position (orange circle) on the wiring diagram (column 3). (): reqtl positioned on branch 3, the only branch downstream of S, which is upstream to both RP and RP2. (2) reqtl positioned on branch 2, the only branch downstream of both S and S2 and upstream of only RP. (3) No known branch exists downstream of S3 and upstream to RP; InCircuit refines the model by adding this branch (dashed line). (4) Both branch and 3 (orange) are downstream of S and upstream of RP, as required. InCircuit positions the reqtl on the branch (Branch ) consisting of a smaller number of downstream RPs, and hence higher specificity.
12 Supp. Figure 7 a Uapl St3gal5 Lta Mina Slc3a4 Ppfibp2 Enah Lrrc8c Clcn7 Lpgat Lhx2 Tor3a Trim2 AI4567 Psen2 Blvra PAM Poly IC LPS PAM Poly IC LPS I II III IV b Group I PAM Group III LPS LPS Poly IC Group II PAM LPS Group IV PAM LPS Poly IC Poly IC Early diverging Late diverging Converging Converging (inverse).4 Absolute responsiveness Supplementary Figure 7. Positioning cis-reqtls in a wiring diagram. (a) Responsiveness and association profiles for genes in each of groups I-IV (as in Fig. 3a). (b) Shown are cisreqtls (black circles) associated with genes in each of these groups (I-IV) and their predicted positioning in relevant wiring diagram (LPS, poly IC, PAM: stimuli; Horizonal grey ovals: inflammatory response; vertical grey ovals: antiviral response). In each case, the cis-reqtl is positioned proximally to its cis-linked gene, and is downstream of the stimuli in which the association is observed (e.g. LPS and PAM in Group II). The linked target gene may be connected to additional signal based on its responsiveness profile (e.g. poly IC in Group II). 2
13 Supp. Figure 8 Enrichment -log P-value Irf3/Stat ReLA PAM-LPS PAM-Poly IC-LPS Poly IC-LPS Responsiveness profiles Supplementary Figure 8. The positioning of responsiveness profiles in the wiring diagram is supported by TF binding. Genes in modules #-#6 were grouped into responsiveness profiles based on their responsiveness to LPS-PAM, LPS-Poly IC and PAM-LPS-Poly IC (Supplementary Fig. 5d). For each group of genes that have the same responsiveness profiles (x axis), shown is their hyper-geometric enrichment score ( log P-value) with NFkB targets (white), and Stat or Irf3 targets (black) (y-axis), as determined by ChIP-Seq or genetic perturbation in DCs (Amit et al, 29; Garber et al, 22). As expected, LPS-PAM and PAM- LPS-Poly IC RPs are enriched with NFκB targets, whereas LPS-Poly IC and PAM-LPS-Poly IC are enriched with Irf3/Stat targets. These strongly support their differential positioning in our circuit. 3
14 Supp. Figure 9 a 25 2 Slfn8 Irf7 Daxx Dhx58 Stat2 Ligp Ifit3 Ifit2 LR score Genomic position shrgs6 (2) effect on transcript level b #2 Neg. Ctrl ** ** ** #2 Neg. Ctrl ** ** ** * #2 Neg. Ctrl ** * #2 Neg. Ctrl Baseline PAM SeV LPS Supplementary Figure 9. Module #2. (a) LR scores for expression traits of eight target genes of module #2. Shown are the LR scores (Y axis) at each genomic position (X axis) for the expression traits of eight target genes (color code, side legend) measured at 6 hours following poly IC stimulation. (b) Rgs6 effect on gene transcript levels in module #2. Shown is a distribution of the effect of knockdown of Rgs6 on transcripts levels. Shown as in Fig. 6c, except that the results are presented for another shrgs6 (here, shrgs6(2)) whose knockdown efficiency is lower (Fig. 6b). *P<. ; **P<.. 4
15 Supp. Figure a Responsiveness (R) Abundance data Fraction Association score ( log p value) b.5.4 Responsiveness (R) Abundance data Fraction Gene x stimuli interaction ( log p value) Supplementary Figure. Responsiveness versus transcript abundance: comparison of genetic associations and gene-environment interaction. Shown is a distribution of the association scores (-log p-value; a) and QTL-stimulus interactions (-log p-value; b), calculated based on a full ANOVA model (Methods), based on either expression traits (eqtls, abundance, red) or responsiveness traits (reqtls, black). Each plot presents data for all variation signature genes that were measured on the Nanostring ncounter. Compared to abundance traits, association of responsiveness tends to be lower (a, KS test P< -5 ), but QTL-stimuli interactions tend to be higher (b, KS test P< -4 ), suggesting the importance of gene-environment interactions in the context of responsiveness traits 5
16 . Predict the InSignature pattern of a gene Supp. Figure Gene expression 2 3 B6 D2 BXD5 BXD4 B6 D2 BXD4 B6 D2 Best mixture of Guassians x x B6 D2 BXD5 BXD4 B6 D2 BXD4 B6 D2 2 3 Predicted InSignature pattern (Gene i) x x 2. Partition genes into insignature classes InSignature Class InSignature Class 2 InSignature Class k InSignature pattern InSignature score Genes ` Gene i... Genes 3. Select representative genes within each class Representative Representative... of Class of Class 2 Representative of Class k Supplementary Figure. The InSignature method. InSignature takes as input genome-wide profiles from different strains and conditions. In step (top) it predicts, for each gene, an underlying InSignature pattern based on a maximum likelihood ratio fitting of two (or three) Gaussians. The maximum likelihood ratio is referred to as InSignature score. In step 2 (middle), genes are grouped based on their InSignature patterns. Each such group is referred to as InSignature class or, in short, class. In step 3 (bottom), InSignature selects the best-scoring genes within each class; these form the variation signature genes. 6
17 Supp. Figure 2.5 Microarray data set Reshuffled data set Fraction InSignature score FDR=.5 Supplementary Figure 2. InSignature scores in microarray data. Shown is a distribution of the InSignature scores (X axis, Methods) calculated from the microarray data (Supplementary Table ; red) and in a randomized dataset (black) generated only by reshuffling the measurements of each gene independently, thus disrupting only the matching between pairs of individuals of the same genetic background. The InSignature score threshold corresponding to an FDR of.5 is 5.5, indicated in gray vertical line. 7
18 Supp. Figure 3 Observed log p-value Nfkb Tnf..5 =.5 = Observed log p-value Expected log p-value Expected log p-value Supplementary Figure 3. A visualization of confounders such as population stratification. Shown are P-P plots for the association of Nfκb (left) and Tnf (right) responsiveness following poly IC stimulation (6 hours). Observed -log P values (y axis) for all SNPs are plotted against the expected null distribution (x axis). To calculate P-values, LR scores were transformed to fit an F distribution. The observed distribution matches the expected distribution closely, with a modest genome-wide inflation (genomic control λ =.5 and.9995) and an excess only at the tail. 8
19 Supplementary Note Supplementary Note. Genome-wide heritable variation in the transcriptional response to pathogenic components We measured 3 genome-wide transcription profiles in resting and stimulated DCs from two parental strains (B6 and D2) and six BXD strains (Supplementary Table ). For the parental strains, we profiled a pool of two individuals from each strain pre-stimulation and at 2 and 6 hours after treatment with LPS, poly IC, or PAM3CSK ( PAM ) (TLR4, 3, and 2 agonists, respectively). For the six BXD strains, we measured expression profiles in two individuals from each strain stimulated with LPS, the stimulus with the broadest effect on gene expression. We used the 3 genome-wide profiles to identify loci with inherited expression variation, by testing whether the parental strains have major gene expression differences following stimulation, and whether these parental differences are inherited. First, we calculated the fold change difference among B6 and D2 (B6/D2 fold change) to evaluate for each gene the extent to which its expression in the parental strains differs under the same stimulation and time point (Methods). There are substantial D2/B6 fold changes, with % of the genes exceeding a 2-fold change in at least one stimulus. In particular, 62, 32, and 97 genes have an absolute log B6/D2 fold change that is higher than 2 at 6 hours after exposure to LPS, poly IC and PAM, respectively. The overall distribution of D2/B6 fold-changes is similar in baseline profiles and under stimulations (SN Fig. a), although the fold changes of individual genes may substantially differ in the baseline versus stimulated state (SN Fig. b). 9
20 SN Figure. D2/B6 fold changes. (a) Shown is the distribution of D2/B6 fold changes (X axis) before activation (red) and at 6 hours following LPS stimulation (black). (b) Shown are the D2/B6 fold changes before activation (X axis) and at 6 hours following LPS stimulation (Y axis). Next, we tested which of the D2/B6 fold changes under stimulation are inherited, by calculating for each gene its broad-sense heritability (H 2 ), as the percentage of between-strain variation out of the total amount of expression variation. Higher heritability indicates a more significant difference between the mean gene expression values of the BXD strains, compared to the withinstrain variation (Methods). The observed heritability values are significantly higher than expected by chance (t-test P< -2, SN Fig. 2). Although no information from the parental strains was used to infer heritability, the D2/B6 fold changes are significantly correlated to H 2 at 2
21 the same condition (LPS at 6 hours, Pearsons s r =.43, P << -2, SN Fig. 2), suggesting that the differences among parental strains are inherited. SN Figure 2. Heritability of variation in gene expression. Shown is the heritability of gene expression in BXD RI strains (Y axis) vs. the fold change in expression between the parental inbred strains (X axis) for data collected at 6h post LPS stimulation (black) and for reshuffling measurements of each gene (gray). The plots were generated by using a moving average of a window of genes. Supplementary Note 2. The variation signature selection pipeline. In this study, we used a signature of 424 genes the maximal number of genes that could be reliably assayed using the meso-scale ncounter technology at the time, given the expression levels and cell numbers used in our system. We used the following selection pipeline. First, we selected genes based on heritable variation in responsiveness, as determined by InSignature analysis of the global profiles across a small 2
22 number of strains (criterion I). This analysis is designed to represent the different underlying genetic factors and their associated genes in an unbiased manner (Methods). To these, we added, in order, genes based on prior knowledge on their biological relevance in this sytem (criterion II), positive control genes based on responsiveness to the relevant stimuli (criterion III), and negative control genes (criterion IV). Supplementary Table 3 summarizes the number of genes that were added in each step of this pipeline. Supplementary Table 2 provides a full list of all signature genes, organized by the particular pipeline criteria. Below we describe each step. Criterion I: An heritability-based (genetic-based) selection: selecting genes that represent different underlying genetic factors in an unbiased manner. A naïve systems genetic approach can select the genes with the highest heritability or highest association to any of the genome-wide genetic variants under one or a few of the stimuli. However, there are two drawbacks to this approach. First, due to the complexity of the genetic landscape, it is impossible to assume that the same genetic variant is functional under all stimuli. As the selection of signature genes is based on only a few genetic backgrounds, it is impossible to apply a standard association test on each of the stimuli. Second, choosing signature genes based on the association or heritability score alone might lead to an imbalanced selection of many genes that are significantly associated with only a few (pleiotropic) genetic variants. InSignature addresses these challenges as follows. First, InSignature associates each expression trait with the most likely genotype in each of the strains and stimulations. The most likely genotype is called a InSignature pattern and its score is called an InSignature score. The prediction and scoring scheme is designed to fit the special case of a different (or no) genetic variant under each stimulus, and as few as one or two profiled strains in some of the stimuli. 22
23 Genes whose InSignature pattern exactly matches the genotype in their proximity are termed cis predictions, and the rest are termed trans predictions. Note that the cis predictions at this stage are not necessarily controlled in cis, since using only a few genetic backgrounds, a large fraction of the genome share the same genotype (e.g., for 6 BXD strains, a fraction of :32 of the genome have exactly the same genetic information). Second, InSignature groups responsiveness traits based on their InSignature pattern. Such a group, referred to as an InSignature class (Methods), consists of all genes that have a certain InSignature pattern: Two genes within the same class have an identical InSignature pattern, whereas two genes at different classes have a distinct InSignature pattern. The partition into classes allows selection of genes that are associated with a variety of genetic factors in an unbiased manner. To this end, InSignature first randomly selects a class and then selecting a gene (if available) within that class. Four alternative criteria have been used to select a gene within a sampled class: I-a: Top in class (232 genes). Selecting the gene with the highest InSignature score within a class. I-b. Biological relevance (2 genes). Selecting a gene in the class that is a key immunerelated or disease-related gene based on prior knowledge. In this study, we used the annotation in the Ingenuity knowledge base (Ingenuity Systems, Mountain View, CA, USA). I-c. Responsiveness (35 genes). Selecting genes in the class with high responsiveness levels under (one or a few) of the stimuli in an unbiased manner. In this study, we used a dataset of 23
24 DCs responsiveness under LPS, PAM or polyic measured at nine time points (Amit et al. 29). I-d. Random (35 genes). Randomly selecting one of the class genes. The researcher needs to decide in advance how many genes are selected based on each of the criteria. Here, the distribution of number of genes that were selected using each of these categories is detailed in Supplementary Table 3, and noted next to each criterion above. The analysis in Supplementary Fig. highlight the eventual relative utility of these different options, and can guide future users to prefer for example high scoring genes over random ones in the class. Supplementary Fig. a summarizes the distribution of number of genes that were from each InSignature class. Each of the 2 genes selected based on biological relevance resides in a different class. In the case of responsiveness-based and random-based selection, two and three classes (respectively) include two selected genes whereas the remaining includes exactly one selected gene. Criterion II: Selection based on biological relevance (knowledge-based selection). In this step, we added genes that have a known key role in immune response or relevant diseases, although they have no significant InSignature score. This serves two purposes. First, previous studies reported an enrichment of genetically-linked genes among such genes (Barreiro et al, 22). Second, such genes are of broad interest to biologists studying the system. In our experience in this and other studies, a biologist researcher knowledgeable in a system would opt 24
25 to include such biological relevance genes in a signature. In this study, we selected 6 genes manually based on their annotation in the Ingenuity Knowledge based (Ingenuity Systems, Mountain View, CA, USA). Analysis of our eventual signature data (Supplementary Fig. ) indicate that such a knowledge-based selection was indeed valuable for identifying additional associations. Criterion III: Selection of positive responsiveness controls. In this step, we added 2 genes based on their high responsiveness levels to the relevant stimuli, despite having no significant InSignature score. Such genes serve two purposes. First, previous studies have reported the enrichment of reqtls underlying highly responding genes (Barreiro et al, 22). Second, they are positive controls for the overall success of the stimulation experiment. In this study, we used a dataset of DCs responsiveness under LPS, PAM or poly IC across nine different time points (Amit et al, 29). Analysis of our eventual signature data (Supplementary Fig. ) indicate that a selection based on responsiveness criteria was indeed valuable for identifying cis- and transassociations. Criterion IV: Selection of negative control genes. The negative controls are genes with low observed variation among time points, strains and individuals. These are used to evaluate false positive and true negative genes in our association study. Since variation depends on the intensity of gene expression, we chose those genes with lowest variance-to-average ratio. To cover various different levels of intensity, genes were grouped into four bins based on their level of gene expression, and five (or six) genes with lowest variance-to-average ratio were selected from each such bin. 25
26 Supplementary Note 3. The vast majority of cis-linked genes are detected at both 2 and 6 hours. To test whether 6-hour data recapitulates cis signals acting during the early response, we measured the expression of the signature genes in DCs from the same 96 mice at 2 hours following stimulation with LPS, and defined their 2-hours responsiveness traits. We find a high agreement between cis-linked responsiveness traits at 2- and 6-hours: each of the 3 cis-linked genes at 6 hours (Fig. 3a) is also cis-linked at 2-hours. 3 of 32 (96%) of the cis-linked genes at 2-hours are also cis-linked at 6 hours. The only exception is Vcan, which is cis-linked only at 2h, consistent with its early response in B6 but not in D2 genetic background (SN Figure 3). SN Figure 3. Expression of the Vcan gene. Shown are the log2 expression levels (Y axis) of Vcan gene at, 2 and 6 hours following LPS (X axis) for parental and BXD strains carrying the B6 (black) and D2 (grey) allele at the proximity of the gene. References: Amit I, Garber M, Chevrier N, Leite AP, Donner Y, Eisenhaure T et al. (29) Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science 326: Barreiro LB, Tailleux L, Pai AA, Gicquel B, Marioni JC, Gilad Y (22) Deciphering the genetic architecture of variation in the immune response to Mycobacterium tuberculosis infection. Proc Natl Acad Sci U S A 9:
27 Supplementary Tables strain stimulation time #individuals # biological repeats B6 no stimulation Baseline 2 D2 no stimulation Baseline 2 BXD2 LPS 6 hours 2 BXD5 LPS 6 hours 2 BXD4 LPS 6 hours 2 BXD6 LPS 6 hours 2 BXD28 LPS 6 hours 2 BXD4 LPS 6 hours 2 B6 LPS 6 hours 2 D2 LPS 6 hours 2 B6 PAM 6 hours D2 PAM 6 hours B6 Poly IC 6 hours D2 Poly IC 6 hours B6 LPS 2 hours D2 LPS 2 hours B6 PAM 2 hours D2 PAM 2 hours B6 Poly IC 2 hours D2 Poly IC 2 hours Total number of arrays: 3 Supplementary Table. Experimental design of microarray study. Shown are mice strains (column ) and the experiment for each strain (stimulation and time point in columns 2 and 3, respectively). Column 4 provides the number of individuals in each array. Column 5 provides the number of biological replicates. For each strain, we have measured one or two individuals; each is a biological replicate and has been monitored in a separate array. 27
28 Supplementary Table 2. Nanostring ncounter design Symbol Entrez ID Chr Strand Start End Selection criterion name Selection criterion Bcl2l heritability (Biological relevance) criterion I-b Ctnnb heritability (Biological relevance) criterion I-b Ctse heritability (Biological relevance) criterion I-b Cxcr heritability (Biological relevance) criterion I-b Ddx heritability (Biological relevance) criterion I-b Fcgr heritability (Biological relevance) criterion I-b Hdac heritability (Biological relevance) criterion I-b Ltb heritability (Biological relevance) criterion I-b Myc heritability (Biological relevance) criterion I-b Nlrp heritability (Biological relevance) criterion I-b Notch heritability (Biological relevance) criterion I-b Pltp heritability (Biological relevance) criterion I-b Ptger heritability (Biological relevance) criterion I-b Ptgs heritability (Biological relevance) criterion I-b Rasgrp heritability (Biological relevance) criterion I-b Sqle heritability (Biological relevance) criterion I-b Tgfbr heritability (Biological relevance) criterion I-b Tk heritability (Biological relevance) criterion I-b Tnf heritability (Biological relevance) criterion I-b cxcl heritability (Biological relevance) criterion I-b Emr heritability (random selection) criterion I-d 242F heritability (random selection) criterion I-d Adi heritability (random selection) criterion I-d Apm heritability (random selection) criterion I-d Cct heritability (random selection) criterion I-d Cdc heritability (random selection) criterion I-d Cep heritability (random selection) criterion I-d Dnajc heritability (random selection) criterion I-d Egln heritability (random selection) criterion I-d Elavl heritability (random selection) criterion I-d Esf heritability (random selection) criterion I-d Exosc heritability (random selection) criterion I-d Fam7b heritability (random selection) criterion I-d Fdft heritability (random selection) criterion I-d Idi heritability (random selection) criterion I-d Il7ra heritability (random selection) criterion I-d Itga heritability (random selection) criterion I-d Kif3b heritability (random selection) criterion I-d Kitl heritability (random selection) criterion I-d Klf heritability (random selection) criterion I-d Lair heritability (random selection) criterion I-d Lss heritability (random selection) criterion I-d Mcm heritability (random selection) criterion I-d Pmvk heritability (random selection) criterion I-d Pqlc heritability (random selection) criterion I-d Rgs heritability (random selection) criterion I-d Setd heritability (random selection) criterion I-d
29 Supplementary Table 2. Nanostring ncounter design (cont) Symbol Entrez ID Chr Strand Start End Selection criteria Selection criterion Sgk heritability (random selection) criterion I-d Slc6a heritability (random selection) criterion I-d Socs heritability (random selection) criterion I-d Socs heritability (random selection) criterion I-d Timp heritability (random selection) criterion I-d Tjp heritability (random selection) criterion I-d Tmem heritability (random selection) criterion I-d Zmat heritability (random selection) criterion I-d Adhfe heritability (responsiveness) criterion I-c Bysl heritability (responsiveness) criterion I-c Ccdc heritability (responsiveness) criterion I-c Cdd heritability (responsiveness) criterion I-c Cd heritability (responsiveness) criterion I-c Cox heritability (responsiveness) criterion I-c Dusp heritability (responsiveness) criterion I-c E2f heritability (responsiveness) criterion I-c H2-Oa heritability (responsiveness) criterion I-c Ikbke heritability (responsiveness) criterion I-c Il2rb heritability (responsiveness) criterion I-c Itgav heritability (responsiveness) criterion I-c Lamc heritability (responsiveness) criterion I-c Lgals heritability (responsiveness) criterion I-c Nr4a heritability (responsiveness) criterion I-c Oasl heritability (responsiveness) criterion I-c Palld heritability (responsiveness) criterion I-c Ppa heritability (responsiveness) criterion I-c Pwp heritability (responsiveness) criterion I-c Ripk heritability (responsiveness) criterion I-c Rrs heritability (responsiveness) criterion I-c Slc3a heritability (responsiveness) criterion I-c Slc7a heritability (responsiveness) criterion I-c Smox heritability (responsiveness) criterion I-c Sp heritability (responsiveness) criterion I-c Stip heritability (responsiveness) criterion I-c Stk heritability (responsiveness) criterion I-c Tgif heritability (responsiveness) criterion I-c Tjp heritability (responsiveness) criterion I-c Tlr heritability (responsiveness) criterion I-c Tnfrsfb heritability (responsiveness) criterion I-c Tor3a heritability (responsiveness) criterion I-c Trem heritability (responsiveness) criterion I-c Zbp heritability (responsiveness) criterion I-c 234L heritability (responsiveness) criterion I-c 92H heritability (top in class) criterion I-a 2IR heritability (top in class) criterion I-a 222G heritability (top in class) criterion I-a
30 Supplementary Table 2. Nanostring ncounter design (cont) Symbol Entrez ID Chr Strand Start End Selection criteria Selection criterion J heritability (top in class) criterion I-a G heritability (top in class) criterion I-a Acox heritability (top in class) criterion I-a Acss heritability (top in class) criterion I-a Adcy heritability (top in class) criterion I-a Adssl heritability (top in class) criterion I-a Afp heritability (top in class) criterion I-a Akrb heritability (top in class) criterion I-a Aldha heritability (top in class) criterion I-a Aps heritability (top in class) criterion I-a Apbb heritability (top in class) criterion I-a Appl heritability (top in class) criterion I-a Arhgdib heritability (top in class) criterion I-a Arid5a heritability (top in class) criterion I-a Atad3a heritability (top in class) criterion I-a Batf heritability (top in class) criterion I-a Baz2a heritability (top in class) criterion I-a BC heritability (top in class) criterion I-a Bcat heritability (top in class) criterion I-a Bckdhb heritability (top in class) criterion I-a Bid heritability (top in class) criterion I-a Blvra heritability (top in class) criterion I-a Bub heritability (top in class) criterion I-a Bxdc heritability (top in class) criterion I-a C5ar heritability (top in class) criterion I-a Calcrl heritability (top in class) criterion I-a Cald heritability (top in class) criterion I-a Cap heritability (top in class) criterion I-a Carhsp heritability (top in class) criterion I-a Ccdc heritability (top in class) criterion I-a Ccl heritability (top in class) criterion I-a Ccnd heritability (top in class) criterion I-a Cct6a heritability (top in class) criterion I-a Cdd heritability (top in class) criterion I-a Cd heritability (top in class) criterion I-a Cd heritability (top in class) criterion I-a Cd heritability (top in class) criterion I-a Cd heritability (top in class) criterion I-a Chi3l heritability (top in class) criterion I-a Chi3l heritability (top in class) criterion I-a Ciita heritability (top in class) criterion I-a Cited heritability (top in class) criterion I-a Clcn heritability (top in class) criterion I-a Clec5a heritability (top in class) criterion I-a Ctla2b heritability (top in class) criterion I-a Ctsk heritability (top in class) criterion I-a Cttn heritability (top in class) criterion I-a
31 Supplementary Table 2. Nanostring ncounter design (cont) Symbol Entrez ID Chr Strand Start End Selection criteria Selection criterion Cxcl heritability (top in class) criterion I-a Dbi heritability (top in class) criterion I-a Ddx heritability (top in class) criterion I-a Ddx heritability (top in class) criterion I-a Dgka heritability (top in class) criterion I-a Dhrs heritability (top in class) criterion I-a Dhx heritability (top in class) criterion I-a Dnajb heritability (top in class) criterion I-a Dtwd heritability (top in class) criterion I-a Dusp heritability (top in class) criterion I-a Dusp heritability (top in class) criterion I-a Egr heritability (top in class) criterion I-a Ehd heritability (top in class) criterion I-a Emilin heritability (top in class) criterion I-a Enah heritability (top in class) criterion I-a Eps heritability (top in class) criterion I-a Ereg heritability (top in class) criterion I-a Ets heritability (top in class) criterion I-a Etv heritability (top in class) criterion I-a Fabp heritability (top in class) criterion I-a Fam5a heritability (top in class) criterion I-a Flnb heritability (top in class) criterion I-a Ftsj heritability (top in class) criterion I-a Gdf heritability (top in class) criterion I-a Gpnmb heritability (top in class) criterion I-a Gpr9a heritability (top in class) criterion I-a Gpr37b-ps heritability (top in class) criterion I-a Gramd heritability (top in class) criterion I-a Gtf2e heritability (top in class) criterion I-a Gusb heritability (top in class) criterion I-a Gyk heritability (top in class) criterion I-a Hebp heritability (top in class) criterion I-a Hfe heritability (top in class) criterion I-a Hice heritability (top in class) criterion I-a Hmg2a heritability (top in class) criterion I-a Hmgn heritability (top in class) criterion I-a Ifi heritability (top in class) criterion I-a Ifna heritability (top in class) criterion I-a Ifnb heritability (top in class) criterion I-a Il heritability (top in class) criterion I-a Ilf heritability (top in class) criterion I-a Il heritability (top in class) criterion I-a Il6st heritability (top in class) criterion I-a Il7r heritability (top in class) criterion I-a Inpp5d heritability (top in class) criterion I-a Ints heritability (top in class) criterion I-a Ipo heritability (top in class) criterion I-a
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
CPM (x 10-3 ) Tregs +Teffs. Tregs alone ICOS CLTA-4
 A 2,5 B 4 Number of cells (x 1-6 ) 2, 1,5 1, 5 CPM (x 1-3 ) 3 2 1 5 1 15 2 25 3 Days of culture 1/1 1/2 1/4 1/8 1/16 1/32 Treg/Teff ratio C alone alone alone alone CD25 FoxP3 GITR CD44 ICOS CLTA-4 CD127
A 2,5 B 4 Number of cells (x 1-6 ) 2, 1,5 1, 5 CPM (x 1-3 ) 3 2 1 5 1 15 2 25 3 Days of culture 1/1 1/2 1/4 1/8 1/16 1/32 Treg/Teff ratio C alone alone alone alone CD25 FoxP3 GITR CD44 ICOS CLTA-4 CD127
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Nature Immunology: doi: /ni.3745
 Supplementary Figure 1 Experimental settings. (a) Experimental setup and bioinformatic data analysis of transcriptomic changes induced in neonatal and adult monocytes by stimulation with 10 ng/ml LPS for
Supplementary Figure 1 Experimental settings. (a) Experimental setup and bioinformatic data analysis of transcriptomic changes induced in neonatal and adult monocytes by stimulation with 10 ng/ml LPS for
An integrated transcriptomic and functional genomic approach identifies a. type I interferon signature as a host defense mechanism against Candida
 Supplementary information for: An integrated transcriptomic and functional genomic approach identifies a type I interferon signature as a host defense mechanism against Candida albicans in humans Sanne
Supplementary information for: An integrated transcriptomic and functional genomic approach identifies a type I interferon signature as a host defense mechanism against Candida albicans in humans Sanne
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response
 Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Supplementary Materials
 1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Single cell RNA Seq reveals dynamic paracrine control of cellular variation
 Single cell RNA Seq reveals dynamic paracrine control of cellular variation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Single cell RNA Seq reveals dynamic paracrine control of cellular variation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Deconvolution of Heterogeneous Wound Tissue Samples into Relative Macrophage Phenotype Composition via Models based on Gene Expression
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2017 Supplementary Information for: Deconvolution of Heterogeneous Wound Tissue Samples into
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2017 Supplementary Information for: Deconvolution of Heterogeneous Wound Tissue Samples into
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Chromatin marks identify critical cell-types for fine-mapping complex trait variants
 Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
* Kyoto Encyclopedia of Genes and Genomes.
 Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
SUPPLEMENTARY INFORMATION
 Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created.
 938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
SUPPLEMENTAL INFORMATIONS
 1 SUPPLEMENTAL INFORMATIONS Figure S1 Cumulative ZIKV production by testis explants over a 9 day-culture period. Viral titer values presented in Figure 1B (viral release over a 3 day-culture period measured
1 SUPPLEMENTAL INFORMATIONS Figure S1 Cumulative ZIKV production by testis explants over a 9 day-culture period. Viral titer values presented in Figure 1B (viral release over a 3 day-culture period measured
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
qpcr-array Analysis Service
 qpcr-array Analysis Service Customer Name Institute Telephone Address E-mail PO Number Service Code Report Date Service Laboratory Department Phalanx Biotech Group, Inc 6 Floor, No.6, Technology Road 5,
qpcr-array Analysis Service Customer Name Institute Telephone Address E-mail PO Number Service Code Report Date Service Laboratory Department Phalanx Biotech Group, Inc 6 Floor, No.6, Technology Road 5,
Supplementary Data for:
 Supplementary Data for: Tumour sampling method can significantly influence gene expression profiles derived from neoadjuvant window studies Dominic A. Pearce 1, Laura M. Arthur 1, Arran K. Turnbull 1,
Supplementary Data for: Tumour sampling method can significantly influence gene expression profiles derived from neoadjuvant window studies Dominic A. Pearce 1, Laura M. Arthur 1, Arran K. Turnbull 1,
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Gene Ontology 2 Function/Pathway Enrichment. Biol4559 Thurs, April 12, 2018 Bill Pearson Pinn 6-057
 Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC
 1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
1 2 1 3 Supplementary figure 1: LII/III GIN-cells show morphological characteristics of MC 4 5 6 7 (a) Reconstructions of LII/III GIN-cells with somato-dendritic compartments in orange and axonal arborizations
Lecture 8 Understanding Transcription RNA-seq analysis. Foundations of Computational Systems Biology David K. Gifford
 Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Lecture 8 Understanding Transcription RNA-seq analysis Foundations of Computational Systems Biology David K. Gifford 1 Lecture 8 RNA-seq Analysis RNA-seq principles How can we characterize mrna isoform
Supplementary materials for: Executive control processes underlying multi- item working memory
 Supplementary materials for: Executive control processes underlying multi- item working memory Antonio H. Lara & Jonathan D. Wallis Supplementary Figure 1 Supplementary Figure 1. Behavioral measures of
Supplementary materials for: Executive control processes underlying multi- item working memory Antonio H. Lara & Jonathan D. Wallis Supplementary Figure 1 Supplementary Figure 1. Behavioral measures of
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Bezzi et al., Supplementary Figure 1 *** Nature Medicine: doi: /nm Pten pc-/- ;Zbtb7a pc-/- Pten pc-/- ;Pml pc-/- Pten pc-/- ;Trp53 pc-/-
 Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Identifying the Zygosity Status of Twins Using Bayes Network and Estimation- Maximization Methodology
 Identifying the Zygosity Status of Twins Using Bayes Network and Estimation- Maximization Methodology Yicun Ni (ID#: 9064804041), Jin Ruan (ID#: 9070059457), Ying Zhang (ID#: 9070063723) Abstract As the
Identifying the Zygosity Status of Twins Using Bayes Network and Estimation- Maximization Methodology Yicun Ni (ID#: 9064804041), Jin Ruan (ID#: 9070059457), Ying Zhang (ID#: 9070063723) Abstract As the
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists.
 chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families
 SUPPLEMENTAL MATERIAL Supplemental Tables: Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families "#$%&' (')*'%$+,-'
SUPPLEMENTAL MATERIAL Supplemental Tables: Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families "#$%&' (')*'%$+,-'
Main Study: Summer Methods. Design
 Main Study: Summer 2000 Methods Design The experimental design is within-subject each participant experiences five different trials for each of the ten levels of Display Condition and for each of the three
Main Study: Summer 2000 Methods Design The experimental design is within-subject each participant experiences five different trials for each of the ten levels of Display Condition and for each of the three
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB
 RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Gene expression analysis. Roadmap. Microarray technology: how it work Applications: what can we do with it Preprocessing: Classification Clustering
 Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Genetics of extreme body size evolution in mice from Gough Island
 Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
SUPPLEMENTARY INFORMATION. Confidence matching in group decision-making. *Correspondence to:
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 1 ARTICLE NUMBER: 0117 Confidence matching in group decision-making an Bang*, Laurence Aitchison, Rani Moran, Santiago
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 1 ARTICLE NUMBER: 0117 Confidence matching in group decision-making an Bang*, Laurence Aitchison, Rani Moran, Santiago
Eosinophils! 40! 30! 20! 10! 0! NS!
 A Macrophages Lymphocytes Eosinophils Neutrophils Percentage (%) 1 ** 4 * 1 1 MMA SA B C Baseline FEV1, % predicted 15 p = 1.11 X 10-9 5 CD4:CD8 ratio 1 Supplemental Figure 1. Cellular infiltrate in the
A Macrophages Lymphocytes Eosinophils Neutrophils Percentage (%) 1 ** 4 * 1 1 MMA SA B C Baseline FEV1, % predicted 15 p = 1.11 X 10-9 5 CD4:CD8 ratio 1 Supplemental Figure 1. Cellular infiltrate in the
Complex Traits Activity INSTRUCTION MANUAL. ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik
 Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Material for
 Supplementary Material for Selective neuronal lapses precede human cognitive lapses following sleep deprivation Supplementary Table 1. Data acquisition details Session Patient Brain regions monitored Time
Supplementary Material for Selective neuronal lapses precede human cognitive lapses following sleep deprivation Supplementary Table 1. Data acquisition details Session Patient Brain regions monitored Time
An expanded view of complex traits: from polygenic to omnigenic
 BIRS 2017 An expanded view of complex traits: from polygenic to omnigenic How does human genetic variation drive variation in complex traits/disease risk? Yang I Li Stanford University Evan Boyle Jonathan
BIRS 2017 An expanded view of complex traits: from polygenic to omnigenic How does human genetic variation drive variation in complex traits/disease risk? Yang I Li Stanford University Evan Boyle Jonathan
15. Supplementary Figure 9. Predicted gene module expression changes at 24hpi during HIV
 Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science. Homework 5
 PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
PSYCH-GA.2211/NEURL-GA.2201 Fall 2016 Mathematical Tools for Cognitive and Neural Science Homework 5 Due: 21 Dec 2016 (late homeworks penalized 10% per day) See the course web site for submission details.
A genome-wide association study identifies vitiligo
 A genome-wide association study identifies vitiligo susceptibility loci at MHC and 6q27 Supplementary Materials Index Supplementary Figure 1 The principal components analysis (PCA) of 2,546 GWAS samples
A genome-wide association study identifies vitiligo susceptibility loci at MHC and 6q27 Supplementary Materials Index Supplementary Figure 1 The principal components analysis (PCA) of 2,546 GWAS samples
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Supplemental Information. Genomic Characterization of Murine. Monocytes Reveals C/EBPb Transcription. Factor Dependence of Ly6C Cells
 Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas
 Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
HTG EdgeSeq Immuno-Oncology Assay Gene List
 A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
Nature Neuroscience: doi: /nn Supplementary Figure 1. Trial structure for go/no-go behavior
 Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
To compare the relative amount of of selected gene expression between sham and
 Supplementary Materials and Methods Gene Expression Analysis To compare the relative amount of of selected gene expression between sham and mice given renal ischemia-reperfusion injury (IRI), ncounter
Supplementary Materials and Methods Gene Expression Analysis To compare the relative amount of of selected gene expression between sham and mice given renal ischemia-reperfusion injury (IRI), ncounter
Psychology Research Process
 Psychology Research Process Logical Processes Induction Observation/Association/Using Correlation Trying to assess, through observation of a large group/sample, what is associated with what? Examples:
Psychology Research Process Logical Processes Induction Observation/Association/Using Correlation Trying to assess, through observation of a large group/sample, what is associated with what? Examples:
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
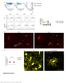 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Medical Statistics 1. Basic Concepts Farhad Pishgar. Defining the data. Alive after 6 months?
 Medical Statistics 1 Basic Concepts Farhad Pishgar Defining the data Population and samples Except when a full census is taken, we collect data on a sample from a much larger group called the population.
Medical Statistics 1 Basic Concepts Farhad Pishgar Defining the data Population and samples Except when a full census is taken, we collect data on a sample from a much larger group called the population.
