Delineation, Functional Validation, and Bioinformatic Evaluation of Gene Expression in Thyroid Follicular Carcinomas with the PAX8-PPARG Translocation
|
|
|
- Terence Simon
- 5 years ago
- Views:
Transcription
1 Delineation, Functional Validation, and Bioinformatic Evaluation of Gene Expression in Thyroid Follicular Carcinomas with the PAX8-PPARG Translocation Thomas J. Giordano, 1 AmyY.M. Au, 4 Rork Kuick, 2 Dafydd G.Thomas, 1,3 Daniel R. Rhodes, 1 Kenneth G.Wilhelm, Jr., 3 MichelleVinco, 1 David E. Misek, 2 Donita Sanders, 1 Zhaowen Zhu, 5 Raffaele Ciampi, 5 Samir Hanash, 2,6 Arul Chinnaiyan, 1 Roderick J. Clifton-Bligh, 4 Bruce G. Robinson, 4 Yuri E. Nikiforov, 5 and Ronald J. Koenig 3 Abstract A subset of follicular thyroid carcinomas contains a balanced translocation, t(2;3)(q13;p25), that results in fusion of the paired box gene 8 (PAX8)and peroxisome proliferator-activated receptor c (PPARG) genes with concomitant expression of a PAX8-PPARg fusion protein, PPFP. PPFP is thought to contribute to neoplasia through a mechanism in which it acts as a dominant-negative inhibitor of wild-type PPARg. To better understand this type of follicular carcinoma, we generated global gene expression profiles using DNA microarrays of a cohortof follicular carcinomas along with other common thyroid tumors and used the data to derive a gene expression profile characteristic of PPFP-positive tumors. Transient transfection assays using promoters of four genes whose expression was highly associated with the translocation showed that each can be activated by PPFP. PPFP had unique transcriptional activities when compared with PAX8 or PPARg, although it had the potential to function in ways qualitatively similar to PAX8 or PPARg depending on the promoter and cellular environment. Bioinformatics analyses revealed that genes with increased expression in PPFP-positive follicular carcinomas include known PPAR target genes; genes involved in fatty acid, amino acid, and carbohydrate metabolism; micro- RNA target genes; and genes on chromosome 3p. These results have implications for the neoplastic mechanism of these follicular carcinomas. Well-differentiated thyroid carcinoma, excluding medullary carcinoma, is divided into papillary, follicular, and oncocytic cell types. Papillary carcinoma is the most common form of thyroid cancer. It has several recognized histologic subtypes that share activating mutations of genes within the RET/RAS/ Authors Affiliations: Departments of 1 Pathology, 2 Pediatrics, and 3 Internal Medicine, University of Michigan Medical School, Ann Arbor, Michigan; 4 Cancer Genetics Unit, Kolling Institute of Medical Research, University of Sydney, New South Wales, Australia; 5 Department of Pathology, University of Cincinnati College of Medicine, Cincinnati, Ohio; and 6 Fred Hutchinson Cancer Research Center, Seattle,Washington Received 9/19/05; revised1/18/06; accepted1/20/06. Grant support: Marilynn Collins Thyroid Cancer Fundand themillie Schembechler Endocrine Cancer Program of the University of Michigan Comprehensive Cancer; the Michigan National Institutes of Diabetes, Digestive, and Kidney Diseases Biotechnology Center (NIH DK58771); the Cell and Molecular Biology Core of the Michigan Diabetes Research and Training Center (NIH DK20572); NIH (NIH CA88041and NIH 5P30CA46592); National Cancer Institute Director s Challenge Program atthe University of Michigan (NIH CA84952); and the University of Michigan Comprehensive Cancer Center Bioinformatics Core (NIH 5P30CA46592). The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with18 U.S.C. Section1734 solely to indicate this fact. Note: Supplementary data for this article are available at Clinical Cancer Research Online ( Requests for reprints: Thomas J. Giordano, Department of Pathology, UH 2G332/0054, Ann Arbor, MI Phone: ; Fax: ; Giordano@umich.edu. F 2006 American Association for Cancer Research. doi: / ccr BRAF/MAPK signaling pathway (1, 2), although each papillary carcinoma subtype has a distinct gene expression signature (3). Follicular carcinoma is either associated with mutations of the RAS gene family (4) or a distinctive translocation of chromosomes 2and 3 that results in fusion of paired box gene 8 (PAX8) with peroxisome proliferator-activated receptor c (PPARG; ref. 5). Unlike the other thyroid carcinomas, the spectrum of mutations present in oncocytic carcinoma remains largely elusive, although mutations of GRIM-19 (NDUFA13), a gene involved in mitochondrial metabolism and regulation of cell death, recently have been identified in a minority of oncocytic carcinomas (6). PAX8 encodes a transcription factor that is expressed at high levels in thyrocytes. It is necessary for normal thyroid development and it directs the expression of many thyroidspecific genes. PPARG encodes a nuclear hormone receptor transcription factor whose activity is related to adipocyte differentiation (7 9), lipid and carbohydrate metabolism (10), and cellular proliferation and differentiation. PPARg is expressed at very low levels in normal thyroid and has no known function in that organ. In follicular carcinomas with the t(2;3)(q13;p25) translocation, the promoter and 5V coding region of PAX8 are fused in-frame with the coding region of PPARc1, resulting in a fusion protein designated PPFP. PPFP is expressed at high levels in PAX8-PPARG translocation-positive follicular carcinomas and is thought to play an oncogenic role through several mechanisms (11, 12)
2 Despite their low incidence compared with papillary carcinomas, follicular carcinomas with the PAX8-PPARG translocation are of great interest for several reasons. First, balanced translocations in epithelial tumors are uncommon compared with mesenchymal and hematologic tumors, for reasons that are not entirely clear. Defining the distinctive characteristics of thyroid tumors that favor chromosomal rearrangements would potentially be very informative. Second, as a nuclear receptor, PPARg displays myriad cellular functions, including potential roles in neoplastic transformation. A better understanding of its role in carcinogenesis is needed, and follicular carcinomas with this translocation should serve as a relevant model. Finally, molecularly targeted treatments for follicular carcinoma are needed; thus, an improved understanding of their biology and relevant therapeutic targets might be gained by elucidation of the consequences of the PAX8-PPARG translocation. With these rationales in mind, we derived the gene expression signature of a cohort of follicular carcinomas with this translocation and did supporting studies to corroborate the signature. Transfection experiments showed that the promoters of several genes in the PAX8-PPARG signature are activated by PPFP in a manner that partially overlaps the functions of PAX8 or PPARg. Bioinformatic analysis of the signature revealed potential roles of several metabolic pathways in the oncogenic action of PPFP. Materials and Methods Tumors, histopathology, and RNA isolation. A total of 93 unique thyroid samples consisting of 4 normal thyroids and 89 thyroid tumors [7 follicular carcinomas with the PAX8-PPARG translocation, 6 follicular carcinomas without the translocation (including Thy 203, described below), 10 follicular adenomas, 8 oncocytic carcinomas, 7 oncocytic adenomas, and 51 papillary carcinoma] were used to generate the gene expression profiles. Cases were derived from the University of Michigan, the University of Cincinnati Medical Center, and the Cooperative Human Tissue Network. The 51 papillary carcinomas and the four normal thyroids were described previously (3). Institutional review board approval was obtained. All tumors were diagnosed using accepted morphologic criteria. Frozen section slides and original permanent sections were reviewed, when available, to confirm the diagnoses and ensure research tissues were in agreement with the final pathologic diagnosis. All tissues were processed and RNAs were extracted similarly, as previously described (13). Microarray analysis. DNA microarray analysis was done using commercially available oligonucleotide DNA microarrays containing 22,283 probe sets (U133A GeneChip, Affymetrix, Santa Clara, CA) as reported (3, 14). crna preparation and hybridization, and scanning and image analysis of the arrays were done according to protocols of the manufacturer and as previously described (13), as was probe set intensity estimation and normalization. Our procedures gave average probe set intensities of f1,500 units, which we log-transformed using log[max(x + 50,0) + 50]. Estimates of fold changes between groups are the antilogarithms of the differences in means of the logtransformed data. Normalized and raw versions of the data are publicly available at index.html. Quantitative reverse transcription-pcr and sequencing. Reverse transcription real-time PCR was done as previously described (15). The probes and labels, shown in Table 1, were designed using Primer Express (ABI, Foster City, CA) and were obtained from Biosearch Technologies (Novato, CA). PCR conditions for each primer-probe combination were optimized for time, temperature, and magnesium concentration and done using a SmartCycler (Cepheid, Sunnyvale, CA). Table 1. Primers, probes, and labels used in quantitative reverse transcription-pcr studies Gene Direction Primer sequence PPARG Forward GGCCAAGGCTTCATGACAA Reverse AACTCAAACTTGGGCTCCATAAAG Probe TAAAGAGCCTGCGAAAGCC TTTTGGTG Label FAM PAX8-PPARG Forward AAAGCACCTTCGCACGGATG Reverse ACGGAGCTGATCCCAAAGTTGG Probe None Label SYBR green PGF Forward TCCTTGTCCCCCGTGATCT Reverse TGGCCGGAAAGAACAATGTC Probe CCCTCACACTTTGCCATTTGC TTGTACTG Label TAMRA ENO3 Forward GGACCGAGAATAAGTCCAAGTTTG Reverse AGCTGCTCCCGCCTTACAC Probe ATGCCATCCTGGGCGTGTCCTTG Label FAM AQP7 Forward ACCCTGCCCCACCCTTAC Reverse GGAATGGATGGGATCACAAATAAT Probe TCCATGGCCCTAGAGCACTTCT AAGCAGA Label FAM ANGPTL4 Forward CATGGTGCTGGTGCTGTTGT Reverse AGGTTGCTTTTATTCCAAGA ACTCTGT Probe CAAGCAGGCGCCAATGGTATCTGG Label FAM NOTE: All primers are written 5Vt o 3V. PCR products were sequenced in both directions by the University of Michigan DNA Sequencing Core. Tissue array and immunohistochemistry. A thyroid tissue array was constructed for validation by immunohistochemisty studies. The four PPFP(+) follicular carcinomas from the University of Michigan were used along with two PPFP( ) follicular carcinomas, four papillary carcinomas, two follicular adenomas, and four normal thyroids. Onemillimeter-diameter cores were arrayed in duplicate. Immunohistochemistry was done using a robotic autostainer (DAKO, Carpinteria, CA) and standard procedures using the Envision detection system (DAKO). The following antibodies and conditions were used: PPARg (Santa Cruz Biotechnology, Santa Cruz, CA), 1:100 dilution, high-ph Tris antigen retrieval, 60 minutes room temperature incubation; enolase 3 (ENO3; BD Transduction Laboratories, San Jose, CA), 1:50 dilution, citrate buffer antigen retrieval, 60 minutes room temperature incubation; and aquaporin 7 (AQP7; Abcam, Cambridge, MA), 1:800 dilution, citrate buffer antigen retrieval, 30 minutes room temperature incubation. Cell culture and transfection assays. All cells were maintained at 37jC with 5% CO 2. JEG-3 human choriocarcinoma cells were cultured in Eagle s MEM with 10% fetal bovine serum and penicillin/ streptomycin. N2a mouse preneuronal cells were cultured in DMEM with 10% fetal bovine serum and penicillin/streptomycin. Rat FRTL-5 thyroid cells were cultured in F12Coon s media with 5% fetal bovine serum, six hormone combination (1 mu/ml bovine TSH, 4 ng/ml insulin, 10 ng/ml somatostatin, 5 Ag/mL apotransferrin, 4 mg/ml
3 Profiles of PAX8-PPARG Follicular Carcinoma hydrocortisone, and 10 ng/ml glycyl-l-histidyl-l-lysine acetate; Sigma, St. Louis, MO) and penicillin/streptomycin. Whole thyroid glands were removed from dogs that had been previously anesthetized and exsanguinated as part of an unrelated, institutionally approved study. Thyroid glands were removed within 10 minutes of exsanguination. Glands were trimmed, minced, and primary cultures of thyrocytes were obtained following the method of Uyttersprot et al. (16). The promoters of four genes that, according to our microarray data, were induced specifically in PPFP(+) follicular carcinomas were selected for analysis by transfection. The PCR was used with Accuprime Pfx polymerase (Invitrogen, Carlsbad, CA) to amplify human AQP7 bp 2,359 to +90 (the transcription start site is +1), angiopoietin-like protein 4 (ANGPTL4) bp 2,565 to +77, placental growth factor (PGF) bp 2,372 to +34, and ENO3 bp 2,808 to +56. The respective templates for these reactions were human genomic DNA and bacterial artificial chromosomes RP11-886P16, RP11-104F2, and RP5-1050D4. The 5V PCR primers contained an Mlu1 restriction enzyme site and the 3V primers contained either an Xho1 orsal1 site. The PCR products were digested with the appropriate enzymes and ligated into the Mlu1 and Xho1 sites of pgl3-basic (Promega, Madison, WI). All constructs were confirmed by sequencing. For transfection, cells were plated into 24-well clusters. The day before transfection, the medium was replaced to include charcoalstripped serum. Transfections were done with LipofectAMINE and Plus reagents according to the protocol of the manufacturer (Invitrogen) in serum-free medium, and included 100 ng of the above-described pgl3- based firefly luciferase reporter plasmids, 100 ng transcription factor expression plasmid (PAX8, PPARg, PPFP, or empty vector pcdna3.1+; Invitrogen), and 0.5 to 1 ng of the internal control Renilla luciferase plasmid prl-sv40 (Promega). After 3 hours of transfection, an equal volume of culture medium containing 20% charcoal-stripped FCS and penicillin/streptomycin was added to the wells. The next day, the culture medium was replaced with medium containing either 10 Amol/L PPARg agonist ciglitazone (17) or vehicle ethanol (again with 10% stripped serum) for an additional 24 hours. The cells were lysed and analyzed for firefly and Renilla luciferase activities using the Promega dual luciferase reagents and protocol. Enriched feature tests. We tested a selected set of 977 probe sets for overrepresentation of any Gene Ontology terms, GenMAPP maps using probe set annotation from Affymetrix ( analysis/index.affx, version of May 31, 2005), as well as pathways defined in the Kyoto Encyclopedia of Genes and Genomes ( using methods similar to those previously reported (Thy 203 was omitted in this analysis; ref. 18). The 22,283 U133A GeneChip probe sets were collapsed to 12,44 distinct genes with unambiguous Entrez gene numbers, which reduced the 977 probe sets to 761 genes (460 up, 301 down). Overrepresentation of each annotation term (e.g., membership in a particular pathway) in this set of genes was tested using one-sided Fisher s exact tests. To estimate the false discovery rates for the most significantly enriched terms, the resulting P values were compared with P values obtained from 100 data sets in which the 761 genes were randomly selected. Bioinformatic analysis using oncomine. The Oncomine data mining platform (19) was used to compare the PPFP(+) and PPFP( ) follicular carcinoma gene expression profiles (including Thy 203 ). Genbank accession IDs corresponding to Affymetrix probe set IDs were downloaded from Netaffx ( Genbank IDs were mapped to Unigene Build 185. A map from Unigene to Entrez Gene ID was downloaded from Entrez Gene ( entrez/query.fcgi?db = gene). The data set was base 2log transformed (negative intensity values were removed) and median centered per array, and the SDs were normalized to one per array. Each gene was assessed for differential expression with Student s t test, done using the R statistical computing package. Tests were conducted both as two-sided for differential expression analysis and one-sided for overexpression analysis. To account for multiple hypothesis testing, Q values (estimated false discovery rates) were calculated as follows: Q ¼ ðnpþ R, where P is P value, N is the total number of genes analyzed, and R is the sorted rank of P value. Gene set collection. All identifiers were mapped to Entrez Gene IDs for analysis. The 22,283 probe sets were collapsed to 13,046 distinct Entrez gene IDs. In the case of multiple probe sets per Entrez gene ID, the probe set with the minimum P value was kept. Sets of biologically related genes were collected or derived from a number of external resources; those relevant to the data presented here are as follows: chromosome arm mappings were downloaded from the National Center for Biotechnology Information Map Viewer ( gov/mapview/), protein-protein interaction sets were downloaded from the Human Protein Reference Database ( and predicted micro-rna (mirna) target genes were downloaded from PicTar ( ref. 20). Gene set analysis. Oncomine gene expression signatures were defined as the top 20% of Entrez gene IDs with enough nonnegative values to perform a t test, rank-ordered by their P values in each differential expression analysis. This constitutes 12,078 distinct genes, giving 2,415 in the top 20%. The association of a gene expression signature and the gene set was assessed with Fisher s exact test. The false discovery rate was again estimated using Q values, calculated as follows: Q ¼ ðnpþ R, where N is the number of gene sets of a given type tested against each gene expression signature and R is the ascending order rank of the respective P value. Interactome. Approximately 16,000 known protein-protein interactions were downloaded from the Human Protein Reference Database ( ref. 21), a manually curated database of pairs of proteins that have experimental evidence for physical interaction. Oncomine reports pairs of differentially expressed genes that encode proteins with documented protein-protein interactions. Oncomine generates interactome maps for the top 10% of genes rank-ordered by their P values in each differential expression analysis. Results Gene expression profiling identifies follicular carcinomas with the PAX8-PPARG translocation. The main focus of this work was to identify the transcriptional changes that are specific to follicular carcinomas that contain the PAX8-PPARG translocation. For this purpose, we obtained gene expression profiles on 93 thyroid samples consisting of 4 normal thyroids and 89 thyroid tumors (13 follicular carcinomas, 10 follicular adenomas, 8 oncocytic carcinomas, 7 oncocytic adenomas, and 51 papillary carcinomas). It was possible to identify cases with the PAX8-PPARG translocation by examining the microarray data for increased expression of PPARg (Fig. 1). High PPARc transcript levels, compared with the other thyroid tumors, were present in seven of the follicular carcinomas. All the follicular patterned tumors (follicular carcinomas, follicular adenomas, oncocytic carcinomas, and oncocytic adenomas) were analyzed by reverse transcription-pcr for the presence of the fusion transcript (data not shown). The fusion transcript was detected in all seven follicular carcinomas with high PPARg expression and in only one other sample, a follicular carcinoma (Thy 203 ) that expressed very low levels of PPARg by microarray (Fig. 1). By reverse transcription real-time PCR, the threshold for detection of the fusion transcript occurred 10 cycles later for Thy 203 than for the seven follicular carcinomas with high PPARg expression, indicating that Thy 203 expresses the fusion transcript at f0.1% the level of those seven follicular carcinomas. Further, real-time PCR for the 3V end of PPARc showed an undetectable level of expression after 40 cycles of amplification (Table 2). Therefore, for analysis, we grouped
4 Fig. 1. Microarray analysis of PPARg expression in benign and malignantthyroid samples. High expression of PPARg correlates with PPFP expression, as analyzed by reverse transcription real-time PCR (not shown). Arrow, follicular carcinomathy 203. Thy 203 with the PPFP( ) follicular carcinomas. As expected from this extremely low-level expression, the microarray profile of Thy 203 was similar to those of the five other PPFP( ) follicular carcinomas. Characterization of transcript fusions in the PPFP(+) follicular carcinomas and Thy 203. PPFP transcripts have been reported to contain PAX8 exons 7, 8, or 9 fused to PPARg1 exon 1 (11). Reverse transcription-pcr using a forward primer in PAX8 exon 7 and a reverse primer in PPARg1 exon 1 followed by sequencing revealed that six of our seven PPFP(+) follicular carcinomas had transcripts with PAX8 exon 8 fused to PPARg1 exon 1, and one (Thy 150 ) had PAX8 exon 7 fused to PPARg1 exon 1. Thy 203 also showed fusion of PAX8 exon 8 to PPARg1 exon 1. Gene expression among follicular patterned lesions is a function of the PAX8-PPARG translocation. Principal component analysis was done to examine global differences in gene expression between samples. Principal component analysis of all follicular neoplasms (23 follicular carcinomas and follicular adenomas) revealed significant separation of the PPFP(+) follicular carcinomas from the PPFP( ) follicular carcinomas and the follicular adenomas (Fig. 2). This result indicates that the PAX8-PPARG translocation is the predominant source of the gene expression variation within this set of tumors. Thy 203 plotted among the other PPFP( ) follicular carcinomas, providing further support for its inclusion in the PPFP( ) follicular carcinoma group. Gene expression profile of follicular carcinomas with the PAX8- PPARG translocation. The most direct way to define the expression profile of follicular carcinomas with the PAX8- PPARG translocation would be comparison of a large number of follicular carcinomas with and without the translocation. However, in general, follicular carcinomas are relatively rare thyroid tumors and microarray analysis requires frozen tissue. Thus, only 13 follicular carcinomas were available for analysis. Therefore, we used all of the data from the various tumor types to identify genes with larger (or smaller) mrna levels in the seven PPFP(+) follicular carcinomas compared with the five PPFP( ) follicular carcinomas without this translocation (Thy 203 was omitted), which also were increased (or decreased) compared with non-follicular carcinoma tumor samples and normal tissue. We asked that two-sample t tests give P < 0.01 for the comparison of PPFP(+) follicular carcinomas to PPFP( ) follicular carcinomas, as well as for the comparison of PPFP(+) follicular carcinomas to the set of nonfollicular carcinoma samples. We further asked that the fold difference between PPFP(+) follicular carcinomas and each of the six groups individually be at least 1.5 and be in the same direction. This selected a set of 322 probe sets, 239 of which had increased values in the PPFP(+) follicular carcinomas. To estimate the false discovery rate for this gene list, we permuted the sample labels 1,000 times, and on average obtained only 3.85 qualifying probe sets in the 1,000 resulting data sets, so that we estimate the false discovery rate to be f1.2%. These 322 probe sets are identified in our Table 2. Quantitative reverse transcription-pcr validation of selected microarray data Tumor PPARG PGF ANGPTL4 AQP7 ENO3 PPFP(+) FC (n = 7) 21.2 (3.1) 24.4 (3.2) 23.7 (1.5) 23.8 (1.0) 25.9 (1.5) All others (n = 70) 38.6 (4.7) 29.1 (6.0) 38.1 (4.7) 37.4 (4.8) 33.5 (4.5) All PPFP( )FC(n = 10) 40.0 (0.0) 28.7 (7.1) 37.4 (5.6) 36.7 (5.5) 33.5 (5.2) PPFP( )FC(n = 6)* 40 (0.0) 29.6 (9.2) 35.6 (6.8) 38.7 (3.2) 32.3 (6.0) PPFP( )FC(n =4) c 40.0 (0.0) 27.4 (2.2) 40 (0.0) 33.7 (7.3) 35.4 (3.7) Thy NT (n = 4) 36.5 (7.0) 24.6 (3.4) (5.3) 33.4 (2.9) FA (n = 14) 40 (0.0) 25.9 (4.6) 38.3 (4.5) 36.4 (5.2) 31.3 (4.1) NH (n =8) 38.4(4.7) 27.9(5.5) 40(0.0) 40(0.0) 35.4(3.2) OA (n = 7) 39.6 (1.1) 34.7 (5.1) 40 (0.0) 32.3 (5.7) 31.6 (3.0) OC (n = 7) 37.6 (9.7) 35.8 (5.6) 38.4 (4.3) 40 (0.0) 35.6 (3.5) PC (n = 19) 38.2 (4.4) 28.8 (4.5) 37.4 (5.3) 39.5 (2.3) 34.8 (5.0) NOTE: Values in table expressed as mean cycle to threshold (SD). PCR reactions that did not reach threshold by the 40th cycle were assigned a value of 40. Abbreviations: FC, follicular carcinoma; NT, normal thyroid; FA, follicular adenoma; OA, oncocytic adenoma; OC, oncocytic carcinoma; PC, papillary carcinoma. *Follicular carcinomas that were used for DNA microarray analysis. cfollicular carcinomas that were not used in the microarray analysis
5 Profiles of PAX8-PPARG Follicular Carcinoma Fig. 2. Principle component analysis of log-transformed data for all probe sets for PPFP(+) follicular carcinomas (PPFP + FC), PPFP( ) follicular carcinomas (PPFP FC), follicular adenomas (FA), and normal thyroids.the first two principal components are plotted.the seven PPFP(+) follicular carcinomas are distinctly separated from the other thyroid follicular patterned tumors and the normal thyroid samples. Supplementary Data. When performing statistical tests for enriched features among sets of genes, below, we desired a somewhat larger list of genes, and for this used a weaker selection criterion that asked that the P values be <0.05 and the fold changes be at least 1.2. This selected 977 probe sets with an estimated false discovery rate of 11.4%. We show a smaller subset in Fig. 3 that qualified under a similar but more stringent selection criteria that required the two P values to be <0.001 and the fold changes be at least 2.0. This selected 80 probe sets (67 up, 13 down) representing 68 distinct genes (55 up, 13 down), and gave an estimated false discovery rate of 0.07% using 1,000 permuted data sets. Note that in our data, PPARG is the most differentially expressed gene, but this reflects the expression of PPFP in tumors with the PAX8-PPARG translocation. We also examined the expression of several thyrocyte differentiation markers genes (SLC5A5, TG, TPO, and TSHR) between the PPFP(+) follicular carcinomas and the other follicular cohorts and found few significant changes. Validation of select genes by reverse transcription real-time PCR. To validate the microarray data, reverse transcription real-time PCR was done using RNA from a set of tumors that partially overlapped with the set used for DNA microarray analysis. PPARG and four additional genes with increased expression in the PPFP(+) follicular carcinomas were selected for validation (ANGPTL4, AQP7, ENO3, and PGF). The results, reported as the number of cycles needed to reach threshold (cycle to threshold, C T ), are shown in Table 2listed by histologic type and translocation status. Overall, the PCR results validate the microarray data, including classification of Thy 203 as PPFP( ). Validation of select proteins by immunohistochemistry. To validate the microarray data at the protein level, immunohistochemistry for PPARg and two proteins (ENO3 and AQP7) identified in the PPFP(+) signature was done using a thyroid tissue array that contained four PPFP(+) follicular carcinomas as well as a 10 other thyroid tumors [including two PPFP( ) follicular carcinomas] and four normal thyroids. The results confirmed increased protein expression in PPFP(+) follicular carcinomas of PPARg (four of four, 100%), ENO3 (three of four, 75%), and aquaporin (three of four, 75%) compared with normal thyroid and the other thyroid tumors (Fig. 4). Functional validation of the gene expression signature by transient transfection assays. Two of the genes most strongly induced specifically in the PPFP(+) follicular carcinomas, AQP7 and ANGPTL4, are induced by PPARg in other tissues (22, 23). This suggests that PPFP might be inducing these genes in a PPARg-like manner, which runs counter to the conventional view that PPFP blocks PPARg action (11). Therefore, we used transient transfection to compare the abilities of PPFP, PPARg, and PAX8 to regulate the AQP7 and ANGPTL4 promoters. The promoters from two additional genes induced specifically in the PPFP(+) follicular carcinomas, PGF and ENO3, were also studied. We transfected three different cell lines and primary cultures of dog thyrocytes to assess whether cell type specific factors might regulate the response. Preliminary studies were done to show the functional capacity of the primary dog thyrocyte cultures. The thyrocytes were transfected with a reporter plasmid in which the rat sodium iodide symporter gene upstream enhancer element and 2kbp proximal promoter direct firefly luciferase expression (NIS-luc), together with a cytomegalovirus-renilla luciferase internal control plasmid. Exposure to 15 miu/ml TSH for 24 hours induced NIS-luc 2.3 F 0.09-fold (n = 3), indicating that the cells are responsive to TSH. Separate immunohistochemical experiments showed uniformly positive thyroglobulin staining (data not shown). Therefore, the dog thyrocytes were deemed appropriate for study. The AQP7 promoter was strongly induced by PPARg and PPFP, but not by PAX8, in all four cell types (Fig. 5). In general, PPARg and PPFP showed similar levels of induction in the presence of the PPARg agonist ciglitazone, although PPFP tended to have stronger ligand-independent activity. For example, in JEG-3 cells, PPARg induced luciferase 2.1-fold in the absence and 14-fold in the presence of ciglitazone, whereas the inductions with PPFP were 5.9- and 14-fold. Similarly, in primary cultures of dog thyrocytes, PPARg induced luciferase 8.8-fold in the absence and 29-fold in the presence of ciglitazone, whereas the inductions with PPFP were 23- and 46-fold. The ANGPTL4 promoter was less responsive than AQP7 and the data showed some cell type specificity, but the overall trend was similar with PPFP being at least as active as PPARg, and PAX8 having no activity (Fig. 6). For example, in JEG-3 cells, PPARg induced luciferase 1.2-fold in the absence and 2.4-fold in the presence of ciglitazone, whereas the inductions with PPFP were 1.9- and 3.4-fold. The ANGPTL4 promoter was not induced by either PPARg or PPFP in FRTL-5 cells, but in dog thyrocytes PPFP expression resulted in a 3.5-fold induction in the absence and a 5.2-fold induction in the presence of ciglitazone. The response in N2a cells was qualitatively similar to that in dog thyrocytes, with PPARg not inducing this promoter but PPFP resulting in inductions of 2.2- and 3.1-fold in the absence and presence of ciglitazone
6 Fig. 3. Gene expression signature of follicular carcinomas with the PAX8-PPARG translocation. Differentially expressed genes were identified by comparing follicular carcinomas with high levels of PPARg to normal thyroid and the other types of thyroid tumors according selection criteria described in Results. This yielded 55 genes whose expression is greater in the PPFP(+) follicular carcinoma group and13 genes whose expression is down in the PPFP(+) follicular carcinoma group. Fold change from the average of the two means for PPFP(+) follicular carcinomas and all other samples combined. FC(+), PPFP(+) follicular carcinoma; FC( ), PPFP( ) follicular carcinoma; FA, follicular adenoma; OA, oncocytic adenoma; OC, oncocytic carcinoma; PC, papillary carcinoma. In JEG-3 cells, the ENO3 promoter was also induced more strongly by PPFP (2.9-fold minus ciglitazone and 6.9-fold plus ciglitazone) than by PPARg (1.3- and 3.8-fold; Fig. 7A). However, this promoter was not induced by PPARg, PPFP, or PAX8 in N2a cells, FRTL-5 cells, or dog thyrocytes (data not shown). The PGF promoter was not induced by PPARg, PPFP, or PAX8 in any of the cell lines (data not shown). However, in dog thyrocytes, PPFP caused inductions of 6.4-fold minus ciglitazone and 10-fold plus ciglitazone, compared with no induction by PPARg and a modest f2.5-fold induction by PAX8 (Fig. 7B). Pathway analysis of the PAX8-PPARG signature genes. We analyzed the larger set of 977 probe sets found to be altered with the PAX8-PPARG translocation for enriched Gene Ontology terms, Kyoto Encyclopedia of Genes and Genomes pathways, and GenMAPP maps (Table 3). The most substantial enrichment was observed for pathways related to fatty acid metabolism. Induced genes in these pathways include several
7 Profiles of PAX8-PPARG Follicular Carcinoma Fig. 4. Immunohistochemistry for PPARg,ENO3,and AQP7. Representative results are shown for PPFP(+) follicular carcinoma, PPFP( ) follicular carcinoma, and papillary carcinoma. Note the nuclear staining for PPARg, the cytoplasmic staining for ENO3, and the membranous staining foraqp7, as well as the absence of staining of intratumoral endothelial cells. (Original magnification, 200). acyl-coa dehydrogenases (ACADL, ACADM, ACADS), acetyl- CoA acyltransferases (ACAA1, ACAA2), and hydroxyacyl-coa dehydrogenases (HADHA, HADHSC), all of which participate in fatty acid h-oxidation. Other metabolic pathways also were enriched, such as Kyoto Encyclopedia of Genes and Genomes pathways valine, leucine, and isoleucine degradation, and glycolysis/gluconeogenesis. These results are striking because PPARg regulates adipogenesis and glucose metabolism. Bioinformatic analysis using Oncomine. The Oncomine data mining platform ( ref. 19) was used to compare the PPFP(+) and PPFP( ) follicular carcinoma gene expression profiles (Thy 203 included), as a means of exploring for differences of potential biological significance between these groups of follicular carcinomas. Genes located on chromosome 3p were found to be overrepresented, with 95 of 341 measured genes on 3p being in the top 20% of the PPFP(+) up-regulated profile (P = 5.1E 5, Q = 0.002; all other chromosome arms had Q values of at least 0.2). Presumably, this is a consequence of the t(2;3)(q13;p25) chromosomal translocation and may reflect strong PAX8 regulatory sequences from chromosome 2exerting effects on chromosome 3p genes or other chromosome structural effects. Interestingly, the genes on 3p that are induced include two genes that are directly involved in fatty acid metabolism carnitine/acylacrnitine translocase (SLC25A20), which transfers fatty acylcarnitines into mitochondria, and acetyl-coa acyltransferase 1 (ACAA1), which participates in peroxisomal fatty acid h oxidation. Thus, although these genes seem to be functionally related to PPARg and it would be plausible to postulate that they are induced directly by PPFP, their membership in the set of induced genes from chromosome 3p suggests they may be induced secondary to the translocation itself. Recently, it has become clear that mirnas down-regulate the expression of a large number of genes posttranscriptionally by binding to short sequences in mrna 3V untranslated regions. Each mirna may regulate multiple mrnas, and one mrna may be regulated by multiple mirnas. Oncomine uses PicTar (20) to analyze for mirna target genes. Putative target genes for Fig. 5. Regulation of theaqp7 promoter. JEG-3 cells, N2a cells, FRTL-5 cells, or primary cultures of dog thyrocytes were cotransfected with an AQP7-luciferase plasmid, an internal control Renilla luciferase plasmid, and an expression vector for PAX8, PPARg, PPFP, or empty vector.the cells were cultured Fciglitazone for 2 days before determining reporter gene activities. Luciferase is normalized to Renilla luciferase, and for each experimentthe normalized activity with empty vector ciglitazone is defined as1. Columns, mean for four to six experiments; bars, SE
8 Fig. 6. Regulation of theangptl4 promoter. JEG-3 cells, N2a cells, FRTL-5 cells, or primary cultures of dog thyrocytes were cotransfected with an ANGPTL4-luciferase plasmid, an internal control Renilla luciferase plasmid, and an expression vector for PAX8, PPARg, PPFP, or empty vector. The cells were cultured Fciglitazone for 2 days before determining reporter gene activities. Luciferase is normalized to Renilla luciferase, and for each experiment the normalized activity with empty vector ciglitazone is defined as 1. Columns, mean for four to six experiments, except FRTL-5 (two experiments, each in triplicate); bars, SE. four mirnas are strongly overrepresented among the upregulated genes in PPFP(+) follicular carcinomas: mir-101 [104 of 329 measured target genes are in the top 20% of the PPFP(+) profile, P = 2.1E 7, Q = 3.6E 5], mir-30a-3p (55 of 160 measured target genes, P = 1.1E 5, Q = 9.3E 4), mir-200a (81 of 262 measured target genes, P = 1.2E 5, Q = 6.7E 4), and mir-199a (92of 309 measured target genes, P = 1.7E 5, Q = 7.1E 4). Twenty-one up-regulated genes are putative targets for at least three of these four mirnas, suggesting coordinate regulation. Included in this list are the oncogenes RUNX1/AML1 and SS18; PUM2, which encodes a protein thought to be involved in stem cell proliferation and self renewal; and NRP2, which encodes the vascular endothelial growth factor/pgf receptor neuropilin 2. The Oncomine Interactome identifies known physically interacting proteins (based up the Human Protein Reference Database; ref. 21) among the differentially expressed genes. This analysis revealed correlations between the expression of PPARg (which also measures PPFP) and two proteins that can function as PPARg coactivators, GADD45G (r 2 = 0.85) and NCOA4/ARA70 (r 2 = 0.48). This suggests that, in PPFP(+) follicular carcinoma, the PPARg-like transcriptional activity of PPFP may be magnified by increased expression of these proteins. Interactome analysis also revealed that the set of genes with increased transcript expression in PPFP(+) follicular carcinomas includes the epidermal growth factor receptor (EGFR) and several EGFR-interacting proteins: BRAF, which is activated by EGFR; CRK, an adapter protein that participates in EGFRmediated BRAF activation; VAV2, an oncogene that is phosphorylated by EGFR; STATs 1 and 5B, which also are activated by EGFR; the ERBB3 oncogene, which dimerizes with EGFR and is amplified in numerous cancers; PTK2, a tyrosine kinase that binds to and helps transmit motility signals from the EGFR; and HBEGF, which binds and activates EGFR with greater potency than EGF. The Interactome analysis also allows one to visualize overall networks of interactions by drawing an interaction map. This reveals that the EGFR is a central node that connects to numerous other up-regulated genes, including the oncogenes BRAF, PTK2, and EPHA2 (Fig. 8). Discussion In their original description of the PPFP fusion protein, Kroll et al. (11) used transiently transfected U2OS (osteosarcoma) cells to show that PPFP does not activate PPAR-responsive promoters. Instead, PPFP functioned as a dominant-negative inhibitor of PPARg-induced reporter gene activation. This led to the hypothesis that inhibition of endogenous PPARg is an important mechanism by which PPFP causes follicular carcinomas to develop or progress. A number of other observations would also be consistent with this hypothesis. There are no thyroid cancer cell lines that express PPFP; however, a PPARg agonist caused growth inhibition of thyroid cancer cell lines Fig. 7. Regulation of the ENO3 and PGF promoters. A, JEG-3 cells were cotransfected with an ENO3-luciferase plasmid, an internal control Renilla luciferase plasmid, and an expression vector for PAX8, PPARg, PPFP, or empty vector.the cells were cultured Fciglitazone for 2 days before determining reporter gene activities. Luciferase is normalized to Renilla luciferase, and for each experimentthe normalized activity with empty vector ciglitazone is defined as1. Columns, mean for four experiments; bars, SE. B, similar to (A), exceptthe reporter plasmid was driven by the PGF promoter and the transfection was in primary cultures of dog thyrocytes. Columns, mean for six experiments; bars, SE
9 Profiles of PAX8-PPARG Follicular Carcinoma Table 3. Pathways enriched in follicular carcinomas with the PAX8-PPARG translocation Annotation source Term/pathway Total number of genes with term Number out of 761 selected genes P GO Fatty acid metabolism E 05 Fatty acid h-oxidation E 05 Transport E 05 Amino acid transport E 04 Oxidoreductase activity E 04 GenMAPP Mitochondrial fatty acid h oxidation E 06 Glycogen metabolism E 05 Fatty acid degradation E 04 Glycolysis and gluconeogenesis E 03 KEGG Fatty acid biosynthesis (path 2) E 05 Fatty acid metabolism E 04 Valine, leucine, and isoleucine degradation E 04 Benzoate degradation via hydroxylation E 03 Glycolysis/Gluconeogenesis E 03 Propanoate metabolism E 03 h-alanine metabolism E 03 Pentose phosphate pathway E 03 NOTE: Results from testing 977 selected probe sets for enriched terms. The probe sets were collapsed to 761genes (out of 12,744 total genes). Five of 4,194 Gene Ontology terms tested gave P < (Fisher s exact test), of which we estimated 0.63 to be false positives by a Monte Carlo simulation described in Materials and Methods. Similarly, testing 59 GenMAPP maps and147 pathways from KEGG gave four and eight terms with P < 0.01, with estimated numbers of false-positive terms of 0.38 and 0.81, respectively. Details are provided as Supplementary Data. that express PPARg (24 26), and forced overexpression of PPARg reduced cell growth in a thyroid cancer cell line that does not express endogenous PPARg (24). Stable transfection of the SV40 T antigen-transformed human thyroid-derived cell line Nthy-ori with PPFP led to increased growth in soft agar, and treatment of Nthy-ori cells with a PPARg antagonist had a similar effect (12). Comparing the gene expression profiles of follicular carcinomas that express PPFP versus those that lack the translocation should provide insight into the mechanism of action of this fusion protein. Although the data summarized above would suggest that PPFP functions by inhibiting thyroid PPARg, two of the genes most strongly induced in PPFP(+) follicular carcinomas, AQP7 and ANGPTL4, are induced by PPARg in other tissues (22, 23) and a PPAR response element has been identified in the mouse AQP7 promoter. Thus, we considered the hypothesis that PPFP might function in a PPARg-like manner, at least on some target genes. To test this hypothesis, we did a reporter gene analysis of AQP7 and ANGPTL4 promoterluciferase constructs cotransfected with PPFP, PPARg, PAX8, or empty vector. The promoters of two additional genes induced in the PPFP(+) follicular carcinomas, ENO3 and PGF, were also evaluated. The major conclusions from these experiments are that PPFP can indeed function in a PPARg-like manner, although it also has transcriptional properties distinct from either PAX8 or PPARg, and the effects of this fusion protein can be cell type dependent. Thus, the concept of PPFP contributing to follicular carcinoma largely by antagonizing endogenous PPARg needs reevaluation. Endogenous PPARg is expressed at very low levels in thyrocytes and has no known function in these cells. Furthermore, there are other ways of blocking endogenous PPARg, yet follicular carcinomas have not been reported to develop these mechanisms. For example, germ line dominantnegative mutations in the ligand-binding domain of PPARg cause severe insulin resistance (27), yet similar mutations have not been described in follicular carcinomas. Because PPFP is a transcription factor, it likely regulates a number of genes that contribute to the phenotype of follicular carcinoma. PGF and ANGPTL4 are angiogenic factors that are overexpressed in nonthyroid malignancies (28 30), and hence could be important in the development or progression of PPFP(+) follicular carcinoma. PGF also is induced in the thyroid glands of patients with Graves disease and rats treated with thiouracil, suggesting that it participates in the physiologic vascular response that accompanies goiter formation (31). The modest induction of the PGF promoter by PAX8 in our transfection studies may in fact be part of the mechanism of this physiologic response, and suggests that PPFP also may have PAX8-like activity on some target genes. AQP7 is a glycerol channel (32, 33) and could provide energy for follicular carcinomas, especially because our microarray data indicate follicular carcinomas express glycerol kinase (as does normal thyroid; ref. 34). The gene expression signature of PPFP(+) follicular carcinomas was most significantly enriched in pathways related to fatty acid h oxidation and metabolism, with significant enrichments also in pathways related to amino acid and carbohydrate metabolism (Table 2). This is striking considering the known roles of PPARg in lipid and carbohydrate metabolism. Interestingly, however, it is primarily PPARa that stimulates fatty acid h oxidation (35), suggesting that PPFP also has PPARa-like activity. PPARs a and g bind to the same response elements, and the factors that dictate their distinct activities are not fully defined
10 Fig. 8. EGFR-related protein interaction network for up-regulated genes in PPFP(+) versus PPFP( ) follicular carcinomas. A protein interaction network was generated by Oncomine illustrating physical interactions between the EGFR and other up-regulated genes in PPFP(+) follicular carcinomas, based up on the Human Protein Reference Database (21). Proteins with direct physical interactions are represented as nodes connected by lines. EPHA2, PTK2, EGFR, and BRAF are indicated by enlarged nodes to signify that they are known oncoproteins. Oncomine and PicTar revealed a striking enrichment of putative mirna target genes among the up-regulated genes in PPFP(+) follicular carcinomas. Included among these are genes plausibly connected to cancer, such as SS18, RUNX1, PUM2, and NRP2. Because mirnas down-regulate gene expression, these data suggest that certain mirnas are under expressed in PPFP(+) follicular carcinoma, which would be consistent with the observed decrease in mirna expression in other cancers (36). Recently, mirna expression profiles have been shown to accurately predict subtypes of acute lymphoblastic leukemia (36). Our analysis suggests that mirna profiling also might distinguish subtypes of follicular thyroid cancer. Interactome analysis revealed that the PPFP(+) follicular carcinoma gene signature includes EGFR and numerous proteins that interact directly with EGFR. In addition, EGFR is a central node in a large network of interacting proteins with increased transcript expression in PPFP(+) follicular carcinomas (Fig. 8). This suggests that inhibition of EGFR could have multiple beneficial downstream effects, making it a potentially attractive target in PPFP(+) follicular carcinomas. Inhibitors of the EGFR tyrosine kinase currently are in use to treat other cancers (37). The potential value of EGFR inhibitors is but one link to clinical oncologic practice provided by our data. We also show increased expression of the angiogenic factors ANGPTL4 and PGF, suggesting that antiangiogenic agents may be therapeutic. Diagnostically, increased expression of PPARg would almost always indicate follicular thyroid cancer because we found that high expression of PPARg uniformly signifies expression of PPFP, which is rarely found in benign lesions. Two recent reports have examined gene expression in PPFP(+) versus PPFP( ) follicular carcinomas, although neither study provided functional validation of their findings. Lui et al. (38) did Affymetrix profiling of four follicular carcinomas with the rearrangement and five without. For unknown reasons, there is virtually no overlap between their PPFP-specific gene signature and ours. While our manuscript was in preparation, Lacroix et al. (39) published gene expression profiling of four follicular tumors containing PPFP (one of which was benign) and eight follicular tumors without the fusion protein (three of which were benign). Our PPFP signature overlaps substantially with theirs, including all four genes that we studied by transfection (AQP7, ANGPTL4, ENO3, and PGF), as well as the enrichments in pathways related to lipid, glucose, and amino acid metabolism. In addition, Lacroix et al. used computational methods to identify putative PPAR binding sites in the promoters of many genes in their PPFP(+) profile. The concordance of our findings, which were done on different platforms, substantiates our conclusions but does not provide insight into the discrepant results of Lui et al. It remains to be determined which PPFP target genes are important in the development and progression of follicular carcinoma. Acknowledgments We thank the many current and former technicians of the Tissue Procurement Service of the University of Michigan Comprehensive Cancer Center, Barbara Lamb for technical assistance with the DNA microarray analysis, Nancy McAnsh for immunohistochemical expertise, Mary Hong for organizational assistance, and Dr. Sissy M. Jhiang (Department of Physiology and Cell Biology, Ohio StateUniversity, Columbus, Ohio) for providing NIS-luc. References 1. Kimura ET, Nikiforova MN, Zhu Z, KnaufJA, Nikiforov YE, Fagin JA. High prevalence of BRAF mutations in thyroid cancer: genetic evidence for constitutive activation of the RET/PTC-RAS-BRAF signaling pathway in papillary thyroid carcinoma. Cancer Res 2003;63: 1454 ^ Melillo RM, Castellone MD, GuarinoV, et al.the RET/ PTC-RAS-BRAF linear signaling cascade mediates the motile and mitogenic phenotype of thyroid cancer cells. JClin Invest2005;115:1068^ Giordano TJ, Kuick R, Thomas DG, etal. Molecular classification of papillary thyroid carcinoma: distinct BRAF, RAS, and RET/PTC mutation-specific gene expression profiles discovered by DNA microarray analysis. Oncogene 2005;24:6646^ Nikiforova MN, Lynch RA, Biddinger PW, etal. RAS pointmutations and PAX8-PPAR g rearrangementin thyroid tumors: evidence for distinct molecular pathways in thyroid follicular carcinoma. J Clin Endocrinol Metab 2003;88:2318^ Kroll TG, Sarraf P, Pecciarini L, etal. PAX8-PPAR g 1 fusion in oncogene human thyroid carcinoma. Science 2000;289:1357^ Maximo V, BotelhoT, Capela J, et al. Somatic and germline mutation in GRIM-19, a dual function gene involved in mitochondrial metabolism and cell death, is linked to mitochondrion-rich (Hurthle cell) tumours of the thyroid. BrJCancer 2005;92:1892^8. 7. Brun RP, Spiegelman BM. PPAR g and the molecular controlofadipogenesis. JEndocrinol1997;155:217 ^ Chawla A, Schwarz EJ, Dimaculangan DD, Lazar MA. Peroxisome proliferator-activated receptor (PPAR) g: adipose-predominantexpression and induction early in adipocyte differentiation. Endocrinology 1994;135:798 ^ Lazar MA. PPAR g, 10 years later. Biochimie 2005; 87:9 ^ Picard F, Auwerx J. PPARg and glucose homeostasis. Annu Rev Nutr 2002;22:167^ Kroll TG, Sarraf P, Pecciarini L, etal. PAX8-1fusion oncogene in human thyroid carcinoma [corrected]. Science 2000;289:1357^ Powell JG, Wang X, Allard BL, etal. The PAX8/ PPARg fusion oncoprotein transforms immortalized human thyrocytes through a mechanism probably involving wild-type PPARg inhibition. Oncogene 2004; 23:3634 ^ Giordano TJ, Shedden KA, Schwartz DR, et al
PAX8-PPARγ Fusion Protein in thyroid carcinoma
 Sidney H. Ingbar Lecture, ATA 10/17/2013: PAX8-PPARγ Fusion Protein in thyroid carcinoma Ronald J. Koenig, MD, PhD Division of Metabolism, Endocrinology & Diabetes University of Michigan Learning Objectives
Sidney H. Ingbar Lecture, ATA 10/17/2013: PAX8-PPARγ Fusion Protein in thyroid carcinoma Ronald J. Koenig, MD, PhD Division of Metabolism, Endocrinology & Diabetes University of Michigan Learning Objectives
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital. NSW Health Pathology University of Sydney
 Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information
 Supporting Information Franco et al. 10.1073/pnas.1015557108 SI Materials and Methods Drug Administration. PD352901 was dissolved in 0.5% (wt/vol) hydroxyl-propyl-methylcellulose, 0.2% (vol/vol) Tween
Supporting Information Franco et al. 10.1073/pnas.1015557108 SI Materials and Methods Drug Administration. PD352901 was dissolved in 0.5% (wt/vol) hydroxyl-propyl-methylcellulose, 0.2% (vol/vol) Tween
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB
 THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB CHARACTERISTIC OF THE IDEAL TUMOR MARKER Specific Sensitive Easy to perform Easy to interpret Adaptable to FNA Reasonable cost (CHEAP) THYROID TUMOR MARKERS
THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB CHARACTERISTIC OF THE IDEAL TUMOR MARKER Specific Sensitive Easy to perform Easy to interpret Adaptable to FNA Reasonable cost (CHEAP) THYROID TUMOR MARKERS
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643
 Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
April 5, :45 AM 1:45 PM MARRIOTT MARQUIS HOTEL & MARINA, MIRAMAR VIGNETTE 1 VIGNETTE 2 VIGNETTE 3* VIGNETTE 4* VIGNETTE 5*
 April 5, 2016 11:45 AM 1:45 PM MARRIOTT MARQUIS HOTEL & MARINA, MIRAMAR CHAIR: DANNY A. MILNER, JR., BRIGHAM & WOMEN S HOSPITAL, BOSTON, MA VIGNETTE 1 VIGNETTE 2 VIGNETTE 3* VIGNETTE 4* VIGNETTE 5* *VIGNETTES
April 5, 2016 11:45 AM 1:45 PM MARRIOTT MARQUIS HOTEL & MARINA, MIRAMAR CHAIR: DANNY A. MILNER, JR., BRIGHAM & WOMEN S HOSPITAL, BOSTON, MA VIGNETTE 1 VIGNETTE 2 VIGNETTE 3* VIGNETTE 4* VIGNETTE 5* *VIGNETTES
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Data Sheet. Notch Pathway Reporter Kit Catalog # 60509
 Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
microrna Presented for: Presented by: Date:
 microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Histogram showing hybridization signals for chicken (left) and quail (right) genomic DNA analyzed by Chicken GeneChip (n=3). www.nature.com/nature 1 Supplementary Figure 2. Independent
Supplementary Figure 1. Histogram showing hybridization signals for chicken (left) and quail (right) genomic DNA analyzed by Chicken GeneChip (n=3). www.nature.com/nature 1 Supplementary Figure 2. Independent
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies
 Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
number Done by Corrected by Doctor Maha Shomaf
 number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events
 Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events Sandrine Rorive, M.D., PhD. Erasme Hospital - Université Libre de Bruxelles (ULB) INTRODUCTION The assessment of thyroid nodules
Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events Sandrine Rorive, M.D., PhD. Erasme Hospital - Université Libre de Bruxelles (ULB) INTRODUCTION The assessment of thyroid nodules
Insulin Resistance. Biol 405 Molecular Medicine
 Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014
 Policy Number 2.04.82 Molecular Markers in Fine Needle Aspirates of the Thyroid Medical Policy Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014 Disclaimer Our medical policies
Policy Number 2.04.82 Molecular Markers in Fine Needle Aspirates of the Thyroid Medical Policy Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014 Disclaimer Our medical policies
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression
 EXPERIMENTAL AND THERAPEUTIC MEDICINE 7: 1757-1761, 2014 mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression ZHI YONG SHEN *, ZI ZHEN ZHANG *, HUA LIU, EN HAO ZHAO
EXPERIMENTAL AND THERAPEUTIC MEDICINE 7: 1757-1761, 2014 mir 375 inhibits the proliferation of gastric cancer cells by repressing ERBB2 expression ZHI YONG SHEN *, ZI ZHEN ZHANG *, HUA LIU, EN HAO ZHAO
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
Cell Culture. The human thyroid follicular carcinoma cell lines FTC-238, FTC-236 and FTC-
 Supplemental material and methods Reagents. Hydralazine was purchased from Sigma-Aldrich. Cell Culture. The human thyroid follicular carcinoma cell lines FTC-238, FTC-236 and FTC- 133, human thyroid medullary
Supplemental material and methods Reagents. Hydralazine was purchased from Sigma-Aldrich. Cell Culture. The human thyroid follicular carcinoma cell lines FTC-238, FTC-236 and FTC- 133, human thyroid medullary
II. The effect of strain and IGF-1 gene transfer using SMGT techniques on some semen characteristics:-
 The experimental work of this study was carried out in poultry farm of the Department of Animal Production, Faculty of Agriculture, Benha University, Egypt with cooperation of Genetic Engineering and Biotechnology
The experimental work of this study was carried out in poultry farm of the Department of Animal Production, Faculty of Agriculture, Benha University, Egypt with cooperation of Genetic Engineering and Biotechnology
Dr Rodney Itaki Lecturer Anatomical Pathology Discipline. University of Papua New Guinea School of Medicine & Health Sciences Division of Pathology
 Neoplasia Dr Rodney Itaki Lecturer Anatomical Pathology Discipline University of Papua New Guinea School of Medicine & Health Sciences Division of Pathology General Considerations Overview: Neoplasia uncontrolled,
Neoplasia Dr Rodney Itaki Lecturer Anatomical Pathology Discipline University of Papua New Guinea School of Medicine & Health Sciences Division of Pathology General Considerations Overview: Neoplasia uncontrolled,
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis
 5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer
 Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Microarray analysis of differentially expressed genes in the liver between Bai er layers and broilers
 Microarray analysis of differentially expressed genes in the liver between Bai er layers and broilers Q.-G. Wang 1,2,4 *, H.-F. Zhang 1,2,3 *, S.-Z. Wang 1,2,3, G.L. Gao 1,2,4, L. Leng 1,2,3, W. Na 1,2,3
Microarray analysis of differentially expressed genes in the liver between Bai er layers and broilers Q.-G. Wang 1,2,4 *, H.-F. Zhang 1,2,3 *, S.-Z. Wang 1,2,3, G.L. Gao 1,2,4, L. Leng 1,2,3, W. Na 1,2,3
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Examination of Mechanisms of Hepatotoxicity of Anti-diabetic PPARγ Agonists Using Applied Biosystems Rat Whole Genome Microarrays
 Examination of Mechanisms of Hepatotoxicity of Anti-diabetic PPARγ Agonists Using Applied Biosystems Rat Whole Genome Microarrays Lu Zhang b, Lei Guo a, Leming Shi a, Weida Tong a, Yongming Sun b, Gary
Examination of Mechanisms of Hepatotoxicity of Anti-diabetic PPARγ Agonists Using Applied Biosystems Rat Whole Genome Microarrays Lu Zhang b, Lei Guo a, Leming Shi a, Weida Tong a, Yongming Sun b, Gary
Supplementary Figure 1:
 Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
Supplementary Figure 1: (A) Whole aortic cross-sections stained with Hematoxylin and Eosin (H&E), 7 days after porcine-pancreatic-elastase (PPE)-induced AAA compared to untreated, healthy control aortas
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Oncogenes/Growth Factors & Environment
 Oncogenes/Growth Factors & Environment 8 th Postgraduate Course in Endocrine Surgery Crete, Greece September, 2006 Orlo H. Clark M.D. Thyroid Cancer Thyroid cancer is the 8 th most common and most rapidly
Oncogenes/Growth Factors & Environment 8 th Postgraduate Course in Endocrine Surgery Crete, Greece September, 2006 Orlo H. Clark M.D. Thyroid Cancer Thyroid cancer is the 8 th most common and most rapidly
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Integration Of Metabolism
 Integration Of Metabolism Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP, NADPH, and building blocks for biosyntheses. 1. ATP is the universal
Integration Of Metabolism Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP, NADPH, and building blocks for biosyntheses. 1. ATP is the universal
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Genetics of Cancer Lecture 32 Cancer II. Prof. Bevin Engelward, MIT Biological Engineering Department
 Genetics of Cancer Lecture 32 Cancer II rof. Bevin Engelward, MIT Biological Engineering Department Why Cancer Matters New Cancer Cases in 1997 Cancer Deaths in 1997 Genetics of Cancer: Today: What types
Genetics of Cancer Lecture 32 Cancer II rof. Bevin Engelward, MIT Biological Engineering Department Why Cancer Matters New Cancer Cases in 1997 Cancer Deaths in 1997 Genetics of Cancer: Today: What types
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung
 Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
TITLE: Investigating the Role of TBX2 in the Inhibition of Senescence in Prostate Cancer
 AD Award Number: W81XWH-07-1-0155 TITLE: Investigating the Role of TBX2 in the Inhibition of Senescence in Prostate Cancer PRINCIPAL INVESTIGATOR: Srinivas Nandana CONTRACTING ORGANIZATION: Vanderbilt
AD Award Number: W81XWH-07-1-0155 TITLE: Investigating the Role of TBX2 in the Inhibition of Senescence in Prostate Cancer PRINCIPAL INVESTIGATOR: Srinivas Nandana CONTRACTING ORGANIZATION: Vanderbilt
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Personalized Medicine: Lung Biopsy and Tumor
 Personalized Medicine: Lung Biopsy and Tumor Mutation Testing Elizabeth H. Moore, MD Personalized Medicine: Lung Biopsy and Tumor Mutation Testing Genomic testing has resulted in a paradigm shift in the
Personalized Medicine: Lung Biopsy and Tumor Mutation Testing Elizabeth H. Moore, MD Personalized Medicine: Lung Biopsy and Tumor Mutation Testing Genomic testing has resulted in a paradigm shift in the
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles
 Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals
 Br. J. Anaesth. (1981), 53, 131 THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals J. C. STANLEY In this paper, the glucose-fatty acid cycle
Br. J. Anaesth. (1981), 53, 131 THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals J. C. STANLEY In this paper, the glucose-fatty acid cycle
Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary Materials and Methods
 Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Resistance to Thyroid Hormone and Down Syndrome: Coincidental Association or. Genetic Linkage?
 Page 1 of 6 1 Resistance to Hormone and Down Syndrome: Coincidental Association or Genetic Linkage? (doi: 10.1089/thy.2011-0316) Resistance to Hormone and Down Syndrome: Coincidental Association or Genetic
Page 1 of 6 1 Resistance to Hormone and Down Syndrome: Coincidental Association or Genetic Linkage? (doi: 10.1089/thy.2011-0316) Resistance to Hormone and Down Syndrome: Coincidental Association or Genetic
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Loss of succinate dehydrogenase activity results in dependency on pyruvate carboxylation for cellular anabolism
 Loss of succinate dehydrogenase activity results in dependency on pyruvate carboxylation for cellular anabolism Charlotte Lussey-Lepoutre, Kate ER Hollinshead, Christian Ludwig, Mélanie Menara, Aurélie
Loss of succinate dehydrogenase activity results in dependency on pyruvate carboxylation for cellular anabolism Charlotte Lussey-Lepoutre, Kate ER Hollinshead, Christian Ludwig, Mélanie Menara, Aurélie
Erzsebet Kokovay, Susan Goderie, Yue Wang, Steve Lotz, Gang Lin, Yu Sun, Badrinath Roysam, Qin Shen,
 Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of
 Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of 1,000 most differentially expressed genes with NKX2-1 amplification in lung adenocarcinoma cell lines and anti-correlated
Supplemental Figure S1. A. Venn diagram depicting overlap between anti-correlated genes of 1,000 most differentially expressed genes with NKX2-1 amplification in lung adenocarcinoma cell lines and anti-correlated
Supplementary Appendix
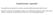 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Objectives. Morphology and IHC. Flow and Cyto FISH. Testing for Heme Malignancies 3/20/2013
 Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
Carcinoma midollare tiroideo familiare
 12 AME Italian Meeting 6 Joint Meeting with AACE Carcinoma midollare tiroideo familiare Profilo genetico e stratificazione del rischio Maria Chiara Zatelli Sezione di Endocrinologia Dipartimento di Scienze
12 AME Italian Meeting 6 Joint Meeting with AACE Carcinoma midollare tiroideo familiare Profilo genetico e stratificazione del rischio Maria Chiara Zatelli Sezione di Endocrinologia Dipartimento di Scienze
THYROID CYTOLOGY THYROID CYTOLOGY FINE-NEEDLE-ASPIRATION ANCILLARY TESTS IN THYROID FNA
 ANCILLARY TESTS IN THYROID FNA Prof. Fernando Schmitt Department of Pathology and Oncology, Medical Faculty of Porto University Head of Molecular Pathology Unit, IPATIMUP General-Secretary of the International
ANCILLARY TESTS IN THYROID FNA Prof. Fernando Schmitt Department of Pathology and Oncology, Medical Faculty of Porto University Head of Molecular Pathology Unit, IPATIMUP General-Secretary of the International
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
Evaluation and Management of Thyroid Nodules. Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada
 Evaluation and Management of Thyroid Nodules Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada Disclosure Consulting Amgen Speaking Amgen Objectives Understand the significance of incidental
Evaluation and Management of Thyroid Nodules Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada Disclosure Consulting Amgen Speaking Amgen Objectives Understand the significance of incidental
Guangdong Medical University, Zhanjiang, China; 5 Guangxi Medical University, Nanning, China; 6 Department of Pathology, University of Michigan
 Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer Jie Zhang 1,2, Shumei Feng 3, Wenmei Su 4, Shengbin
Lecture 29: Membrane Transport and metabolism
 Chem*3560 Lecture 29: Membrane Transport and metabolism Insulin controls glucose uptake Adipose tissue and muscles contain a passive glucose transporter GluT4 which takes up glucose from blood. (This is
Chem*3560 Lecture 29: Membrane Transport and metabolism Insulin controls glucose uptake Adipose tissue and muscles contain a passive glucose transporter GluT4 which takes up glucose from blood. (This is
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Systems biology approaches and pathway tools for investigating cardiovascular disease
 Systems biology approaches and pathway tools for investigating cardiovascular disease Craig E. Wheelock 1,2*, Åsa M. Wheelock 2,3,4, Shuichi Kawashima 5, Diego Diez 2, Minoru Kanehisa 2,5, Marjan van Erk
Systems biology approaches and pathway tools for investigating cardiovascular disease Craig E. Wheelock 1,2*, Åsa M. Wheelock 2,3,4, Shuichi Kawashima 5, Diego Diez 2, Minoru Kanehisa 2,5, Marjan van Erk
Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4
 Khozoie et al. BMC Genomics 2012, 13:665 RESEARCH ARTICLE Open Access Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4 Combiz Khozoie
Khozoie et al. BMC Genomics 2012, 13:665 RESEARCH ARTICLE Open Access Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4 Combiz Khozoie
