SUPPLEMENTARY INFORMATION
|
|
|
- Kenneth Webb
- 5 years ago
- Views:
Transcription
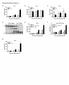
1 doi: 1.138/nature89 IFN- (ng ml ) Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) *** ** Lymph node cells Supplementary Figure 1 IFN-γ and IL- production in splenocytes and lymph node cells from immunized mice. IFN- and IL- production in splenocytes and lymph node cells after restimulation with anti-cd3 and anti-cd8. Error bars denote mean ± s.e.m. **P <.1; ***P <.5; NS, not significant. 1
2 doi: 1.138/nature NS NS 18 NS TNF- (ng ml ) 1..5 IL-6 (ng ml ) 1 6 * Nfkbiz / None LPS None CpG None LPS None CpG cdcs pdcs cdcs pdcs Supplementary Figure LPS- or CpG DNA-induced TNF- and IL-6 production in or Nfkbiz / DCs. cdcs or pdcs isolated from or Nfkbiz / mice were stimulated with LPS or CpG DNA. Twenty-four hours later, TNF- and IL-6 production was assessed by ELISA. Error bars denote mean ± s.e.m. *P <.5; NS, not significant.
3 doi: 1.138/nature89 Nfkbiz / Rag / Rag / Rag / 1.8%.% 6.% 5.5% CD4 CD3 Supplementary Figure 3 Similar reconstitution efficiency in mice that received or Nfkbiz / CD4 T cells. The populations of CD3! CD4 + T cells in the spleen of Rag / mice reconstituted with or Nfkbiz / CD4 T cells. Representative data from three mice 1 days after immunization. 3
4 doi: 1.138/nature89 5 NS BrdU + (% of CD4 + ) Nfkbiz -/- (5 5.%) (7.6%) Supplementary Figure 4 No difference in proliferation between and Nfkbiz / CD4 + T cells in vivo after MOG immunization. Rag / mice reconstituted with or Nfkbiz / CD4 T cells were administered with BrdU 1 days after MOG immunization, and then the proliferation of CD4 T cells was measured by BrdU incorporation. NS, not significant. 4
5 doi: 1.138/nature Runx1 Ahr Irf4 Socs3 Batf () () () () () 16 Ccr6 3 Ccl 18 S1pr1 4 Il6ra 4 Il6st () () () 3 1 () 3 1 () 3 Il1r.5 Ilrg 1 Il1r1 1.5 Il1rap 1 () () () 1..5 () Nfkbiz / Supplementary Figure 5 The mrna expression level of genes related to cells in Nfkbiz / T cells. The mrna expression of genes related to development (Runx1, Ahr, Irf4, Socs3 and Batf), as well as genes related to the migration of cells (Ccr6, Ccl and S1pr1) and -related cytokine receptors (Il6ra, Il6st, Il1r, Ilrg, Il1r1 and Il1rap), in or Nfkbiz / CD4 T cells activated under - and -polarizing conditions were analyzed by quantitative RT-PCR. CCR6 expression was slightly decreased (but not statistically significant) in Nfkbiz / T cells under -polarizing conditions. The resistance of Nfkbiz / mice to EAE is mainly due to a defect in the early development of cells, although a contribution of a decrease in CCR6-mediated migration of cells to the central nervous system cannot be completely excluded. Error bars denote mean ± s.e.m. ignificant. 5
6 doi: 1.138/nature89 IL-6 min 3 min 6 min 18 min Control IgG phospho-stat3 Nfkbiz / phospho-stat3 Supplementary Figure 6 IL-6-induced phosphorylation of STAT3 in Nfkbiz / T cells. and Nfkbiz / naive CD4 T cells were stimulated with anti-cd3 and anti-cd8 in the presence of IL-6 for the indicated periods, and then stained with anti-phospho-stat3 or control IgG. 6
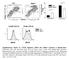
7 doi: 1.138/nature89 a Thymus Spleen Lymph nodes.6% 9.% 8.4% Nfkbiz / Foxp3.7% 8.1% 7.9% CD5 b Neutral TGF- IL-3 IL-1 TGF- IL-3.9% 43.% 8.8% 19.1% 3.4% 1.1% 49.9% 14.3% 16.9% 3.9% Nfkbiz / Foxp3 Supplementary Figure 7 Targeted disruption of the Nfkbiz gene has no effect on the development of natural T reg and inducible T reg cells. a, CD4 CD5 Foxp3 natural T reg cells in the thymus, spleen or lymph nodes of mice and Nfkbiz / mice. b, Foxp3 expression in or Nfkbiz / CD4 T cells activated under the indicated conditions. There was no defect in the development of inducible T reg cells from Nfkbiz / naive CD4 T cells. Under -polarizing conditions, there was no difference in Foxp3 expression between and Nfkbiz / T cells. 7
8 doi: 1.138/nature89 a Relative Nfkbiz mrna level RPMI IMDM * * None IL-3 IL-3 anti-il None IL-3 IL-3 anti-il b M IB (L) IB (D) LPS-stimulated M 14 bp 818 bp IL-3 14 bp 818 bp c Relative Nfkbiz mrna level RPMI IMDM Supplementary Figure 8 Nfkbiz mrna is highly expressed in cells. a, Quantitative RT-PCR analysis of Nfkbiz mrna expression in CD4 T cells activated under different -polarizing conditions. cells expressed a high level of Nfkbiz mrna when cultured in RPMI containing IL-6 and TGF-. The expression of Nfkbiz mrna was further increased if cultured in the presence of anti-il or in IMDM. The primer set used in Figs 1a, f and g and Supplementary Fig.8a was designed to detect both IB (L) and (S). b, The mrna expression of IB (D) in bone marrow-derived macrophage (M), LPS-stimulated M, cell subsets (top). RT-PCR was performed using a primer set [detecting both (D) and (L) isoforms] designed to amplify the region between nucleotides 141 and 154 of the cdna for IB (L) (bottom). IB (D) expression in cells was found to be under the detection limit of RT-PCR. c, The expression of mrna for IB (L) was analyzed using an IB (L)-specific primer set by quatitative RT-PCR. Error bars (a and c) denote mean ± s.e.m. *P <.5. 9 IL-3 * IL-3 anti-il it reg * 9 Naive IL-3 IL-3 anti-il 8
9 doi: 1.138/nature89 a (kda) Naive 1 Anti-IB it reg 9 IB(L) IB(S) ns b (kda) 75 5 None IL-6 TGF- IL-3 Anti-IB IL-3 IB(L) IB(S) ns Relative expression (IB (L)/ -actin) Anti--actin *** Relative expression (IB (L)/ -actin) Anti--actin ** ** c Relative expression (IB (L)/ -actin) (kda) None RPMI IL-3 IL-3 anti-il- RPMI IL-3 Anti-IB Anti--actin * None * IMDM IL-3 IL-3 anti-il- ** IB(L) IB(S) ns d Relative expression (IB (L)/ -actin) (kda) Myd88 / Ila / Anti-IB Anti--actin *** Rorc / Stat3 flox/ Lck-Cre IB(L) IB(S) ns Supplementary Figure 9 Protein expression level of the IB splicing variants in cells. a, Protein expression level of the IB splicing variants [IB (L) (79 kda), IB (S) (69 kda) and IB (D) (57 kda)] in naive CD4 T cells, cell subsets and inducible T reg (it reg ) cells. Consistent with the results of RT-PCR analysis (Fig. 1a and Supplementary Fig. 8c), IB expression was moderately increased in 9 cells. b, Effects of the cytokines on the protein expression of the IB splicing variants in CD4 T cells. c, Protein expression level of IB in CD4 T cells activated under different -polarizing conditions. cells expressed high levels of IB (L) and (S) when cultured in RPMI containing IL-6 and TGF-. The expression of IB was further increased if cultured in the presence of anti-il or in IMDM. d, Protein expression level of IB in CD4 T cells derived from Myd88 -/-, Ila -/-, Rorc -/- or Stat3 flox/- Lck-Cre mice. ns, non-specific band. The expression of IB in cells was more obvious at the protein level than at the mrna level, suggesting the involvement of the post-transcriptional regulatory mechanisms that were reported in previous studies 1,. Error bars denote mean ± s.e.m. *P <.5; **P <.1; ***P <.5; NS, not significant. 9
10 doi: 1.138/nature89 pmx 1.% 3.8% pmx-ib (L).1% 14.% IL- 4.6% 54.7% 5.6% 33.3% EGFP Supplementary Figure 1 Ectopic expression of IB enhances IL- production in a cell-intrinsic manner. Retroviral expression of IB (L) led to a high production of IL-. EGFP- nontransduced cells did not exhibit IL- production. 1
11 doi: 1.138/nature89 pmx pmx-rela pmx-crel pmx-p5 1.3%.%.% 1.7% pmx pmx-rela pmx-crel pmx-p5 8.8% 9.3% 9.4% 8.8% pmx pmx- IB (L) IL-.6%.5%.5%.6% FSC 19.% 9.% 14.3% 6.9% Supplementary Figure 11 Effects of the NF-B subunits on the ability of IB to induced IL- production. Naive CD4 T cells were co-transduced with an NF-B subunit (RelA, crel or p5)-expressing retrovirus (IRES-EGFP) and IB (L)-expressing retrovirus (IRES-hCD), and then IL- production in EGFP hcd cells was assessed. FSC, forward scatter. 11
12 doi: 1.138/nature89 a Neutral IL-3 IL-3 IL-1 TNF-.% 5.3% 5.7% 1.% 1.%.4%.3% 1.6%.1% 8.% 8.8%.5% Nfkb1 / IL- 3.%.5%.4% 1.5% b IFN- Nfkb1 / Ilf Il Il3r Il Neutral IL-3 Neutral IL-3 Neutral IL-3 Neutral IL-3 Supplementary Figure 1 Normal differentiation of Nfkb1 / naive CD4 T cells into cells. a, Intracellular expression of IFN- and IL- in or Nfkb1 / CD4 T cells activated under - or -polarizing conditions. b, Ilf, Il1, Il3r and Il mrna expression in or Nfkb1 / CD4 T cells activated under - or -polarizing conditions. Error bars denote mean ± s.e.m. 1
13 doi: 1.138/nature89 a pmx pmx-ib (L) 4.4%.3% 6.1% 19.3% Nfkb1 / IL- EGFP b Ilf Il1 Il3r pmx pmx-ib (L) pmx pmx-ib (L) pmx pmx-ib (L) Nfkb1 / Supplementary Figure 13 NF-B p5 is not essential for IB-mediated regulation of development. a, Effects of retroviral expression of IB (L) on IL- production in or Nfkb1 / CD4 T cells activated under -polarizing conditions. b, Effects of retroviral expression of IB (L) on Ilf, Il1 and Il3r mrna expression in or Nfkb1 / CD4 T cells activated under -polarizing conditions. Error bars denote mean ± s.e.m. 13
14 doi: 1.138/nature89 Control Control.9% Control IB(L) 3.5% Control IB(S) 1.6% RORt Control.9% RORt IB(L) 48.6% RORt IB(S) 3.% ROR Control 16.4% ROR IB(L) 38.8% ROR IB(S) 3.9% (IRES-EGFP) (IRES-hCD) (+ anti-tgf-) IL- FSC Supplementary Figure 14 Effects of overexpression of the ROR nuclear receptors and IB in the presence of anti-tgf-. Intracellular expression of IL- in CD4 T cells transduced with IB (IRES-hCD) and RORt or ROR (IRES-EGFP) after activation under -polarizing conditions with anti-tgf-. Even in the presence of anti-tgf-, the ROR nuclear receptors and IB synergistically promoted development. When the anti-tgf- was added to the cultures, it was found that RORt or ROR induced IL- production significantly, possibly because of an inhibition of Foxp3 expression 3. FSC, forward scatter. 14
15 doi: 1.138/nature89 a 3 1 NS Il9 NS Nfkbiz / IL-3 b pmx- pmxpmx RORt ROR 1.6% 51.5% 34.%.5% 1.% 16.% IL-1 Nfkbiz / 9.6% 54.8% 4.3% 3.9% 9.5% 13.% IL-9 Nfkbiz / IL- EGFP Supplementary Figure 15 The defect in the ROR nuclear receptor-induced IL- production in Nfkbiz / T cells is independent of IL-1 or IL-9 expression. a, Il9 mrna expression was normal in Nfkbiz / T cells under -polarizing conditions. Error bars denote mean ± s.e.m. NS, not significant. b, Even when 8 ng ml 1 IL-1 or ng ml 1 IL-9 were added into the cultures, IL- production by RORt or ROR overexpression was impaired in Nfkbiz / CD4 T cell. 15
16 doi: 1.138/nature89 5 *** *** *** *** IL- cells (% of EGFP cells) ** *** Rorc / Rora sg/sg Rorc / Rora sg/sg Rorc / Rora sg/sg pmx pmx- IB (L) pmx- IB (S) Supplementary Figure 16 Effects of retroviral expression of IB on the generation of IL--producing cells in, Rorc / or Rora sg/sg CD4 T cells. Bar graphs represent the frequency of IL--producing cells among EGFP cells (see Fig. 4c for the dot plots). The data are shown as mean ± s.e.m of three independent experiments. **P <.1; ***P <.5. 16
17 doi: 1.138/nature89 a CNS TK pro Luc Luc Il-Luc Il CNS-Luc : ROR response element : IB response element Luc Il 4371-Luc Luc Il 1547-Luc b Relative luciferase activity 1 +1 Il CNS-Luc IB (L) IB (S) IB (D) RORt ROR RORt ROR c Relative luciferase activity Il 4371-Luc IB (L) IB (S) d Relative luciferase activity Il 1547-Luc IB (L) IB (S) RORt ROR RORt ROR RORt ROR RORt ROR Supplementary Figure IB promotes the CNS enhancer activity together with RORt and/or ROR. a, Schematic of a series of reporter constructs containing the regions upstream of the mouse Ila transcriptional initiation site. Il CNS-Luc carries the CNS region upstream of a minimal thymidine kinase promoter (TK pro) linked to the luciferase (Luc) gene. Il 4371-Luc and Il 1547-Luc contain the 4371 bp and the 1547 bp promoter region of the mouse Ila gene, respectively. Il 4371-Luc and Il 1547-Luc lack the CNS region. b, c, d, Effects of the IB variants, RORt and/or ROR on the reporter activity of Il CNS-Luc (b), Il 4371-Luc (c) or Il 1547-Luc (d). Error bars (b-d) denote mean ± s.e.m.
18 doi: 1.138/nature89 a CNS ISE1 ISE ISE TK pro Luc Il CNS-Luc TK pro Luc Il CNS mut1-luc TK pro Luc Il CNS mut-luc TK pro Luc Il CNS mut3-luc : ROR response element : IB response element b Relative luciferase activity 1 Il CNS-Luc Il CNS mut1-luc Il CNS mut-luc Il CNS mut3-luc - IB (S) RORt RORt IB (S) ROR ROR IB (S) RORt ROR RORt ROR IB (S) Supplementary Figure 18 Requirement of ISE1, an IB response element, for IB-mediated CNS enhancer activity. a, Three IB response elements were located in the CNS region (ISE1, -635 to -6339; ISE, to -4786; ISE3, to -4445). Il CNS mut1-luc, Il CNS mut-luc and Il CNS mut3-luc carry a mutation of ISE1, ISE and ISE3, respectively. b, Effects of IB (S), RORt and/or ROR on the reporter activity of Il CNS-Luc, Il CNS mut1-luc, Il CNS mut-luc and Il CNS mut3-luc. Error bars denote mean ± s.e.m. 18
19 doi: 1.138/nature89 a Ilf -154 to kb +6kb +4kb +kb +1 -kb -4kb -6kb -8kb CNS7 Human vs Mouse Relative binding Ilf promoter -154 to -163 Relative binding CNS7-744 to Nfkbiz/ Nfkbiz/ Con IgG Anti-IB b Il1 promoter -31 to -4 +1kb +8kb +6kb Il1 +4kb -31 to -4 +kb +1 -kb -4kb Human vs Mouse Relative binding Con IgG Anti-IB.1 c Il3r Nfkbiz / -1kb +8kb +6kb +4kb +kb to to kb Human vs Mouse Rat vs Mouse Relative binding Il3r promoter -13 to -14 Relative binding Il3r promoter to -143 Nfkbiz/ Nfkbiz/ Con IgG Anti-IB Supplementary Figure 19 Recruitment of IB to the Ilf, Il1 and Il3r promoter in cells. a, rvista alignment plot displaying the sequence similarity between the human and mouse Ilf loci (left). The transcriptional initiation site was designated as +1. Putative IB response sites were found within the proximal promoter region of the Ilf gene (-154 to -163) and the CNS7 region located -7 kb from the transcriptional start site (-744 to -7433). ChIP analysis revealed binding of IB to the CNS7 region, but not the proximal promoter, in cells (right). b, rvista alignment plot displaying the sequence similarity between the human and mouse Il1 loci (left). Recruitment of IB in cells to the conserved proximal promoter region of the Il1 gene (-31 to -4) (right). c, rvista alignment plot displaying the sequence similarity between the human, rat and mouse Il3r loci (left). Two putative IB response sites were found within the promoter region of the mouse Il3r gene (-13 to -14 and to -143). IB was recruited only to the distal element in the mouse Il3r promoter, which is conserved between mice and rats (right). It is also possible that the decreased expression of Il3r mrna in Nfkbiz / T cells is partly due to the impaired expression of IL-1. Error bars denote mean ± s.e.m. 19
20 doi: 1.138/nature89 Supplementary Table 1 Primers used for RT-PCR. Gene Forward primer Reverse primer Gapdh 5 -AACTTTGGCATTGTGGAAGG-3 5 -GGATGCAGGGATGATGTTCT-3 Nfkbiz # 5 -TCCAGAATGTCCCAGTCTCC-3 5 -GAGTCTCAGTTTGGGGTGGA-3 Nfkbiz (LD) 5 -CGACACCGGCTACCTGTC-3 5 -CACCAAGGTTTACCAGATCTTG-3 Nfkbiz (L) 5 -CGCTCAACCTGGCTTACTTC-3 5 -GGGCTCAACTTGAGGGCG-3 Rorc 5 -TGCAAGACTCATCGACAAGG-3 5 -AGGGGATTCAACATCAGTGC-3 Rora 5 -GAACACCTTGCCCAGAACAT-3 5 -AGCTGCCACATCACCTCTCT-3 Ilf 5 -CAAAACCAGGGCATTTCTGT-3 5 -ATGGTGCTGTCTTCCTGACC-3 Il1 5 -CGCCTCCTGATTAGACTTCG-3 5 -GCCCCTTTACATCTTGTGGA-3 Il 5 -TCATCGGGGAGAAACTGTTC-3 5 -CATGTAGGGCTGGAACCTGT-3 Il3r 5 -GGTCTTCTTGGCCATCATGT-3 5 -GCCACTTTGGGATCATCAGT-3 Il9 5 -ATCACGTGTCCGTCCTTTTC-3 5 -TCTTCATGGTCGGCTTTTCT-3 Runx1 5 -TACCTGGGATCCATCACCTC-3 5 -GACGGCAGAGTAGGGAACTG-3 Ahr 5 -AGCAGCTGTGTCAGATGGTG-3 5 -CTGAGCAGTCCCCTGTAAGC-3 Irf4 5 -GCAGCTCACTTTGGATGACA-3 5 -CCAAACGTCACAGGACATTG-3 Socs3 5 -AGCTCCAAAAGCGAGTACCA-3 5 -TGACGCTCAACGTGAAGAAG-3 Batf 5 -CCAGAAGAGCCGACAGAGAC-3 5 -GAGCTGCGTTCTGTTTCTCC-3 Ccr6 5 -TCCAGGCAACCAAATCTTTC-3 5 -GATGAACCACACTGCCACAC-3 Ccl 5 -CGACTGTTGCCTCTCGTACA-3 5 -CACCCAGTTCTGCTTTGGAT-3 S1pr1 5 -TTGCCACCCCAGATCTATTC-3 5 -TTGCCACCCCAGATCTATTC-3 Il6ra 5 -CCAGGTGCCCTGTCAGTATT-3 5 -CTGGACTTGCTTCCCACACT-3 Il6st 5 -ACCAGATTCCTGTGGACGAC-3 5 -AGAATCCACATGCACAACCA-3 Il1r 5 -TGTCAATGTGACGGACCAGT-3 5 -CACGTAGTTGGAGGGTTCGT-3 Ilrg 5 -TTGCCACCCCAGATCTATTC-3 5 -TTGCCACCCCAGATCTATTC-3 Il1r1 5 -TGAAGAGCACAGAGGGGACT-3 5 -CATTGATCCTGGGTCAGCTT-3 Il1rap 5 -TTGCCACCCCAGATCTATTC-3 5 -TTGCCACCCCAGATCTATTC-3 # The primer set designed to detect mrna for both IκBζ (L) and (S) was used in Figs 1a, f and g and Supplementary Fig. 8a. Supplementary Fig. 8b. The primer set designed to detect mrna for IκBζ (L) and (D) was used in The IκBζ (L)-specific primer set used in Supplementary Fig. 8c.
21 doi: 1.138/nature89 Supplementary Table PCR Primers used for ChIP assays. Position of Gene IκBζ response element # Primer Ila -635 to Forward: 5 -CAGATGCATGCAGAACTGACT-3 (ISE1) Reverse: 5 -AAGGCTCTGGAGAGCAGACA-3 Ila to Forward: 5 -AGTCTGGCCCCTACACACAC-3 (outside CNS) Reverse: 5 -ATGGGGGACTTTTGGGATAG-3 Ilf -154 to -163 Forward: 5 -CACTTCCTGAAGGGGAATCA-3 (promoter) Reverse: 5 -GGGTGGGCTTAGAAGAGAGG-3 Ilf -744 to Forward: 5 -CTGAGTTGGGGGCTGTGTAT-3 (CNS7) Reverse: 5 -CATATCGAGGGTGTCGGACT-3 Il1-31 to -4 Forward: 5 -ACCTTGGTGAATGCTGAAAACTGG-3 Reverse: 5 -TGCGTGTGGGGGCAGGGATGGATA-3 Il3r -13 to -14 Forward: 5 -AACACTGACAAGGCAGCTCA-3 Reverse: 5 -GTCAGCAGAGCCCTGACCTA-3 Il3r to -143 Forward: 5 -CCCTCCCTGCTTGATCACTA-3 Reverse: 5 -TCAGCTTGACACAGGATGGA-3 # The numbering is relative to the transcriptional initiation site. 1
22 doi: 1.138/nature89 Supplementary References 1 Muta, T. IκB-ζ: an inducible regulator of nuclear factor-κb. Vitam. Horm. 74, (6). Watanabe, S., Takeshige, K. & Muta, T. A cis-element in the 3'-untranslated region of IκB-ζ mrna governs its stimulus-specific expression. Biochem Biophys Res Commun 356, (7). 3 Zhou, L. et al. TGF-β-induced Foxp3 inhibits cell differentiation by antagonizing RORγt function. Nature 453, 36-4 (8).
Supplementary. presence of the. (c) mrna expression. Error. in naive or
 Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured
 Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was
 Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Materials for. Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to. FTY720 during neuroinflammation
 Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Nature Medicine: doi: /nm.3922
 Title: Glucocorticoid-induced tumor necrosis factor receptor-related protein co-stimulation facilitates tumor regression by inducing IL-9-producing helper T cells Authors: Il-Kyu Kim, Byung-Seok Kim, Choong-Hyun
Title: Glucocorticoid-induced tumor necrosis factor receptor-related protein co-stimulation facilitates tumor regression by inducing IL-9-producing helper T cells Authors: Il-Kyu Kim, Byung-Seok Kim, Choong-Hyun
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1
 ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF
 CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12
 1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL IL-1 signaling modulates activation of STAT transcription factors to antagonize retinoic acid signaling and control the T H 17 cell it reg cell balance Rajatava Basu 1,5, Sarah K.
SUPPLEMENTARY MATERIAL IL-1 signaling modulates activation of STAT transcription factors to antagonize retinoic acid signaling and control the T H 17 cell it reg cell balance Rajatava Basu 1,5, Sarah K.
D CD8 T cell number (x10 6 )
 IFNγ Supplemental Figure 1. CD T cell number (x1 6 ) 18 15 1 9 6 3 CD CD T cells CD6L C CD5 CD T cells CD6L D CD8 T cell number (x1 6 ) 1 8 6 E CD CD8 T cells CD6L F Log(1)CFU/g Feces 1 8 6 p
IFNγ Supplemental Figure 1. CD T cell number (x1 6 ) 18 15 1 9 6 3 CD CD T cells CD6L C CD5 CD T cells CD6L D CD8 T cell number (x1 6 ) 1 8 6 E CD CD8 T cells CD6L F Log(1)CFU/g Feces 1 8 6 p
Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis. is mediated by SOCS3 regulatory pathway
 Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis is mediated by SOCS3 regulatory pathway Yuan Zhang 1,2, Xing Li 1,2, Bogoljub Ciric 1, Cun-gen Ma 3, Bruno Gran 4, Abdolmohamad
Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis is mediated by SOCS3 regulatory pathway Yuan Zhang 1,2, Xing Li 1,2, Bogoljub Ciric 1, Cun-gen Ma 3, Bruno Gran 4, Abdolmohamad
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Intracellular MHC class II molecules promote TLR-triggered innate. immune responses by maintaining Btk activation
 Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
Akt and mtor pathways differentially regulate the development of natural and inducible. T H 17 cells
 Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
Akt and mtor pathways differentially regulate the development of natural and inducible T H 17 cells Jiyeon S Kim, Tammarah Sklarz, Lauren Banks, Mercy Gohil, Adam T Waickman, Nicolas Skuli, Bryan L Krock,
<10. IL-1β IL-6 TNF + _ TGF-β + IL-23
 3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep
 SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Hasan et al analyse the transcriptional program used by pathogenic Th17 cells raised in the presence of IL-23 as compared to
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Hasan et al analyse the transcriptional program used by pathogenic Th17 cells raised in the presence of IL-23 as compared to
Supplementary Figure 1. Characterization of basophils after reconstitution of SCID mice
 Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1554 a TNF-α + in CD4 + cells [%] 1 GF SPF 6 b IL-1 + in CD4 + cells [%] 5 4 3 2 1 Supplementary Figure 1. Effect of microbiota on cytokine profiles of T cells in GALT. Frequencies of TNF-α
doi:1.138/nature1554 a TNF-α + in CD4 + cells [%] 1 GF SPF 6 b IL-1 + in CD4 + cells [%] 5 4 3 2 1 Supplementary Figure 1. Effect of microbiota on cytokine profiles of T cells in GALT. Frequencies of TNF-α
SUPPLEMENTARY INFORMATION
 Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS Histological analysis. Colonic tissues were collected from 5 parts of the middle colon on day 7 after the start of DSS treatment, and then were cut into segments, fixed with 4% paraformaldehyde,
SUPPLEMENTARY METHODS Histological analysis. Colonic tissues were collected from 5 parts of the middle colon on day 7 after the start of DSS treatment, and then were cut into segments, fixed with 4% paraformaldehyde,
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Eosinophils! 40! 30! 20! 10! 0! NS!
 A Macrophages Lymphocytes Eosinophils Neutrophils Percentage (%) 1 ** 4 * 1 1 MMA SA B C Baseline FEV1, % predicted 15 p = 1.11 X 10-9 5 CD4:CD8 ratio 1 Supplemental Figure 1. Cellular infiltrate in the
A Macrophages Lymphocytes Eosinophils Neutrophils Percentage (%) 1 ** 4 * 1 1 MMA SA B C Baseline FEV1, % predicted 15 p = 1.11 X 10-9 5 CD4:CD8 ratio 1 Supplemental Figure 1. Cellular infiltrate in the
Supplementary Figure 1
 d f a IL7 b IL GATA RORγt h HDM IL IL7 PBS Ilra R7 PBS HDM Ilra R7 HDM Foxp Foxp Ilra R7 HDM HDM Ilra R7 HDM. 9..79. CD + FOXP + T reg cell CD + FOXP T conv cell PBS Ilra R7 PBS HDM Ilra R7 HDM CD + FOXP
d f a IL7 b IL GATA RORγt h HDM IL IL7 PBS Ilra R7 PBS HDM Ilra R7 HDM Foxp Foxp Ilra R7 HDM HDM Ilra R7 HDM. 9..79. CD + FOXP + T reg cell CD + FOXP T conv cell PBS Ilra R7 PBS HDM Ilra R7 HDM CD + FOXP
Supplementary Figure 1. ETBF activate Stat3 in B6 and Min mice colons
 Supplementary Figure 1 ETBF activate Stat3 in B6 and Min mice colons a pstat3 controls Pos Neg ETBF 1 2 3 4 b pstat1 pstat2 pstat3 pstat4 pstat5 pstat6 Actin Figure Legend: (a) ETBF induce predominantly
Supplementary Figure 1 ETBF activate Stat3 in B6 and Min mice colons a pstat3 controls Pos Neg ETBF 1 2 3 4 b pstat1 pstat2 pstat3 pstat4 pstat5 pstat6 Actin Figure Legend: (a) ETBF induce predominantly
NMED-A65251A. Supplementary Figures.
 NMED-A65251A Supplementary Figures. Sup. Fig. 1. ILC3 cells are the main source of in obese mice a. We gated on T cells (upper panels) or T cells (lower panels), and examined production. b. CD45 + - IL-13
NMED-A65251A Supplementary Figures. Sup. Fig. 1. ILC3 cells are the main source of in obese mice a. We gated on T cells (upper panels) or T cells (lower panels), and examined production. b. CD45 + - IL-13
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the
 Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
Supplementary Figure 1. Generation of knockin mice expressing L-selectinN138G. (a) Schematics of the Sellg allele (top), the targeting vector, the targeted allele in ES cells, and the mutant allele in
Nature Medicine doi: /nm.3957
 Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
for six pairs of mice. (b) Representative FACS analysis of absolute number of T cells (CD4 + and
 SUPPLEMENTARY DATA Supplementary Figure 1: Peripheral lymphoid organs of SMAR1 -/- mice have an effector memory phenotype. (a) Lymphocytes collected from MLNs and Peyer s patches (PPs) of WT and SMAR1
SUPPLEMENTARY DATA Supplementary Figure 1: Peripheral lymphoid organs of SMAR1 -/- mice have an effector memory phenotype. (a) Lymphocytes collected from MLNs and Peyer s patches (PPs) of WT and SMAR1
CD4 + T cells recovered in Rag2 / recipient ( 10 5 ) Heart Lung Pancreas
 a CD4 + T cells recovered in Rag2 / recipient ( 1 5 ) Heart Lung Pancreas.5 1 2 4 6 2 4 6 Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas b Heart Lung Pancreas Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas
a CD4 + T cells recovered in Rag2 / recipient ( 1 5 ) Heart Lung Pancreas.5 1 2 4 6 2 4 6 Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas b Heart Lung Pancreas Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Nature Immunology: doi: /ni Supplementary Figure 1. Cellularity of leukocytes and their progenitors in naive wild-type and Spp1 / mice.
 Supplementary Figure 1 Cellularity of leukocytes and their progenitors in naive wild-type and Spp1 / mice. (a, b) Gating strategies for differentiated cells including PMN (CD11b + Ly6G hi and CD11b + Ly6G
Supplementary Figure 1 Cellularity of leukocytes and their progenitors in naive wild-type and Spp1 / mice. (a, b) Gating strategies for differentiated cells including PMN (CD11b + Ly6G hi and CD11b + Ly6G
Transcription factor Foxp3 and its protein partners form a complex regulatory network
 Supplementary figures Resource Paper Transcription factor Foxp3 and its protein partners form a complex regulatory network Dipayan Rudra 1, Paul deroos 1, Ashutosh Chaudhry 1, Rachel Niec 1, Aaron Arvey
Supplementary figures Resource Paper Transcription factor Foxp3 and its protein partners form a complex regulatory network Dipayan Rudra 1, Paul deroos 1, Ashutosh Chaudhry 1, Rachel Niec 1, Aaron Arvey
Supplementary Figures. T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ
 Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplemental Information. T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism
 Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
Supplementary Figure 1. Deletion of Smad3 prevents B16F10 melanoma invasion and metastasis in a mouse s.c. tumor model.
 A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
Peli1 negatively regulates T-cell activation and prevents autoimmunity
 Peli1 negatively regulates T-cell activation and prevents autoimmunity Mikyoung Chang 1,*, Wei Jin 1,5,*, Jae-Hoon Chang 1, Yi-chuan Xiao 1, George Brittain 1, Jiayi Yu 1, Xiaofei Zhou 1, Yi-Hong Wang
Peli1 negatively regulates T-cell activation and prevents autoimmunity Mikyoung Chang 1,*, Wei Jin 1,5,*, Jae-Hoon Chang 1, Yi-chuan Xiao 1, George Brittain 1, Jiayi Yu 1, Xiaofei Zhou 1, Yi-Hong Wang
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs
 Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author):
 Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Supplementary Figure 1.
 Supplementary Figure 1. Female Pro-ins2 -/- mice at 5-6 weeks of age were either inoculated i.p. with a single dose of CVB4 (1x10 5 PFU/mouse) or PBS and treated with αgalcer or control vehicle. On day
Supplementary Figure 1. Female Pro-ins2 -/- mice at 5-6 weeks of age were either inoculated i.p. with a single dose of CVB4 (1x10 5 PFU/mouse) or PBS and treated with αgalcer or control vehicle. On day
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplemental Table I.
 Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
Supporting Information
 Supporting Information Stegbauer et al. 10.1073/pnas.0903602106 SI Methods Analysis of Plasma Renin Activity (PRA) and ACE Activity. PRA and serum ACE activity levels were determined by RIA (RENCTK, DiaSorin;
Supporting Information Stegbauer et al. 10.1073/pnas.0903602106 SI Methods Analysis of Plasma Renin Activity (PRA) and ACE Activity. PRA and serum ACE activity levels were determined by RIA (RENCTK, DiaSorin;
Supplementary Information:
 Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
Kerdiles et al - Figure S1
 Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Kerdiles et al - Figure S1 a b Homo sapiens T B ce ce l ls c l M ls ac r PM oph N ag es Mus musculus Foxo1 PLCγ Supplementary Figure 1 Foxo1 expression pattern is conserved between mouse and human. (a)
Immune response to infection
 Immune response to infection Dr. Sandra Nitsche (Sandra.Nitsche@rub.de ) 20.06.2018 1 Course of acute infection Typical acute infection that is cleared by an adaptive immune reaction 1. invasion of pathogen
Immune response to infection Dr. Sandra Nitsche (Sandra.Nitsche@rub.de ) 20.06.2018 1 Course of acute infection Typical acute infection that is cleared by an adaptive immune reaction 1. invasion of pathogen
% of live splenocytes. STAT5 deletion. (open shapes) % ROSA + % floxed
 Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Tbk1-TKO! DN cells (%)! 15! 10!
 a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the
 Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Bezzi et al., Supplementary Figure 1 *** Nature Medicine: doi: /nm Pten pc-/- ;Zbtb7a pc-/- Pten pc-/- ;Pml pc-/- Pten pc-/- ;Trp53 pc-/-
 Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c)
 1 Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c) Serum IgG1 (a), IgM (b) and IgG2 (c) concentrations in response to papain immediately before primary immunization (day
1 Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c) Serum IgG1 (a), IgM (b) and IgG2 (c) concentrations in response to papain immediately before primary immunization (day
Effector T Cells and
 1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
Eosinophils are required. for the maintenance of plasma cells in the bone marrow
 Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Supplementary Figure 1. Antibiotic partially rescues mice from sepsis. (ab) BALB/c mice under CLP were treated with antibiotic or PBS.
 1 Supplementary Figure 1. Antibiotic partially rescues mice from sepsis. (ab) BALB/c mice under CLP were treated with antibiotic or PBS. (a) Survival curves. WT Sham (n=5), WT CLP or WT CLP antibiotic
1 Supplementary Figure 1. Antibiotic partially rescues mice from sepsis. (ab) BALB/c mice under CLP were treated with antibiotic or PBS. (a) Survival curves. WT Sham (n=5), WT CLP or WT CLP antibiotic
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
Supplementary Table 1. List of primers used in this study
 Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
MicroRNA mir-326 regulates T H -17 differentiation and is associated with the pathogenesis of multiple sclerosis
 correction notice Nat. Immunol. 10, 1252 1259 (2009) MicroRNA mir-326 regulates T H -17 differentiation and is associated with the pathogenesis of multiple sclerosis Changsheng Du, Chang Liu, Jiuhong Kang,
correction notice Nat. Immunol. 10, 1252 1259 (2009) MicroRNA mir-326 regulates T H -17 differentiation and is associated with the pathogenesis of multiple sclerosis Changsheng Du, Chang Liu, Jiuhong Kang,
SUPPLEMENTARY FIGURES AND TABLE
 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
Supplementary Information. Tissue-wide immunity against Leishmania. through collective production of nitric oxide
 Supplementary Information Tissue-wide immunity against Leishmania through collective production of nitric oxide Romain Olekhnovitch, Bernhard Ryffel, Andreas J. Müller and Philippe Bousso Supplementary
Supplementary Information Tissue-wide immunity against Leishmania through collective production of nitric oxide Romain Olekhnovitch, Bernhard Ryffel, Andreas J. Müller and Philippe Bousso Supplementary
W/T Itgam -/- F4/80 CD115. F4/80 hi CD115 + F4/80 + CD115 +
 F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Supplemental Materials
 Supplemental Materials Programmed death one homolog maintains the pool size of regulatory T cells by promoting their differentiation and stability Qi Wang 1, Jianwei He 1, Dallas B. Flies 2, Liqun Luo
Supplemental Materials Programmed death one homolog maintains the pool size of regulatory T cells by promoting their differentiation and stability Qi Wang 1, Jianwei He 1, Dallas B. Flies 2, Liqun Luo
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study.
 Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
Supplementary Table 1. Oligonucleotide primers used in quantitative and qualitative PCRs of this study. Primer Sequence Qualitative PCR primers Universal 16S rdna forward 5 - GACGCTGAGGCATGAGAGCA-3 Universal
Supplementary Figure 1. NAFL enhanced immunity of other vaccines (a) An over-the-counter, hand-held non-ablative fractional laser (NAFL).
 Supplementary Figure 1. NAFL enhanced immunity of other vaccines (a) An over-the-counter, hand-held non-ablative fractional laser (NAFL). (b) Depiction of a MTZ array generated by NAFL. (c-e) IgG production
Supplementary Figure 1. NAFL enhanced immunity of other vaccines (a) An over-the-counter, hand-held non-ablative fractional laser (NAFL). (b) Depiction of a MTZ array generated by NAFL. (c-e) IgG production
RORγt and IL-17 Responses`
 Falk Workshop Mechanisms of Intestinal Inflammation October 10, 2007 and IL-17 Responses` Dan Littman HHMI, Skirball Institute New York University School of Medicine New paradigm for T Helper Cell Differentiation
Falk Workshop Mechanisms of Intestinal Inflammation October 10, 2007 and IL-17 Responses` Dan Littman HHMI, Skirball Institute New York University School of Medicine New paradigm for T Helper Cell Differentiation
Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x
 Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x magnification). B: Second HC skin stained for TSLP receptor in brown
Supplementary Figure 1: TSLP receptor skin expression in dcssc. A: Healthy control (HC) skin with TSLP receptor expression in brown (10x magnification). B: Second HC skin stained for TSLP receptor in brown
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created.
 938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
Nature Immunology: doi: /ni.3866
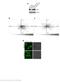 Nature Immunology: doi:10.1038/ni.3866 Supplementary Figure 1 The effect of TIPE2 on chemotaxis. a, The expression of TIPE2 in dhl-60c, dhl-60t, TIPE2-expressing and 15/16Q-expressing dhl-60t neutrophils
Nature Immunology: doi:10.1038/ni.3866 Supplementary Figure 1 The effect of TIPE2 on chemotaxis. a, The expression of TIPE2 in dhl-60c, dhl-60t, TIPE2-expressing and 15/16Q-expressing dhl-60t neutrophils
Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression.
 Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression. (a) Characterization of c-independent SP8 cells. Stainings for maturation markers (top)
Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression. (a) Characterization of c-independent SP8 cells. Stainings for maturation markers (top)
Simultaneous correlation of cytokine production with Treg and Th17 cell proliferation
 Simultaneous correlation of cytokine production with Treg and Th17 cell proliferation Jurg Rohrer, PhD Director, R&D BD Biosciences 23-11773-00 For Research Use Only. Not for use in diagnostic or therapeutic
Simultaneous correlation of cytokine production with Treg and Th17 cell proliferation Jurg Rohrer, PhD Director, R&D BD Biosciences 23-11773-00 For Research Use Only. Not for use in diagnostic or therapeutic
pplementary Figur Supplementary Figure 1. a.
 pplementary Figur Supplementary Figure 1. a. Quantification by RT-qPCR of YFV-17D and YFV-17D pol- (+) RNA in the supernatant of cultured Huh7.5 cells following viral RNA electroporation of respective
pplementary Figur Supplementary Figure 1. a. Quantification by RT-qPCR of YFV-17D and YFV-17D pol- (+) RNA in the supernatant of cultured Huh7.5 cells following viral RNA electroporation of respective
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice
 Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
D2 inhibits TLR2- initiated 12p40 transcription (-) TLR2 PGN MDP. MyD88 IRAK ECSIT TRAF6 NIK. Smallest unit of PGN muramyl dipeptide IKK.
 D2 inhibits TLR2- initiated 12p40 transcription CARD CARD NOD2 LRR RICK/Rip2 NIK MDP TRAF6 PGN TLR2 MyD88 IRAK ECSIT (-) IKK Smallest unit of PGN muramyl dipeptide IκB NF-κB atanabe et al, 2004 NF-κB IL-12p40
D2 inhibits TLR2- initiated 12p40 transcription CARD CARD NOD2 LRR RICK/Rip2 NIK MDP TRAF6 PGN TLR2 MyD88 IRAK ECSIT (-) IKK Smallest unit of PGN muramyl dipeptide IκB NF-κB atanabe et al, 2004 NF-κB IL-12p40
Supplementary Data. (continued)
 Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature775 4 O.D. (595-655) 3 1 -ζ no antibody isotype ctrl Plated Soluble 1F6 397 7H11 Supplementary Figure 1 Soluble and plated anti- Abs induce -! signalling. B3Z cells stably expressing!
doi: 1.138/nature775 4 O.D. (595-655) 3 1 -ζ no antibody isotype ctrl Plated Soluble 1F6 397 7H11 Supplementary Figure 1 Soluble and plated anti- Abs induce -! signalling. B3Z cells stably expressing!
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Predictive PP1Ca binding region in BIG3 : 1,228 1,232aa (-KAVSF-) HEK293T cells *** *** *** KPL-3C cells - E E2 treatment time (h)
 Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel
 Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1%
 FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
Transcription Factor IRF8 Directs a Silencing Programme for TH17 Cell Differentiation
 Transcription Factor Directs a Silencing Programme for TH7 Cell Differentiation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
Transcription Factor Directs a Silencing Programme for TH7 Cell Differentiation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
Nature Immunology: doi: /ni Supplementary Figure 1. Cytokine pattern in skin in response to urushiol.
 Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder I Nakaya, Kousik
 SUPPLEMENTARY FIGURES 1-19 T H 2 response to cysteine-proteases requires dendritic cell-basophil cooperation via ROS mediated signaling Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder
SUPPLEMENTARY FIGURES 1-19 T H 2 response to cysteine-proteases requires dendritic cell-basophil cooperation via ROS mediated signaling Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic
 Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
