Promiscuous RNA binding by Polycomb Repressive Complex 2
|
|
|
- Ann Tate
- 5 years ago
- Views:
Transcription
1 Promiscuous RNA binding by Polycomb Repressive Complex 2 Chen Davidovich 1,2, Leon Zheng 3, Karen J. Goodrich 1,2, and Thomas R. Cech 1,2,4 1 Department of Chemistry & Biochemistry, Howard Hughes Medical Institute, University of Colorado, Boulder, Colorado, 80309, USA 2 BioFrontiers Institute, University of Colorado, Boulder, Colorado, 80309, USA 3 Medical Scientist Training Program, University of Colorado School of Medicine, Aurora, Colorado, 80045, USA 4 Correspondence to: thomas.cech@colorado.edu Supplementary Materials Supplementary Figure 1. In vitro histone methyltransferase (HMTase) assay Supplementary Figure 2. Sequence source for HOTAIR RNA and HOTAIR 400 RNA Supplementary Figure 3. PRC2-RNA direct binding assay after prolonged incubation Supplementary Figure 4. Direct binding assay of PRC2 with various RNAs Supplementary Figure 5. Secondary structure prediction of MBP Supplementary Figure 6. Gene Ontology (GO) analysis for genes that were up-regulated following SUZ12 knockdown Supplementary Figure 7. Original western blots used to generate Figure 7a Supplementary Table 1. Genes observed with Ezh2-FE larger than 3 Supplementary Table 2. GO analysis for genes with high Ezh2-FE and coverage Supplementary Table 3. Data for Fig 5b Supplementary Table 4. PRC2-Regulated and EZH2-Associated genes Supplementary Table 5. GO analysis for genes up-regulated by SUZ12 KD Supplementary Table 6. ChIP-seq datasets used for Figure 6 Supplementary Table 7. NGS datasets generated in the scope of this study
2 Supplementary Figure 1. In vitro histone methyltransferase (HMTase) assay. Assays carried out under identical conditions, except for the exclusion of PRC2, H3.1 or both as highlighted. The higher radioactive signal following TCA precipitation, appearing only in the presence of both PRC2 and H3.1 histone, confirmed HMTase activity of PRC2. Some low signal was observed in the absence of histone substrate, possibly because of loaded SAM that was co-precipitated with PRC2 or a weak auto HMTase activity of PRC2, as previously observed 4. Error bars represents standard deviations that were generated based on 3 independent samples.
3 Supplementary Figure 2. Sequence source for HOTAIR RNA and HOTAIR 400 RNA. The sequence of HOTAIR (in red) originated from a previously generated cdna clone 58 and is in accord with the HOTAIR RNA examined in previous studies 24,34. We found this RNA construct prone to aggregation under various conditions, evidenced by the presence of radiolabeled RNA in wells and across lanes following non-denaturing gel electrophoresis, in the absence or presence of protein (Fig 1b). RNA aggregation was eliminated by adding approximately 50 nucleotides upstream and downstream of HOTAIR 1-300, forming HOTAIR 400 RNA (see Supplementary Methods for complete details of DNA templates used for in vitro RNA transcription). Importantly, all bases that were added (in light blue) were not arbitrarily tailored but appear within cellular HOTAIR transcripts (in blue).
4 Supplementary Figure 3. PRC2-RNA direct binding assay after prolonged incubation. To verify that dissociation constants were derived under equilibrium conditions, incubation time was increased eightfold, from 30 min to 240 min. The fraction of bound RNA was not increased following this prolonged incubation (panels a). An increase of 43% in equilibrium dissociation constant (panel b) is most likely due to loss of similar fraction of RNA binding activity by PRC2 due to the prolonged incubation period (4 hours at 30 ⁰C) prior to the EMSA experiment.
5 Supplementary Figure 4. Direct binding assay of PRC2 with various RNAs. (a) EMSA of different RNAs after incubation in the presence or absence of PRC2 complex confirms similar affinity of PRC2 to the following: a 400 base RNA from the 5 domain of HOTAIR RNA (HOTAIR 400 for sense RNA and ashotair 400 for antisense RNA), an approximately 500 base RNA representing the entire A-region from the RepA gene including all tandem repeats (A-region for sense and asa-region for antisense strand), the 397 base mouse telomerase RNA (mouse TR), a 157 base RNA representing the wild type (wt) P4-P6 region within the Tetrahymena group I intron, and a mutant P4-P6 RNA that cannot form tertiary structure 49. (b) Direct binding EMSA of the antisense strand of HOTAIR (ashotair 1-300, binding curve presented in Fig 1c).
6 Supplementary Figure 5. Secondary structure prediction of MBP Secondary structure prediction of MBP performed by mfold using default settings. Presented are the top three hits, those with the lowest Gs. These predictions failed to identify the two-hairpin motif (in dashed frame) that was previously suggested to be enriched in short ncrnas associated with PRC2 28.
7 Supplementary Figure 6. Gene Ontology (GO) analysis for genes that were up-regulated following SUZ12 knockdown. GO analysis yielded a notable network of significant GO terms for developmental processes, with specification at neuron development. GO term IDs indicated in brackets, p-values indicated under each GO term name.
8 Supplementary Figure 7. Original western blots used to generate Figure 7a. Full lanes shown as captured, without further cropping.
9 Supplementary Table 6 ChIP-seq datasets used for Figure 6 Dataset (cell line, method, additional information) GEO Accession Number Data source and/or reference Human Dnd41, ChIP seq, Input GSM ENCODE 59,60 Human Dnd41, ChIP seq, EZH2 GSM ENCODE 59,60 Human Dnd41, ChIP seq, H3K4me3 GSM ENCODE 59,60 Human Dnd41, ChIP seq, H3K27me3 GSM ENCODE 59,60 Human Dnd41, ChIP seq, H3K36me3 GSM ENCODE 59,60 Human HSMMtube, ChIP seq, Control GSM ENCODE 59,60 Human HSMMtube, ChIP seq, EZH2 GSM ENCODE 59,60 Human HSMMtube, ChIP seq, H3K4me3 GSM ENCODE 59,60 Human HSMMtube, ChIP seq, H3K27me3 GSM ENCODE 59,60 Human HSMMtube, ChIP seq, H3K36me3 GSM ENCODE 59,60 Human NHEK, ChIP seq, Input GSM ENCODE 59,60 Human NHEK, ChIP seq, EZH2 GSM ENCODE 59,60 Human NHEK, ChIP seq, H3K4me3 GSM ENCODE 59,60 Human NHEK, ChIP seq, H3K27me3 GSM ENCODE 59,60 Human NHEK, ChIP seq, H3K36me3 GSM ENCODE 59,60 Human NHLF, ChIP seq, Input GSM ENCODE 59,60 Human NHLF, ChIP seq, EZH2 GSM ENCODE 59,60 Human NHLF, ChIP seq, H3K4me3 GSM ENCODE 59,60 Human NHLF, ChIP seq, H3K27me3 GSM ENCODE 59,60 Human NHLF, ChIP seq, H3K36me3 GSM ENCODE 59,60 Mouse E14, RNA Seq GSM Mouse E14, ChIP seq, H3K36me3 GSM Mouse E14, ChIP seq, H3K27me3 GSM Mouse E14, ChIP seq, H3K4me3 GSM Mouse E14, ChIP seq, EZH2 GSM
10 Supplementary Table 7 NGS datasets generated in the scope of this study Cell line Treatment Method Specifications Number of reads aligned to the human genome HEK293T/17 untreated ChIP seq H3K4me2/3 30 M HEK293T/17 untreated ChIP seq EZH2 13 M HEK293T/17 sisuz12 RNA seq biological replicate 1, Poly A isolated 33 M HEK293T/17 sisuz12 RNA seq biological replicate 2, Poly A isolated 37 M HEK293T/17 sictrl RNA seq biological replicate 1, Poly A isolated 29 M HEK293T/17 sictrl RNA seq biological replicate 2, Poly A isolated 33 M
11 References 1. Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, (2011). 2. Cao, R. & Zhang, Y. SUZ12 is required for both the histone methyltransferase activity and the silencing function of the EED EZH2 complex. Mol Cell 15, (2004). 3. Margueron, R. et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461, (2009). 4. Xu, C. et al. Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc Natl Acad Sci U S A 107, (2010). 5. Yuan, W. et al. Dense chromatin activates Polycomb repressive complex 2 to regulate H3 lysine 27 methylation. Science 337, (2012). 6. Schmitges, F.W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol Cell 42, (2011). 7. Yuan, W. et al. H3K36 methylation antagonizes PRC2 mediated H3K27 methylation. J Biol Chem 286, (2011). 8. Boyer, L.A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, (2006). 9. Lee, T.I. et al. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125, (2006). 10. Ram, O. et al. Combinatorial patterning of chromatin regulators uncovered by genome wide location analysis in human cells. Cell 147, (2011). 11. Bernstein, B.E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, (2006). 12. Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nat Cell Biol 8, (2006). 13. Roh, T.Y., Cuddapah, S., Cui, K. & Zhao, K. The genomic landscape of histone modifications in human T cells. Proc Natl Acad Sci U S A 103, (2006). 14. Mousavi, K., Zare, H., Wang, A.H. & Sartorelli, V. Polycomb protein Ezh1 promotes RNA polymerase II elongation. Mol Cell 45, (2012). 15. Herz, H.M. et al. Polycomb repressive complex 2 dependent and independent functions of Jarid2 in transcriptional regulation in Drosophila. Mol Cell Biol 32, (2012). 16. Brookes, E. et al. Polycomb associates genome wide with a specific RNA polymerase II variant, and regulates metabolic genes in ESCs. Cell Stem Cell 10, (2012). 17. Ballare, C. et al. Phf19 links methylated Lys36 of histone H3 to regulation of Polycomb activity. Nat Struct Mol Biol (2012). 18. Musselman, C.A. et al. Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1. Nat Struct Mol Biol (2012). 19. Brien, G.L. et al. Polycomb PHF19 binds H3K36me3 and recruits PRC2 and demethylase NO66 to embryonic stem cell genes during differentiation. Nat Struct Mol Biol (2012). 20. Schwartz, Y.B. & Pirrotta, V. Polycomb silencing mechanisms and the management of genomic programmes. Nat Rev Genet 8, 9 22 (2007). 21. Cuddapah, S. et al. A novel human polycomb binding site acts as a functional polycomb response element in Drosophila. PLoS One 7, e36365 (2012). 22. Sing, A. et al. A vertebrate Polycomb response element governs segmentation of the posterior hindbrain. Cell 138, (2009).
12 23. Woo, C.J., Kharchenko, P.V., Daheron, L., Park, P.J. & Kingston, R.E. A region of the human HOXD cluster that confers polycomb group responsiveness. Cell 140, (2010). 24. Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, (2010). 25. Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, (2007). 26. Chu, C., Qu, K., Zhong, F.L., Artandi, S.E. & Chang, H.Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA chromatin interactions. Mol Cell 44, (2011). 27. Zhao, J., Sun, B.K., Erwin, J.A., Song, J.J. & Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, (2008). 28. Kanhere, A. et al. Short RNAs are transcribed from repressed polycomb target genes and interact with polycomb repressive complex 2. Mol Cell 38, (2010). 29. Zhao, J. et al. Genome wide identification of polycomb associated RNAs by RIP seq. Mol Cell 40, (2010). 30. Khalil, A.M. et al. Many human large intergenic noncoding RNAs associate with chromatinmodifying complexes and affect gene expression. Proc Natl Acad Sci U S A 106, (2009). 31. Guttman, M. et al. lincrnas act in the circuitry controlling pluripotency and differentiation. Nature 477, (2011). 32. Maenner, S. et al. 2 D structure of the A region of Xist RNA and its implication for PRC2 association. PLoS Biol 8, e (2010). 33. Duszczyk, M.M., Wutz, A., Rybin, V. & Sattler, M. The Xist RNA A repeat comprises a novel AUCG tetraloop fold and a platform for multimerization. RNA 17, (2011). 34. Kaneko, S. et al. Phosphorylation of the PRC2 component Ezh2 is cell cycle regulated and upregulates its binding to ncrna. Genes Dev 24, (2010). 35. Kowalczykowski, S.C. et al. Cooperative and noncooperative binding of protein ligands to nucleic acid lattices: experimental approaches to the determination of thermodynamic parameters. Biochemistry 25, (1986). 36. Epstein, I.R. Kinetics of nucleic acid large ligand interactions: exact Monte Carlo treatment and limiting cases of reversible binding. Biopolymers 18, (1979). 37. Broderick, J.A., Salomon, W.E., Ryder, S.P., Aronin, N. & Zamore, P.D. Argonaute protein identity and pairing geometry determine cooperativity in mammalian RNA silencing. RNA 17, (2011). 38. Record, M.T., Jr., Lohman, M.L. & De Haseth, P. Ion effects on ligand nucleic acid interactions. J Mol Biol 107, (1976). 39. Makino, D.L., Baumgartner, M. & Conti, E. Crystal structure of an RNA bound 11 subunit eukaryotic exosome complex. Nature 495, 70 5 (2013). 40. Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA Seq. Nat Methods 5, (2008). 41. Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, (2012). 42. Shaw, G., Morse, S., Ararat, M. & Graham, F.L. Preferential transformation of human neuronal cells by human adenoviruses and the origin of HEK 293 cells. FASEB J 16, (2002). 43. Schwartz, J.C. et al. FUS binds the CTD of RNA polymerase II and regulates its phosphorylation at Ser2. Genes Dev 26, (2012). 44. Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet 4, e (2008).
13 45. Hansen, K.H. et al. A model for transmission of the H3K27me3 epigenetic mark. Nat Cell Biol. 10, (2008). 46. Sun, S. et al. Jpx RNA Activates Xist by Evicting CTCF. Cell 153, (2013). 47. Mazzone, J. & Pickett, J. The Household Diary Study, Mail Use & Attitudes in FY (2011). 48. Khersonsky, O. & Tawfik, D.S. Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu Rev Biochem 79, (2010). 49. Murphy, F.L., Wang, Y.H., Griffith, J.D. & Cech, T.R. Coaxially stacked RNA helices in the catalytic center of the Tetrahymena ribozyme. Science 265, (1994). 50. Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory efficient alignment of short DNA sequences to the human genome. Genome Biol 10, R25 (2009). 51. Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, (2010). 52. Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, (2009). 53. Robinson, J.T. et al. Integrative genomics viewer. Nat Biotechnol 29, 24 6 (2011). 54. Zhang, Y. et al. Model based analysis of ChIP Seq (MACS). Genome Biol 9, R137 (2008). 55. Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate a Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B Methodological 57, (1995). 56. Schwartz, J.C. et al. FUS binds the CTD of RNA polymerase II and regulates its phosphorylation at Ser2. Genes Dev 26(2012). 57. Reimand, J., Kull, M., Peterson, H., Hansen, J. & Vilo, J. g:profiler a web based toolset for functional profiling of gene lists from large scale experiments. Nucleic Acids Res 35, W (2007). 58. Gupta, R.A. et al. Long non coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, (2010). 59. Consortium, E.P. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, (2012). 60. Consortium, E.P. A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol 9, e (2011).
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Long noncoding RNAs in DNA methylation: new players stepping into the old game
 DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
Hox genes. Discovered in Drosophila in 1923 by Bridges and Morgan Antennapaedia complex ANT-C Bithorax complex BX-C
 Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Figure S1, Beyer et al.
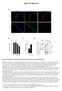 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Computational aspects of ChIP-seq. John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory
 Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION. Jude Fitzgibbon
 FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
ubiquitinylation succinylation butyrylation hydroxylation crotonylation
 Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
LncRNAs: The Ideal Composer of the Melody for Life
 RNA and Transcription 2016; 2(2): 11-15 http://www.sciencepublishinggroup.com/j/rnat doi: 10.11648/j.rnat.20160202.11 Review Article LncRNAs: The Ideal Composer of the Melody for Life Qingyu Cheng 1, Shengwei
RNA and Transcription 2016; 2(2): 11-15 http://www.sciencepublishinggroup.com/j/rnat doi: 10.11648/j.rnat.20160202.11 Review Article LncRNAs: The Ideal Composer of the Melody for Life Qingyu Cheng 1, Shengwei
H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells
 Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC)
 Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
ChIP-seq data analysis
 ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Peak-calling for ChIP-seq and ATAC-seq
 Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-06-1-0093 TITLE: An Epigenetic Link to Prostate Cancer PRINCIPAL INVESTIGATOR: Raphael F Margueron, Ph.D. CONTRACTING ORGANIZATION: University of Medicine and Dentistry of New Jersey
AD Award Number: W81XWH-06-1-0093 TITLE: An Epigenetic Link to Prostate Cancer PRINCIPAL INVESTIGATOR: Raphael F Margueron, Ph.D. CONTRACTING ORGANIZATION: University of Medicine and Dentistry of New Jersey
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
The genetics of heterochromatin. in metazoa. mutations by means of X-ray irradiation" "for the discovery of the production of
 The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
GeneOverlap: An R package to test and visualize
 GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins?
 REVIEW ARTICLE The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins? Andrea Scelfo, Andrea Piunti and Diego Pasini Department
REVIEW ARTICLE The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins? Andrea Scelfo, Andrea Piunti and Diego Pasini Department
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
SUPPLEMENTARY INFORMATION. Intron retention is a widespread mechanism of tumor suppressor inactivation.
 SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells
 Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
DNA methylation and differentiation: HOX genes in muscle cells
 Tsumagari et al. Epigenetics & Chromatin 2013, 6:25 RESEARCH Open Access DNA methylation and differentiation: HOX genes in muscle cells Koji Tsumagari 1, Carl Baribault 2, Jolyon Terragni 3, Sruti Chandra
Tsumagari et al. Epigenetics & Chromatin 2013, 6:25 RESEARCH Open Access DNA methylation and differentiation: HOX genes in muscle cells Koji Tsumagari 1, Carl Baribault 2, Jolyon Terragni 3, Sruti Chandra
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to
 Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq Data
 BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
Transcriptional repression of Xi
 Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
p53 cooperates with DNA methylation and a suicidal interferon response to maintain epigenetic silencing of repeats and noncoding RNAs
 p53 cooperates with DNA methylation and a suicidal interferon response to maintain epigenetic silencing of repeats and noncoding RNAs 2013, Katerina I. Leonova et al. Kolmogorov Mikhail Noncoding DNA Mammalian
p53 cooperates with DNA methylation and a suicidal interferon response to maintain epigenetic silencing of repeats and noncoding RNAs 2013, Katerina I. Leonova et al. Kolmogorov Mikhail Noncoding DNA Mammalian
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Omaha, NE PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AWARD NUMBER: W81XWH-13-1-0075 TITLE: LincRNAs and AR Reactivation after Androgen Deprivation in Prostate Cancer Cells PRINCIPAL INVESTIGATOR: Xian-Ming Chen, MD CONTRACTING ORGANIZATION: Creighton University
AWARD NUMBER: W81XWH-13-1-0075 TITLE: LincRNAs and AR Reactivation after Androgen Deprivation in Prostate Cancer Cells PRINCIPAL INVESTIGATOR: Xian-Ming Chen, MD CONTRACTING ORGANIZATION: Creighton University
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Distinct patterns of epigenetic marks and transcription factor binding sites across promoters of sense-intronic long noncoding RNAs
 c Indian Academy of Sciences RESEARCH ARTICLE Distinct patterns of epigenetic marks and transcription factor binding sites across promoters of sense-intronic long noncoding RNAs SOURAV GHOSH 1,2, SATISH
c Indian Academy of Sciences RESEARCH ARTICLE Distinct patterns of epigenetic marks and transcription factor binding sites across promoters of sense-intronic long noncoding RNAs SOURAV GHOSH 1,2, SATISH
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome
 The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
V24 Regular vs. alternative splicing
 Regular splicing V24 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Regular splicing V24 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Participating Institutions Insitut Gustave Roussy, Villejuif, France Institut Bergonie, Bourdeaux, France. Sponsor Epizyme, Inc
 Phase 1 Study of EPZ-6438 (E7438), an Enhancer of Zeste Homolog-2 (EZH2) Inhibitor: Dose Determination and Preliminary Activity in Non-Hodgkin Lymphoma V. Ribrag, J-C. Soria, B. Thomson, L. Reyderman,
Phase 1 Study of EPZ-6438 (E7438), an Enhancer of Zeste Homolog-2 (EZH2) Inhibitor: Dose Determination and Preliminary Activity in Non-Hodgkin Lymphoma V. Ribrag, J-C. Soria, B. Thomson, L. Reyderman,
Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance
 Stringer et al. BMC Biology (2018) 16:104 https://doi.org/10.1186/s12915-018-0569-5 RESEARCH ARTICLE Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance Open Access
Stringer et al. BMC Biology (2018) 16:104 https://doi.org/10.1186/s12915-018-0569-5 RESEARCH ARTICLE Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance Open Access
Marta Puerto Plasencia. microrna sponges
 Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
gliomas. Fetal brain expected who each low-
 Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
STEM CELL GENETICS AND GENOMICS
 STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
Regulatory Roles of Noncoding RNA in Development and Disease
 Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 940 Regulatory Roles of Noncoding RNA in Development and Disease GAURAV KUMAR PANDEY ACTA UNIVERSITATIS UPSALIENSIS
Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 940 Regulatory Roles of Noncoding RNA in Development and Disease GAURAV KUMAR PANDEY ACTA UNIVERSITATIS UPSALIENSIS
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Genomic Methods in Cancer Epigenetic Dysregulation
 Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples
 DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Selective depletion of abundant RNAs to enable transcriptome analysis of lowinput and highly-degraded RNA from FFPE breast cancer samples LIBRARY
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
