Identification and characterization of a novel splice variant of the PLCζ1 gene in bull testis tissues
|
|
|
- Byron Hunt
- 5 years ago
- Views:
Transcription
1 Identification and characterization of a novel splice variant of the PLCζ1 gene in bull testis tissues Z.H. Ju 1 *, Q. Pan 1,2 *, Y. Zhang 1 *, J.M. Huang 1, C. Qi 1, X.G. Wang 1, Q.L. Li 1, J.F. Zhong 1, M. Liu 2 and C.F. Wang 1 1 Dairy Cattle Research Center, Shandong Academy of Agricultural Science, Jinan, China 2 College of Life Science, Nanjing Normal University, Nanjing, China *These authors contributed equally to this study. Corresponding author: C.F. Wang wangcf1967@163.com Genet. Mol. Res. 13 (4): (2014) Received October 28, 2013 Accepted January 24, 2014 Published November 27, 2014 DOI ABSTRACT. Phospholipase C zeta 1 (PLCζ1), which transcribes a key protein, has an important function in oocyte activation and embryo development because PLCζ1 can trigger a series of intracellular Ca 2+ oscillations in mammals. In this study, a novel splice variant in the testis tissues of adult and fetal Chinese Holstein bulls was characterized by reverse transcription-polymerase chain reaction (RT-PCR) and sequencing analysis. The novel splice variant PLCζ1-sv1 was derived from the PLCζ1 complete transcript (PLCζ1-complete) by alternative splicing; the alternative splicing pattern exhibited alternative 5'-splice sites. The full-length transcript, PLCζ1-complete, is the main transcript found in fetal and adult cow testis tissue. Quantitative real-time PCR (qpcr) analysis demonstrated that the expression levels of the PLCζ1- complete transcript were significantly higher than those of the PLCζ1- sv1 splice variant in bovine testis tissues. PLCζ1 protein sequencing analysis showed that the amino acids at positions 453 to 457 were
2 Z.H. Ju et al deleted in PLCζ1-sv1, thereby terminating transcription prematurely. In summary, this study provided information to elucidate the structure and function of the bovine PLCζ1 gene. Key words: PLCζ1 gene; Chinese Holstein bull; Splice variant; Quantitative real-time polymerase chain reaction; Protein sequencing analysis INTRODUCTION Bovine sperm-specific phospholipase C zeta 1 (PLCζ1), mapped on SSC5q11-12, comprises 14 exons and encodes a protein with 634 amino acids, which is a key protein during sperm-egg fusion. The PLCζ1 gene sequence have been cloned in bovine, mouse, human, monkey, chicken, pig, medaka fish, rat, and hamster (Cox et al., 2002; Saunders et al., 2002; Coward et al., 2005; Yoneda et al., 2006; Ito et al., 2008; Young et al., 2009). PLCζ1 contains X and Y catalytic domains that are common to all phosphoinositide-specific PLCs, two pairs of EF hand domains that are possibly involved in Ca 2+ binding, a C2 domain, which appears to bind to phosphoinositide-containing lipids (Katan, 1998; Rebecchi and Pentyala, 2000), and a putative nuclear localization signal in the linker region between the X and Y catalytic domains. Similar to other PLC family members (Rebecchi and Pentyala, 2000), the PLCζ1 protein catalyzes the hydrolysis of phosphatidylinositol 4,5-bisphosphate and the production of 1,4,5-inositol trisphosphate; diacylglycerol (Kouchi et al., 2005; Nomikos et al., 2005) is necessary to initiate and maintain Ca 2+ oscillations, which are also observed during fertilization in normal embryo development (Brind et al., 2000; Jellerette et al., 2000; Wu et al., 2001). Ca 2+ oscillations are observed in experiments in which PLCζ1 is introduced to somatic cells by microinjection (Berrie et al., 1996) and to cell-free systems, such as sea urchin egg homogenates (Jones et al., 1998). Despite the seemingly conserved nature of PLCζ1 among different species, small sequence differences are sufficient to affect structural activity significantly in a species-specific manner (Cox et al., 2002; Ross et al., 2008). Considering these results, we focused on domain regions that determine the specificity of the catalytic activity of bovine PLCζ1. Alternative splicing has a very important function in protein diversity; therefore, alternative splicing may result in discrepancy between gene expression and individual complexity (Garcia-Blanco et al., 2004; Garcia-Blanco, 2006). This phenomenon is universal to numerous protein-coding genes of multicellular organisms. Several studies have indicated that alternative splicing should be determined (Ast, 2004; Chacko and Ranganathan, 2009). In general, alternatively spliced exons have 5'- and 3'-splice site motifs that differ significantly from the normal motifs. In addition to these splice sites, exons are defined by cis-acting regulatory elements, which have been divided into four functional categories: exonic splicing enhancers, exonic splicing silencers, intronic splicing enhancers (also known as intronic activators of splicing), and intronic splicing silencers. These cis-acting elements interact directly or indirectly with trans-acting activators or repressors of splicing. In mice, a novel alternative splicing variant of the PLCζ1 gene that contains a 30-bp insertion following base 159 of the wild-type open reading frame is present in the nucleotide sequence database (accession No. AK006672), which results in the expression
3 Splice variant of the PLCz1 gene in bull testis tissues 9901 of a different protein (Kouchi et al., 2005; Nomikos et al., 2005). During the cloning of the human and rat PLCζ1 from testis, Saunders et al. (2002) observed several other incompletely spliced cdnas that contained internal stop codons. However, the splice variants of bovine PLCζ1 and PLCζ1 mrna expression in bulls have not been characterized yet. In this study, PLCζ1 splice variants in testis tissues of bulls and the differential expression pattern in adult and fetal cows were identified. MATERIAL AND METHODS Tissue sample collection Testis tissue samples were aseptically collected from eight Chinese Holstein bulls (four adult bulls were three to six years old, and four fetal bulls were newly born). The tissues were snap frozen in liquid nitrogen in a slaughterhouse and transported to our laboratory. Isolation of total RNA and cdna synthesis Total RNA was isolated from 0.1 to 0.2 g testis tissue using TRIzol reagent (Tiangen, China) according to manufacturer instructions. RNA was digested by RNase-free DNase (Takara, Japan) to remove genomic DNA contamination. The RNA concentration was measured using a Biophotometer (Implen, Germany). RNA integrity was determined by electrophoresis on a 1% agarose gel. The RevertAid TM First-Strand cdna Synthesis kit (Fermentas, Canada) was used to convert approximately 1 µg RNA from each sample to cdna according to manufacturer specifications. Identification of PLCζ1 gene splice variants Primers P A-C (Table 1) were designed and used to amplify the coding region of the bovine PLCζ1 gene according to the reference sequence (GenBank accession No. NM_ ). Polymerase chain reaction (PCR) products were purified and subcloned in a peasy-t3 cloning vector, which was transformed into Escherichia coli DH5α competent cells. The cells were subsequently propagated in Luria broth medium overnight at 37 C. The plasmids were then extracted using a Plasmid Miniprep kit (Biomiga), and the cdna segment inserts were sequenced using an ABI PRISM 3730 DNA sequencer (Applied Biosystems) and BigDye Terminator v3.1 Sequencing kit (Shanghai Sangon, China). Multiple sequence alignments were performed using the DNAMAN v5.2.2 software (LynonBiosoft) to identify possible splice variants. Protein structure predictions of different PLCζ1 splice variants The secondary protein structure of bovine PLCζ1 splice variants was predicted according to the three-dimensional protein structure was predicted according to Alterations in the binding site of the splicing factor were predicted using ESEfinder 3.0 ( esefinder.cgi).
4 Z.H. Ju et al Table 1. Primers used in this study. Primers Primer sequences (5' 3') Annealing Fragment Target temperatures ( C) size (bp) P A AGTGGCAACAGCGGACGA PLCζ1 gene cdna CCCCAAATGTAGCGAGCAAA amplification (position 136 to 1078) P B CGATACATTCACCAGCAAG PLCζ1 gene cdna CGAAGAGCCATAGGGACA amplification (position 925 to 1692) P C GAAACGGGTGGGCGGAAT PLCζ1 gene cdna AGGGAAGCGGGCTCAAGAC amplification (position 1338 to 2147) PLCζ1-complete CAGGAGATTCATTACCAGAG PLCζ1 gene qrt-pcr AAGTTCCATAGGGACACC PLCζ1-sv1 TGTAGGTTGTCAGATGGTGG PLCζ1 gene qrt-pcr CTCGTAGGAAGCGTGGTT β-actin TTAGCTGCGTTACACCCTT β-actin gene qrt-pcr TGTCACCTTCACCGTTCC Nucleotide sequences are numbered relative to the translational start site. Quantitative real-time PCR analysis of PLCζ1 splice variants To determine the relative expression levels of the PLCζ1-complete and PLCζ1-sv1 transcripts in bovine testis, we performed quantitative real-time PCR (q-pcr) using SYBR Green PCR Master Mix (TaKaRa, Dalian, China). The q-pcr mixture with a final volume of 20 µl consisted of the following components: 10.0 µl 2X SYBR Pre-mix Ex Taq TM, 50 ng cdna (1.5 µl), 0.5 µl 10 µm sense and antisense primers, and 7.5 µl ddh 2 O. After initial denaturation at 95 C for 30 s, the following incubation protocol was used: 40 cycles of denaturation at 95 C for 5 s and annealing at 60 C for 30 s. PCR was carried out using a Light- Cycler 480 II Real-time PCR system. Each sample was run in triplicate. The q-pcr primers of the PLCζ1 gene and housekeeping internal control gene (β-actin gene; GenBank accession No. BT ) are listed in Table 1. The relative expression was determined as described by Ju et al. (2012). Statistical analysis PLCζ1 mrna expression was quantified relative to a housekeeping gene, and the results are reported as fold-change. Analysis was performed using SPSS v.10.0 (SPSS Inc.). The means of the two groups were compared using a paired-sample Student t-test. P < 0.05 was considered to be significant. RESULTS AND DISCUSSION Identification of a novel PLCζ1 gene splice variant in bull testis tissues mrna splicing occurs during as post-transcriptional step, in which certain segments of pre-mrnas are spliced, and the remaining exons are arranged to generate a mature mrna product. As a result, the same genetic locus can produce various transcripts, thereby producing hundreds, thousands, or even tens of thousands of protein isoforms (Graveley, 2001). The number of proteins expressed by an organism can considerably exceed the number of genes contained in a genome. Among the bovine genes, 21% undergo alternative splicing events. In this study, a novel splice variant of the PLCζ1 gene was found in bull testis tissues.
5 Splice variant of the PLCz1 gene in bull testis tissues 9903 The full-length PLCζ1 transcript (including three fragments) was amplified using cdna from the testis tissues as a template. To further identify the sequence, we used purified PCR products for subcloning and sequencing. We compared the sequence obtained from each sample with the bovine PLCζ1 genomic sequence (GenBank accession No. NC_ ) and with the PLCζ1 mrna sequence (GenBank accession No. NM_ ). We identified a novel transcript, which we named PLCζ1-sv1 (splice variant; Figure 1). The complete transcript was designated as PLCζ1-complete. On the basis of isoform frequencies, PLCζ1- complete was considered to be the major transcript comprising 90% of 50 sequenced colonies. In contrast, five other transcripts were identified as PLCζ1-sv1 with a frequency of 10% (5/50). The PLCζ1-sv1 sequence was submitted to the National Center for Biotechnology Information (GenBank accession No. JX442544). Figure 1. Diagrammatic maps of bovine PLCζ1 genomic structure and alternative splicing patterns of PLCζ1 transcripts. Boxes in different colors represent the exons; yellow boxes represent the 5'-untranslated regions (UTRs), including exon 1 and part of exon 2; blue boxes represent the 3'-UTRs containing part of exon 14; and red boxes represent the coding region. The alternatively spliced region is shown as a white box. Transcription pattern of the novel splice variant In this study, sequence alignment demonstrated that the splice variant PLCζ1-sv1 was truncated at the 3'-end of exon 10 and lacked 16 nucleotides (Figure 1). The alternative splicing pattern of PLCζ1-sv1 is found in the alternative 5'-splice site (Keren et al., 2010). Sixteen missense nucleotide deletions correspond to the deleted amino acids located in the primary structure (amino acids at positions 453 to 457); as a result, a transcription termination codon is encountered at an early stage as revealed by bioinformatic analysis. PLCζ1-complete encodes a complete protein containing 634 amino acids, whereas PLCζ1-sv1 putatively encodes 454 amino acids and lacks 180 amino acids (Figure 2). The deleted section is located in a highly conserved catalytic structure of the Y domain.
6 Z.H. Ju et al Figure 2. Predicted amino acid sequences of PLCζ1-complete and PLCζ1-sv1. The X catalytic domain is represented by a continuous red underline. The X-Y linker domain is represented by a boxed region in red. The Y catalytic domain is represented by a red broken underline. The EF hand and C2 domains correspond to unmarked N- and C-terminal regions, respectively. Difference between the predicted protein structures of PLCζ1-complete and PLCζ1-sv1 The results demonstrated that bovine PLCζ1-complete was a mixed-type protein, in which α-helices, β-pleated sheets, and random coils account for approximately 29.3, 16.1, and 54.6% of the protein, respectively. Hydrophilic and hydrophobic regions were also widely distributed in the secondary structure. However, the protein structure of PLCζ1-sv1 was composed of the following: 38.1% α-helices, 8.8% β-pleated sheets, and 53.1% random coils. These results showed that the 16 missense mutations in PLCζ1-sv1 altered the secondary protein structure. The domain, secondary structures, and three-dimensional structures of bovine PLCζ1-complete and PLCζ1-sv1 proteins were predicted by SWISS-MODEL (Figures 3 and 4). PLCζ1-complete contains six domains: calcium-binding EF-hand domain (PS50222: 35-70); phosphatidylinositol-specific phospholipase C, X region domain (PF00388: ; PS50007: ); phosphatidylinositol-specific phospholipase C, Y region domain (PF00387: ; PS50008: ); C2 calcium-dependent membrane-targeting domain (PF00168: ; PS50004: ); C2 calcium/lipid-binding region, CaLB domain (SSF49562: ); and phosphatidylinositol-specific phospholipase C, X and Y box domain (PD001202: ). However, the regions encoding the C2 calcium-dependent membrane-targeting domain and the C2 calcium/lipid-binding region (CaLB domain) were completely spliced out of the PLCζ1-sv1 transcript (Figures 3a and 4a). In addition to the differences in the secondary structures of PLCζ1-complete and PLCζ1-sv1 (Figures 3b and 4b), differences in the three-dimensional structures were predicted. The QMEAN-scores of PLCζ1-complete and PLCζ1-sv1 were (mean Z-score: -4.98) and (mean Z-score: -4.15), respectively (Figures 3c and 4c).
7 Splice variant of the PLCz1 gene in bull testis tissues 9905 These results demonstrated that the structure of the product encoded by PLCζ1-sv transcripts underwent significant changes, thereby forming unusual protein conformations. Two forms of bovine PLCζ1 have been identified; such forms differ from those showing a conservative deletion in the C2 domain. Although the deleted C2 domain exhibits enzymatic activity, this domain fails to cause Ca 2+ oscillations, possibly because of the loss of a lipid-targeting domain, which is required to direct PLCζ1 to its substrate (Saunders et al., 2007). Deregulated splice variant expression has been identified as the cause of numerous genetic disorders; certain forms of cancer have been linked to an unbalanced isoform expression of genes involved in cell cycle regulation or apoptosis (Faustino and Cooper, 2003; Giesecke et al., 2010). Figure 3. Putative protein structure and functional domains of bovine PLCζ1-complete with the QMEAN results. (a) Putative functional domains of bovine PLCζ1-complete. (b) Predicted protein structure of bovine PLCζ1-complete. (c) The total QMEAN-score of PLCζ1-complete is (mean Ζ-score: -4.98; estimated model reliability between 0 and 1). The red x indicates the differences in the three-dimensional structures of PLCζ1-complete and PLCζ1-sv1. Figure 4. Putative protein structure and functional domains of bovine PLCζ1-sv1 with the QMEAN results. (a) Putative functional domains of bovine PLCζ1-sv1. (b) Predicted protein structure of bovine PLCζ1-sv1. (c) The total QMEAN-score of PLCζ1-sv1 is (mean Z-score: -4.15; estimated model reliability between 0 and 1). The red x indicates the differences in the three-dimensional structures of PLCζ1-sv1 and PLCζ1-complete.
8 Z.H. Ju et al Expression of the two transcripts in testis tissues of Chinese Holstein bulls Northern and Western blot analyses confirmed that PLCζ1 is expressed mainly in the testis (Saunders et al., 2002). In this study, the samples were collected from the testis tissue of fetal and adult bovine to investigate the expression levels of the two PLCζ1 transcripts. Differential expression of PLCζ1-complete and PLCζ1-sv1 mrnas in these adult and fetal cow testis tissues was investigated by q-pcr using the specific primers shown in Table 1. Differential expression of PLCζ1-complete transcript between fetal and adult tissues is shown in Figure 5. PLCζ1-complete was abundantly expressed in adult testis but was marginally detected in fetal testis (P < 0.05). However, no significant differences in PLCζ1- sv1 mrna expression were observed in adult and fetal bulls. The expression level of the PLCζ1-complete transcript was significantly higher than that of the PLCζ1-sv1 transcript in adult bull testis (P < 0.05). The PLCζ1-complete transcript is dominantly expressed by the PLCζ1 gene. Although the PLCζ1-sv1 transcript, which was generated by RNA splicing, was found in bovine testis, defective pre-mrna splicing may generate non-functional and toxic proteins with formidable effects on cell homeostasis by removing certain parts of exons. Aberrant RNA splicing is associated with many diseases (Wang et al., 2003; Nagao et al., 2005). Therefore, studies should be conducted to investigate the association between a novel PLCζ1 transcript and bovine reproduction-related diseases. Figure 5. Relative expression of the two transcripts of the PLCζ1 gene obtained by quantitative real-time polymerase chain reaction (PCR) of bovine testis. Blue represents the relative expression of the bovine PLCζ1- complete transcript in adult and fetal bovine testis tissues; purple represents the relative expression of the bovine PLCζ1-sv1 transcript in adult and fetal bovine testis tissues. The vertical bars represent standard error. Exon splicing enhancer prediction of the two transcripts The mechanism by which the novel splice variant PLCζ1-sv1 exists is unknown. Splicing signals, such as the 5'-splice site, 3'-splice site, branch point, and polypyrimidine
9 Splice variant of the PLCz1 gene in bull testis tissues 9907 tract site, are required for pre-mrna splicing (Lee et al., 2012). Additional RNA sequence elements, such as exon splicing enhancers, are also important to initiate the pre-mrna splicing of many genes. Exon splicing enhancers activate exons and promote their inclusion in mature transcripts. Several different exon splicing enhancer families have been recognized; among these families, purine-rich sequences that function by recruiting members of the serine/arginine-rich family of splicing factors (e.g., ASF/SF2) are widely studied. Exon splicing enhancers have been implicated in the regulation of alternative splicing and possibly define many constitutive exons. Using ESEfinder 3.0, we found that alternatively spliced exons contain RNA sequences that are important for pre-mrna splicing; these sequences are similar to exons that undergo regular splicing (Figure 6). The PLCζ1-sv1 transcript did not change the binding sites of SRSF1, SRSF2 (IgM-BRCA1), SRSF3, SRSF4, and SRSF6. However, the PLCζ1-sv1 transcript did not contain potential binding sites for the splicing factor SRSF5, which differed from the PLCζ1-complete transcript in terms of auxiliary splicing proteins (Figure 6). In addition to the essential splicing signals on the pre-mrna, enhancers or inhibitors/silencers that are found on pre-mrna are particularly important to regulate alternative splicing. Figure 6. Potential exonic splice enhancer motif threshold scores of the two transcripts of the PLCζ1 gene. The PLCζ1-complete sequence: ccagggaatttattcttcacaccaggaggttcattaccagagtatatcccaaagcattaagagctgactcttctaatttt aatcctcaagaattttggaatgtaggttgtcaaatgg. The PLCζ1-sv1 sequence: ccagggaatttattcttcacaccaggaggttcattaccagagtatatc ccaaagcattaagagctgactcttctaattttaatcctcaagaattttggaat. (A) Red, purple, blue, green, and yellow represent the exonic splice enhancers of PLCζ1, including SRSF1, SRSF2, SRSF3, SRSF4, and SPSF5, respectively. (B) The novel PLCζ1-sv1 transcript contains an additional SRSF5 site (TCAAATC).
10 Z.H. Ju et al A novel splice variant, PLCζ1-sv1, was identified in bovine testis tissues. The fulllength PLCζ1-complete transcript is the main transcript present in fetal and adult bull testis tissues. PLCζ1-complete transcript expression was significantly higher than that of the PLCζ1-sv1 transcript. The mrna expression levels of the PLCζ1-complete transcript in adult bull testis tissues were higher than those in fetal bull testis tissues. The expression levels of the splice variant PLCζ1-sv1 in adult and fetal bull testis tissues did not significantly differ. However, the predicted three-dimensional structures of the products of the two transcripts showed that the PLCζ1-sv1 structure varied from that of PLCζ1-complete. Therefore, PLCζ1- complete may be used as a suitable transcript to initiate responses to the environment and bovine reproduction. This study may facilitate a better understanding of the structure and function of the PLCζ1 gene. Further studies should be conducted to determine the reason why novel splice variants of PLCζ1 are present in bovine. ACKNOWLEDGMENTS Research supported by the Program of the Ministry of Science and Technology, China (Grants #2011BAD19B04, #2011BAD19B02, and #2011BAD28B02), the Modern Agro-Industry Technology Research System (Grant #CARS-37), the National Natural Science Foundation of China (Grants # , # , and # ), and the Shandong Province Science & Technology Development Program (Grant #2013GNC11023). REFERENCES Ast G (2004). How did alternative splicing evolve? Nat. Rev. Genet. 5: Berrie CP, Cuthbertson KS, Parrington J, Lai FA, et al. (1996). A cytosolic sperm factor triggers calcium oscillations in rat hepatocytes. Biochem. J. 313 (Pt 2): Brind S, Swann K and Carroll J (2000). Inositol 1,4,5-trisphosphate receptors are downregulated in mouse oocytes in response to sperm or adenophostin A but not to increases in intracellular Ca 2+ or egg activation. Dev. Biol. 223: Chacko E and Ranganathan S (2009). Genome-wide analysis of alternative splicing in cow: implications in bovine as a model for human diseases. BMC Genomics 10 (Suppl 3): S11. Coward K, Ponting CP, Chang HY, Hibbitt O, et al. (2005). Phospholipase Czeta, the trigger of egg activation in mammals, is present in a non-mammalian species. Reproduction 130: Cox LJ, Larman MG, Saunders CM, Hashimoto K, et al. (2002). Sperm phospholipase Czeta from humans and cynomolgus monkeys triggers Ca 2+ oscillations, activation and development of mouse oocytes. Reproduction 124: Faustino NA and Cooper TA (2003). Pre-mRNA splicing and human disease. Genes Dev. 17: Garcia-Blanco MA (2006). Alternative splicing: therapeutic target and tool. Prog. Mol. Subcell. Biol. 44: Garcia-Blanco MA, Baraniak AP and Lasda EL (2004). Alternative splicing in disease and therapy. Nat. Biotechnol. 22: Giesecke K, Sieme H and Distl O (2010). Infertility and candidate gene markers for fertility in stallions: a review. Vet. J. 185: Graveley BR (2001). Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17: Ito M, Shikano T, Oda S, Horiguchi T, et al. (2008). Difference in Ca 2+ oscillation-inducing activity and nuclear translocation ability of PLCZ1, an egg-activating sperm factor candidate, between mouse, rat, human, and medaka fish. Biol. Reprod. 78: Jellerette T, He CL, Wu H, Parys JB, et al. (2000). Down-regulation of the inositol 1,4,5-trisphosphate receptor in mouse eggs following fertilization or parthenogenetic activation. Dev. Biol. 223: Jones KT, Cruttwell C, Parrington J and Swann K (1998). A mammalian sperm cytosolic phospholipase C activity generates inositol trisphosphate and causes Ca 2+ release in sea urchin egg homogenates. FEBS Lett. 437: Ju Z, Wang C, Li Q, Hou M, et al. (2012). Alternative splicing and mrna expression analysis of bovine SLAMF7 gene
11 Splice variant of the PLCz1 gene in bull testis tissues 9909 in healthy and mastitis mammary tissues. Mol. Biol. Rep. 39: Katan M (1998). Families of phosphoinositide-specific phospholipase C: structure and function. Biochim. Biophys. Acta 1436: Keren H, Lev-Maor G and Ast G (2010). Alternative splicing and evolution: diversification, exon definition and function. Nat. Rev. Genet. 11: Kouchi Z, Shikano T, Nakamura Y, Shirakawa H, et al. (2005). The role of EF-hand domains and C2 domain in regulation of enzymatic activity of phospholipase Czeta. J. Biol. Chem. 280: Lee J, Zhou J, Zheng X, Cho S, et al. (2012). Identification of a novel cis-element that regulates alternative splicing of Bcl-x pre-mrna. Biochem. Biophys. Res. Commun. 420: Nagao K, Togawa N, Fujii K, Uchikawa H, et al. (2005). Detecting tissue-specific alternative splicing and diseaseassociated aberrant splicing of the PTCH gene with exon junction microarrays. Hum. Mol. Genet. 14: Nomikos M, Blayney LM, Larman MG, Campbell K, et al. (2005). Role of phospholipase C-zeta domains in Ca 2+ - dependent phosphatidylinositol 4,5-bisphosphate hydrolysis and cytoplasmic Ca 2+ oscillations. J. Biol. Chem. 280: Rebecchi MJ and Pentyala SN (2000). Structure, function, and control of phosphoinositide-specific phospholipase C. Physiol. Rev. 80: Ross PJ, Beyhan Z, Iager AE, Yoon SY, et al. (2008). Parthenogenetic activation of bovine oocytes using bovine and murine phospholipase C zeta. BMC Dev. Biol. 8: 16. Saunders CM, Larman MG, Parrington J, Cox LJ, et al. (2002). PLC zeta: a sperm-specific trigger of Ca 2+ oscillations in eggs and embryo development. Development 129: Saunders CM, Swann K and Lai FA (2007). PLCzeta, a sperm-specific PLC and its potential role in fertilization. Biochem. Soc. Symp Wang H, Hubbell E, Hu JS, Mei G, et al. (2003). Gene structure-based splice variant deconvolution using a microarray platform. Bioinformatics 19 (Suppl 1): i315-i322. Wu H, Smyth J, Luzzi V, Fukami K, et al. (2001). Sperm factor induces intracellular free calcium oscillations by stimulating the phosphoinositide pathway. Biol. Reprod. 64: Yoneda A, Kashima M, Yoshida S, Terada K, et al. (2006). Molecular cloning, testicular postnatal expression, and oocyteactivating potential of porcine phospholipase Czeta. Reproduction 132: Young C, Grasa P, Coward K, Davis LC, et al. (2009). Phospholipase C zeta undergoes dynamic changes in its pattern of localization in sperm during capacitation and the acrosome reaction. Fertil. Steril. 91:
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4)
 Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
Identification and characterization of multiple splice variants of Cdc2-like kinase 4 (Clk4) Vahagn Stepanyan Department of Biological Sciences, Fordham University Abstract: Alternative splicing is an
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus:
 RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
General Biology 1004 Chapter 11 Lecture Handout, Summer 2005 Dr. Frisby
 Slide 1 CHAPTER 11 Gene Regulation PowerPoint Lecture Slides for Essential Biology, Second Edition & Essential Biology with Physiology Presentation prepared by Chris C. Romero Neil Campbell, Jane Reece,
Slide 1 CHAPTER 11 Gene Regulation PowerPoint Lecture Slides for Essential Biology, Second Edition & Essential Biology with Physiology Presentation prepared by Chris C. Romero Neil Campbell, Jane Reece,
The Alternative Choice of Constitutive Exons throughout Evolution
 The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
The Alternative Choice of Constitutive Exons throughout Evolution Galit Lev-Maor 1[, Amir Goren 1[, Noa Sela 1[, Eddo Kim 1, Hadas Keren 1, Adi Doron-Faigenboim 2, Shelly Leibman-Barak 3, Tal Pupko 2,
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy
 Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Generation and validation of mtef4-knockout mice.
 Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
Bin Liu, Lei Yang, Binfang Huang, Mei Cheng, Hui Wang, Yinyan Li, Dongsheng Huang, Jian Zheng,
 The American Journal of Human Genetics, Volume 91 Supplemental Data A Functional Copy-Number Variation in MAPKAPK2 Predicts Risk and Survival of Lung Cancer Bin Liu, Lei Yang, Binfang Huang, Mei Cheng,
The American Journal of Human Genetics, Volume 91 Supplemental Data A Functional Copy-Number Variation in MAPKAPK2 Predicts Risk and Survival of Lung Cancer Bin Liu, Lei Yang, Binfang Huang, Mei Cheng,
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Department of Pharmaceutical Sciences, School of Pharmacy, Northeastern University, Boston, MA 02115, USA 2
 Pancreatic Cancer Cell Exosome-Mediated Macrophage Reprogramming and the Role of MicroRNAs 155 and 125b2 Transfection using Nanoparticle Delivery Systems Mei-Ju Su 1, Hibah Aldawsari 2, and Mansoor Amiji
Pancreatic Cancer Cell Exosome-Mediated Macrophage Reprogramming and the Role of MicroRNAs 155 and 125b2 Transfection using Nanoparticle Delivery Systems Mei-Ju Su 1, Hibah Aldawsari 2, and Mansoor Amiji
Senior Thesis. Presented to. The Faculty of the School of Arts and Sciences Brandeis University
 Greenwald 1 Mouse intercellular adhesion molecule 1 (ICAM-1) isoforms demonstrate different binding affinities to mouse macrophage-1 antigen (Mac-1) and preliminary evidence for alternatively-spliced variants
Greenwald 1 Mouse intercellular adhesion molecule 1 (ICAM-1) isoforms demonstrate different binding affinities to mouse macrophage-1 antigen (Mac-1) and preliminary evidence for alternatively-spliced variants
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
capacitation hyperactivation acrosome hyperactivation AR bovine serum albumin BSA non-genomic effect isothiocyanate; FITC PR mrna P hyperactivation HA
 17 2 47 54 2002 P PRP total RNA cdna PCR primer set PR mrna P hyperactivation HA AR Ca PR P HA AR P Ca PR mrna P DNA C PR PR P P HA AR Ca mrna capacitation hyperactivation acrosome reaction; AR hyperactivation
17 2 47 54 2002 P PRP total RNA cdna PCR primer set PR mrna P hyperactivation HA AR Ca PR P HA AR P Ca PR mrna P DNA C PR PR P P HA AR Ca mrna capacitation hyperactivation acrosome reaction; AR hyperactivation
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Chapter 11. How Genes Are Controlled. Lectures by Edward J. Zalisko
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Campbell Essential Biology, Fifth Edition, and Campbell Essential Biology with Physiology, Fourth Edition Eric J. Simon, Jean L. Dickey, and
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3.
 Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Pre-mRNA has introns The splicing complex recognizes semiconserved sequences
 Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
II. The effect of strain and IGF-1 gene transfer using SMGT techniques on some semen characteristics:-
 The experimental work of this study was carried out in poultry farm of the Department of Animal Production, Faculty of Agriculture, Benha University, Egypt with cooperation of Genetic Engineering and Biotechnology
The experimental work of this study was carried out in poultry farm of the Department of Animal Production, Faculty of Agriculture, Benha University, Egypt with cooperation of Genetic Engineering and Biotechnology
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Product Manual. Omni-Array Sense Strand mrna Amplification Kit, 2 ng to 100 ng Version Catalog No.: Reactions
 Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Supplemental Data. Integrating omics and alternative splicing i reveals insights i into grape response to high temperature
 Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Supplemental Data Integrating omics and alternative splicing i reveals insights i into grape response to high temperature Jianfu Jiang 1, Xinna Liu 1, Guotian Liu, Chonghuih Liu*, Shaohuah Li*, and Lijun
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons
 The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr)
 Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
Single Cell Quantitative Polymer Chain Reaction (sc-qpcr) Analyzing gene expression profiles from a bulk population of cells provides an average profile which may obscure important biological differences
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Shangqiu Medical College, Shangqiu, China 2. Corresponding author: M.Z. Zhang
 Comparison of the expression of human equilibrative nucleotide transporter 1 (hent1) and ribonucleotide reductase subunit M1 (RRM1) genes in seven non-hodgkin lymphoma cell lines H.B. Zhao 1,2, X.F. Zhang
Comparison of the expression of human equilibrative nucleotide transporter 1 (hent1) and ribonucleotide reductase subunit M1 (RRM1) genes in seven non-hodgkin lymphoma cell lines H.B. Zhao 1,2, X.F. Zhang
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
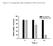 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Biochemistry 2000 Sample Question Transcription, Translation and Lipids. (1) Give brief definitions or unique descriptions of the following terms:
 (1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
(1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002
 Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A. Real-Time RT-PCR Detection.
 For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
Translation Activity Guide
 Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Effects of methionine-containing dipeptides on α s1 casein expression in bovine mammary epithelial cells *
 Journal of Animal and Feed Sciences, 16, Suppl. 2, 2007, 325 329 Effects of methionine-containing dipeptides on α s1 casein expression in bovine mammary epithelial cells * H.H. Wu 1, J.Y. Yang 1,2, K.
Journal of Animal and Feed Sciences, 16, Suppl. 2, 2007, 325 329 Effects of methionine-containing dipeptides on α s1 casein expression in bovine mammary epithelial cells * H.H. Wu 1, J.Y. Yang 1,2, K.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Chapter 5: The Structure and Function of Large Biological Molecules
 Chapter 5: The Structure and Function of Large Biological Molecules 1. Name the four main classes of organic molecules found in all living things. Which of the four are classified as macromolecules. Define
Chapter 5: The Structure and Function of Large Biological Molecules 1. Name the four main classes of organic molecules found in all living things. Which of the four are classified as macromolecules. Define
Sex Determination and Gonadal Sex Differentiation in Fish
 Sex Determination and Gonadal Sex Differentiation in Fish Yoshitaka Nagahama Okazaki National Research Institutes, Japan This first slide shows the processes of gonadal sex differentiation and gametogenesis
Sex Determination and Gonadal Sex Differentiation in Fish Yoshitaka Nagahama Okazaki National Research Institutes, Japan This first slide shows the processes of gonadal sex differentiation and gametogenesis
Fertilization depends on mechanisms that help sperm meet eggs of the same species.
 Fertilization depends on mechanisms that help sperm meet eggs of the same species. www.uchsc.edu/ltc/fertilization.html Fertilization union of sperm and egg Is a chain of events. Interruption of any step
Fertilization depends on mechanisms that help sperm meet eggs of the same species. www.uchsc.edu/ltc/fertilization.html Fertilization union of sperm and egg Is a chain of events. Interruption of any step
Relationship between SPOP mutation and breast cancer in Chinese population
 Relationship between SPOP mutation and breast cancer in Chinese population M.A. Khan 1 *, L. Zhu 1 *, M. Tania 1, X.L. Xiao 2 and J.J. Fu 1 1 Key Laboratory of Epigenetics and Oncology, The Research Center
Relationship between SPOP mutation and breast cancer in Chinese population M.A. Khan 1 *, L. Zhu 1 *, M. Tania 1, X.L. Xiao 2 and J.J. Fu 1 1 Key Laboratory of Epigenetics and Oncology, The Research Center
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Received 26 January 1996/Returned for modification 28 February 1996/Accepted 15 March 1996
 MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
Oligo Sequence* bp %GC Tm Hair Hm Ht Position Size Ref. HIVrt-F 5 -CTA-gAA-CTT-TRA-ATg-CAT-ggg-TAA-AAg-TA
 Human immunodeficiency virus (HIV) detection & quantitation by qrt-pcr (Taqman). Created on: Oct 26, 2010; Last modified by: Jul 17, 2017; Version: 3.0 This protocol describes the qrt-pcr taqman based
Human immunodeficiency virus (HIV) detection & quantitation by qrt-pcr (Taqman). Created on: Oct 26, 2010; Last modified by: Jul 17, 2017; Version: 3.0 This protocol describes the qrt-pcr taqman based
Bioinformatics Laboratory Exercise
 Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Alternative splicing control 2. The most significant 4 slides from last lecture:
 Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Muscular Dystrophy. Biol 405 Molecular Medicine
 Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Muscular Dystrophy Biol 405 Molecular Medicine Duchenne muscular dystrophy Duchenne muscular dystrophy is a neuromuscular disease that occurs in ~ 1/3,500 male births. The disease causes developmental
Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast cancer cells
 Int J Clin Exp Pathol 2017;10(5):5039-5062 www.ijcep.com /ISSN:1936-2625/IJCEP0052419 Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast
Int J Clin Exp Pathol 2017;10(5):5039-5062 www.ijcep.com /ISSN:1936-2625/IJCEP0052419 Original Article Differential expression profile analysis of mirnas with HER-2 overexpression and intervention in breast
SALSA MLPA probemix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four probes have been adjusted.
 mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
2.2 Cell Construction
 2.2 Cell Construction Elemental composition of typical bacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
2.2 Cell Construction Elemental composition of typical bacterial cell C 50%, O 20%, N 14%, H 8%, P 3%, S 1%, and others (K +, Na +, Ca 2+, Mg 2+, Cl -, vitamin) Molecular building blocks Lipids Carbohydrates
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell
 Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
AIDS - Knowledge and Dogma. Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/ , Vienna, Austria
 AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
Processing of RNA II Biochemistry 302. February 13, 2006
 Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described
 Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
Supplemental Methods In vitro DNase I foot printing. In vitro DNase I footprinting was performed as described previously 1 2 using 32P-labeled 211 bp fragment from 3 HS1. Footprinting reaction mixes contained
Doctor of Philosophy
 Regulation of Gene Expression of the 25-Hydroxyvitamin D la-hydroxylase (CYP27BI) Promoter: Study of A Transgenic Mouse Model Ivanka Hendrix School of Molecular and Biomedical Science The University of
Regulation of Gene Expression of the 25-Hydroxyvitamin D la-hydroxylase (CYP27BI) Promoter: Study of A Transgenic Mouse Model Ivanka Hendrix School of Molecular and Biomedical Science The University of
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
