TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder
|
|
|
- Catherine Campbell
- 5 years ago
- Views:
Transcription
1 TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder Jian Chen, Zhi-Xing Yao, Jiun-Sheng Chen, Young Jin Gi, Nina M. Muñoz, Suchin Kundra, H. Franklin Herlong, Yun Seong Jeong, Alexei Goltsov, Kazufumi Ohshiro, Nipun A. Mistry, Jianping Zhang, Xiaoping Su, Sanaa Choufani, Abhisek Mitra, Shulin Li, Bibhuti Mishra, Jon White, Asif Rashid, Alan Yaoqi Wang, Milind Javle, Marta Davila, Peter Michaely, Rosanna Weksberg, Wayne L. Hofstetter, Milton J Finegold, Jerry W. Shay, Keigo Machida, Hidekazu Tsukamoto, and Lopa Mishra Correspondence to: Lopa Mishra, MD Director, Center for Translational Research Department of Surgery and GW Cancer Center, George Washington University, 2150 Pennsylvania Avenue, NW, Washington, DC & The Del & Dennis McCarthy Distinguished Professor in Gastrointestinal Cancer Research, Department of Gastroenterology, Hepatology, and Nutrition Unit 1466 The University of Texas MD Anderson Cancer Center 1515 Holcombe Boulevard, Houston, TX lmishra@mdanderson.org, lmishra@gwu.edu Tel: ,
2 Supplemental Materials and Methods Whole-transcriptome Sequencing Analysis. Whole-transcriptome RNA sequencing were performed and analyzed at the MD Anderson Cancer Center DNA core facility. We generated a standard bam file by aligning the raw paired-end reads in FASTQ format to the human reference genome, GRCh37/hg19, using MOSAIK alignment software (1). We then counted the uniquely mapped reads in genomic features such as genes annotated in refseq to generate the raw counts for each gene, where raw reads count data were subsequently normalized across samples using DESeq with variance stabilization (2). Database analyses. Affymetrix mrna microarray data GSE3228 were downloaded from NCBI s Gene Expression Omnibus (GEO). Smad3 / primary keratinocytes were derived from Smad3 null mice. The transcriptome data of two independent Smad3 / keratinocytes were downloaded and further normalized with wild type mouse keratinocytes. Most of the decreased TGF-β/Smad3- regulated genes presented on Figure 2A are validated by IPA analysis (cutoff: adjusted P value < 0.05) in Smad3 / primary keratinocytes. The information about somatic mutations in SPTBN1 and SMAD3 reported in the Catalogue of s in Cancer and The Cancer Genome Atlas (TCGA) liver cancers was downloaded in May 2014 (Supplemental Table 3). Preparation of Nuclear and Cytoplasmic Proteins. Nuclear and cytoplasmic proteins were prepared as previously described (3). Briefly, the cells were harvested and incubated in buffer A (10 mmol/l Hepes, ph 7.8, 10 mmol/l KCl, 0.1 mmol/l EDTA, 1 mmol/l dithiothreitol, 2 mg/ml aprotinin, 0.5 mmol/l phenylmethyl sulfonyl fluoride, and 0.5% Triton X-100). After centrifugation at 5000 rpm, buffer A was collected, 2
3 including cytoplasmic proteins, as the cytoplasmic protein sample. Buffer C (50 mmol/l Hepes, ph 7.8, 420 mmol/l KCl, 0.1 mmol/l EDTA, 5 mmol/l MgCl2, 10% glycerol, 1 mmol/l dithiothreitol, 2 mg/ml aprotinin, and 0.5 mmol/l phenylmethyl sulfonyl fluoride) was added to the pellet. After rotation for 30 minutes and centrifugation at 15,000 rpm, supernatants were collected as the nuclear protein sample. Then, lysates were immunoprecipitated with the indicated antibodies. The following antibodies were used for immunoblotting and immunoprecipitation analyses: V5 antibody (R961-25, Invitrogen), Flag M2 antibody (Sigma Aldrich, F3165), His antibody (2366, Cell Signaling), Actin antibody (A2066, Sigma Aldrich), Tubulin antibody (T8328, Sigma Aldrich), c-myc antibody (sc-40, Santa Cruz Biotechnology), CTCF antibody (2899, Cell Signaling), Smad3 antibody (9523, Cell Signaling), phosphor-smad3 antibody (Ser423/425, 9520, Cell Signaling), Smad2 antibody (3122, Cell Signaling), phosphor- Smad2 antibody (Ser465/467, 8828, Cell Signaling), β2sp antibody (customized antibody from biosythesis). Primers for Quantitative RT-PCR Analyses and ChIP Assays. The sequences used for quantitative RT-PCR analysis were as follows: htert, forward 5 - AAATGCGGCCCCTGTTTCT-3 ; reverse 5 - CAGTGCGTCTTGAGGAGCA-3. mtert, forward 5 - TCTACCGCACTTTGGTTGCC-3 ; reverse 5 - CAGCACGTTTCTCTCGTTGC-3. IGF2, forward 5 - GTGGCATCGTTGAGGAGTG-3 ; reverse 5 - CACGTCCCTCTCGGACTTG-3. CTCF, forward 5 - CAGTGGAGAATTGGTTCGGCAT-3 ; reverse 5 - CTGGCGTAATCGCACATGGA-3. Smad3, forward 5 - TGGACGCAGGTTCTCCAAAC-3 ; reverse 5 - CCGGCTCGCAGTAGGTAAC-3. m HEY1, forward 5 - CCGACGAGACCGAATCAATAAC-3 ; reverse 5 - TCAGGTGATCCACAGTCATCTG-3. 3
4 mesr1, forward 5 - CCCGCCTTCTACAGGTCTAAT-3 ; reverse 5 - CTTTCTCGTTACTGCTGGACAG-3. mrunx3, forward 5 - GACTCCTTCCCCAACTATACACC-3 ; reverse 5 - CGCTGTTCTCGCCCATCTT-3. mslc1a2, forward 5 - GCCAACAATATGCCCAAGCAG-3 ; reverse 5 - GACACCAAACACAGTCAGTGA-3. hrunx1, forward 5 - CTGCCCATCGCTTTCAAGGT- 3 ; reverse 5 - GCCGAGTAGTTTTCATCATTGCC-3. hcdkn1a, forward 5 - TGTCCGTCAGAACCCATGC-3 ; reverse 5 -AAAGTCGAAGTTCCATCGCTC-3. hcdkn1b, forward 5 -AACGTGCGAGTGTCTAACGG-3 ; reverse 5 - CCCTCTAGGGGTTTGTGATTCT-3. hserpine1, forward 5 - ACCGCAACGTGGTTTTCTCA-3 ; reverse 5 - TTGAATCCCATAGCTGCTTGAAT-3. hirf7, forward 5 - GCTGGACGTGACCATCATGTA-3 ; reverse 5 - GGGCCGTATAGGAACGTGC-3. hfos, forward 5 -CCGGGGATAGCCTCTCTTACT-3 ; reverse 5 -CCAGGTCCGTGCAGAAGTC-3. htgif1, forward 5 - GGGATTGGCTGTATGAGCACC-3 ; reverse 5 - GGCGGGAAATTGTGAACTGA-3. hmmp9, forward 5 - TGTACCGCTATGGTTACACTCG-3 ; reverse 5 - GGCAGGGACAGTTGCTTCT-3. hmyc, forward 5 -GGCTCCTGGCAAAAGGTCA -3 ; reverse 5 - CTGCGTAGTTGTGCTGATGT-3. Gapdh, forward 5 -TGCACCACCAACTGCTTAGC-3 ; Gapdh, reverse 5 -GTCTTCTGGGTGGCAGTGATG-3 5. halb, forward 5 -TGCAACTCTTCGTGAAACCTATG-3 ; halb, reverse 5 -ACATCAACCTCTGGTCTCACC-3 ; malb, forward 5 -CAAGAGTGAGATCGCCCATCG-3 ; 4
5 malb, reverse 5 -TTACTTCCTGCACTAATTTGGCA-3 ; hkrt19, forward 5 -AACGGCGAGCTAGAGGTGA-3 ; hkrt19, reverse 5 -GGATGGTCGTGTAGTAGTGGC-3 ; mkrt19, forward 5 -GTTCAGTACGCATTGGGTCAG-3 ; mkrt19, reverse 5 -GAGGACGAGGTCACGAAGC-3 ; hprom1(cd133, forward 5 -AGTCGGAAACTGGCAGATAGC-3 ; hprom1(cd133), reverse 5 -GGTAGTGTTGTACTGGGCCAAT-3 ; mprom1(cd133, forward 5 -ACTGGGGCTGTGTGGAAAG-3 ; mprom1(cd133), reverse 5 -GCATTGAAGGTATCTTGGGTCTC-3 ; hnanog, forward 5 -TTTGTGGGCCTGAAGAAAACT-3 ; hnanog, reverse 5 -AGGGCTGTCCTGAATAAGCAG-3 ; mnanog, forward 5 -CACAGTTTGCCTAGTTCTGAGG-3 ; mnanog, reverse 5 -GCAAGAATAGTTCTCGGGATGAA-3 ; hpou5f1(oct4), forward 5 -CTGGGTTGATCCTCGGACCT-3 ; hpou5f1(oct4), reverse 5 -CCATCGGAGTTGCTCTCCA-3 ; mpou5f1(oct4), forward 5 -AGAGGATCACCTTGGGGTACA-3 ; mpou5f1(oct4), reverse 5 -CGAAGCGACAGATGGTGGTC-3 ; hsox2, forward 5 -GCCGAGTGGAAACTTTTGTCG-3 ; hsox2, reverse 5 -GGCAGCGTGTACTTATCCTTCT-3 ; msox2, forward 5 -GCGGAGTGGAAACTTTTGTCC-3 ; msox2, reverse 5 -GGGAAGCGTGTACTTATCCTTCT-3 ; hepcam, forward 5 -AATCGTCAATGCCAGTGTACTT-3 ; hepcam, reverse 5 -TCTCATCGCAGTCAGGATCATAA-3 ; 5
6 mepcam, forward 5 -CTGGCGTCTAAATGCTTGGC-3 ; mepcam, reverse 5 -CCTTGTCGGTTCTTCGGACTC-3 ; hafp, forward 5 -CTTTGGGCTGCTCGCTATGA-3 ; hafp, reverse 5 -GCATGTTGATTTAACAAGCTGCT-3 ; mafp, forward 5 -AGCTTCCACGTTAGATTCCTCC-3 ; mafp, reverse 5 -ACAAACTGGGTAAAGGTGATGG-3 ; The sequences used for ChIP assays were as follows (Supplementary Figure S10): htert ChIP-1-F: CTCTGCAGTCCGAGGCTTG; htert ChIP-1-R: CCCGTCATTTCTCTTTGCAG. htert ChIP-2-F: CCGGACCTGGAGGCAGCC; htert ChIP-2-R: CCCCAGCGGAGAGAGGTC. htert ChIP-3-F: ACCTCTCTCCGCTGGGGC; htert ChIP-3-R: CGTCTGTGCCCGCGAATC. htert ChIP-4-F: GGACCGCGCTTCCCACGT; htert ChIP-4-R: AGGAAGGGGAGGGGCTGG. htert ChIP-5-F: AGCCCCTCCCCTTCCTTTCC; htert ChIP-5-R: AGCGCACGGCTCGGCAGC. htert ChIP-6-F: AGCCGTGCGCTCCCTGCT; htert ChIP-6-R: GCACGCACACCAGGCACT. mtert ChIP-1-F: AAATTCAGCAGCCCCTCTCT; mtert ChIP-1-R: TCTCAACTCACGCACCCATA. mtert ChIP-2-F: ATGGTCGCACCACAATAAAG; mtert ChIP-2-R: GGGCAAGCAAAGAAGCCTAT. mtert ChIP-3-F: GGATAGGCTTCTTTGCTTGC; mtert ChIP-3-R: TTGATGGTCACAATGCTGGT. mtert ChIP-4-F: TTCGTCGTGGACTCTCAGTG; mtert ChIP-4-R: GCCACACCTCCCGGTATC. mh19-icr ChIP-4-F: TCATGGGTCACTCAGGCATA; mh19-icr ChIP-4-R: CGTCTGCCGAGCAATATGTA. hpai-1 Chip-F: GTCCTAGGCTTTTTGGGTCA; hpai-1 Chip-R: ACAATTGAGCAAACCCCAAT. mpai-1 6
7 Chip-F: AGGGAACCAGAGTTTGCTCA; mpai-1 Chip-R: GCCCCACCCACTTTCTAACT. Supplemental References 1. Anders S & Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):R Marth GT, et al. (1999) A general approach to single-nucleotide polymorphism discovery. Nature genetics 23(4): Chen J, et al. (2009) Hypoxia-mediated up-regulation of Pim-1 contributes to solid tumor formation. The American journal of pathology 175(1):
8 Supplemental Figure1. Sptbn1 +/ /Smad3 +/ mice phenocopy BWS. Representative enlargement of organs in Sptbn1 +/ /Smad3 +/ mice. I: ears; II: kidneys; III: Tongues; IV: livers; V: brains; VI: hearts. The arrow indicates an anterior linear ear lobe crease. 8
9 Supplemental Figure 2. Validation of TGF-β pathway disruption in Smad3 / primary keratinocytes. (A) Q-PCR analysis of defective TGF-β response in Sptbn1 +/ /Smad3 +/ MEFs (n=2). Error bars are shown as standard deviations. Each result shown is representative of three independent experiments. *: P < 0.01, Student s t-test. (B) The transcriptome data of two independent Smad3 / keratinocytes were downloaded from NCBI s Gene Expression Omnibus (GEO) cohort GSE3228, and further normalized with wild type mouse keratinocytes and then analyzed by IPA. TGF-β/Smad3-regulated genes are detected by fold change cutoff (P < 0.05) and presented on heat map. 9
10 Supplemental Figure 3. Analyses of whole-transcriptome sequencing of three BWS cell lines. (A) The differential gene expressions in three BWS cells were compared to the gene expression in a fibroblast cell line established from a normal person by fold change (significance cutoff, P value < 0.05). (B) Downregulation of the TGF-β pathway in CDKN1C+ BWS cells. The differential gene expression in CDKN1C+ BWS cells was compared to the gene expression in a fibroblast cell line established from normal person and then detected by fold change (cutoff, q value < 0.3). These data were uploaded to the IPA program. The diagram lists all the TGF-β1- related genes in the input dataset. Red indicates upregulated and green indicates downregulated genes. The arrows point to validated TGF-β/Smad3-regulated genes. (C) Q-PCR analysis validated a defective TGF-β response in CDKN1C+ cells. Normal fibroblast and BWS CDKN1C+ cells were treated with TGF-β1 for 2 hours. Q-PCR was performed to detect TGF-β target gene expression. The result shown is representative of three independent experiments. 10
11 11
12 Supplemental Figure 4. β2sp interacts with SMAD3. (A) β2sp interacts with SMAD3, but not with SMAD2. HepG2 cells were treated with 200 pm TGF-β for 2 h. Cell lysates were immunoprecipitated with an anti-smad3 antibody, an anti-smad2 antibody or anti-β2sp antibody, and immunoblotted with indicated antibodies. * designates non-specific bands. (B) Overexpression of β2sp rescues SMAD3 nuclear translocation in BWS cells. BWS KvDMR+ cells were transfected with full-length V5-β2SP for 24 hours and then were treated with 200 pm TGF-β1 for 2 hours. Immunofluorescent staining was performed to detect β2sp and Smad3. Scale bars = 50 μm. Representative confocal images are shown. Data are representative of two independent experiments (A B). (C) Proposed model of the role of β2sp required for Smad3 nuclear translocation in response to TGF-β. 12
13 13
14 Supplemental Figure 5. TGF-β/Smad3/β2SP regulates CTCF levels posttranscriptionally. (A) Lower CTCF levels in BWS cells compared with normal hepatocytes. CTCF levels were detected by immunoblotting analysis. Seven human BWS cell lines were developed by Dr. Weksberg (Ontario, Canada). CDKN1C+: CDKN1C mutation, no tumor; KvDMR-: loss of methylation in KCNQ1OT1, no tumor; KvDMR+: loss of methylation in KCNQ1OT1, hepatoblastoma; UPD+NT: uniparental disomy, no tumor; UPD+T: uniparental disomy, hepatoblastoma; UPD+1: uniparental disomy, tongue tissue 1; UPD+2: uniparental disomy, tongue tissue 2. UPD+1 and UPD+2 cell lines were derived from the same case with biopsies at separate sites. KvDMR- and KvDMR+ were derived from normal monozygotic twin (absence of KvDMR molecular defect but it had some clinical signs of BWS) and BWS monozygotic twin with KvDMR molecular defect, respectively. Normal human hepatocytes were received from the Liver Tissue Cell Distribution System, University of Pittsburgh and University of Minnesota. (B) Knock down Smad3 decreased CTCF protein stability. HepG2-sh-Ctrl or HepG2-sh-Smad3 cells were treated with cycloheximide (CHX; 100 μg/ml) for the indicated times. The density of CTCF and the integrated optical density were measured. The turnover of CTCF is indicated graphically. (C) Smad3-mediated CTCF downregulation was proteasome-dependent. HepG2-sh- Ctrl or HepG2-sh-Smad3 cells were treated with or without 50 µg/ml of MG132 for 6 hours. The cell lysates were then immunoblotted with the indicated antibodies. All blots are representative of experiments performed 3 times (A C). (D, E) CTCF mrna level was not affected by TGF-β1 treatment in MEFs (D) or HepG2 cells (E). The Sptbn1 +/ /Smad3 +/ MEFs (D) or β2spknockdown HepG2 cells (E) were treated with 200 pm TGF-β1 for the indicated times. Q-PCR was performed to detect CTCF mrna expression. (n=3). Each blot is representative of three independent experiments (A, B, D & E), 2(C). 14
15 15
16 Supplemental Figure 6. β2sp/smad3 interact with CTCF in cell nucleus. (A) β2sp and/or Smad3 increased CTCF levels in a TGF-β-dependent manner. HepG2 cells were co-transfected with the indicated plasmids and were treated with TBR1 inhibitor SB (5 um) overnight or with TGF-β1 (200 pm) for 2 h. (B) CTCF interacts with Smad3 MH1 domain. SNU398 cells were co-transfected with indicated plasmids. Cell lysates were immunoprecipitated with an anti- His antibody and immunoblotted with indicated antibodies. (C) Interaction of β2sp/smad3 with CTCF is TGF-β dependent. SNU398 cells were co-transfected with the indicated plasmids and were treated with 5 µm SB overnight. Cell lysates were immunoprecipitated with an anti-histidine antibody and immunoblotted with indicated antibodies. Each blot is representative of two independent experiments (A C). (D) Nuclear translocation of β2sp and CTCF upon treatment of TGF-β. SNU475 cells were treated with 200 pm TGF-β1 for 2 h. Immunofluorescent staining was performed to detect β2sp and CTCF. Scale bars, 20 μm. 16
17 Supplemental Figure 7. The expression of TERT, c-myc and IGF2 are increased in human BWS-associated tumors, TGF-β defective mouse livers and cell lines established from individual tumors in Sptbn1 +/- /Smad3 +/- mice. (A) Increased levels of TERT and c-myc in kidney tumors of BWS patients were observed. Representative immunohistochemical staining of human TERT and c-myc in normal kidney or in kidney tumors of BWS patients. (B) Increased mrna expressions of c-myc in Sptbn1 +/ /Smad3 +/, Smad3 +/, and Sptbn1 +/ mouse livers. (C) Reduction of TGF-β-induced TERT expression levels were observed in human normal fibroblasts but not CDKN1C+ BWS cells. The cells were treated with 200 pm TGF-β for 2 h. (D) The TERT mrna level was increased in β2sp, Smad3 or CTCF knockdown cells compared with control cells. The cells were treated with 200 pm TGF-β for 2 h. Error bars are shown as standard deviations. Each result shown is representative of three independent experiments (B D). *: P < 0.001, one-way ANOVA with post-hoc Bonferroni test 17
18 Supplemental Figure 8. Knock down CTCF or Smad3 in SNU398 cells. SNU398 cells were infected with lentivirus-mediated CTCF-shRNA, Smad3-shRNA or control-shrna. (A) Q-PCR and (B) Western blot analyses were performed to detect the expression of CTCF and Smad3 in SNU398 cells. Results are the average of three independent experiments and are presented as mean ± SD in (A). *: P < 0.01, Student s t-test. 18
19 1
20 Supplemental Figure 9. β2sp, Smad3, and CTCF bind on TERT Promoter Region. (A) Diagrammatic representation shows the potential SBE motifs, CTCF binding motifs, and Myc binding motifs on the human or mouse TERT promoter-exon1 region. (B) TGF-β increases β2sp/smad3 binding activities on the sites ( -335 htert but not htert ) on human TERTpromoter region. Genomic DNA was isolated from normal hepatocytes treated with TGFβ1 (200 pm) for 2 h. (C) TGF-β1 treatment increases β2sp and SMAD3 binding on the TERT promoter in normal hepatocytes but not in BWS KvDMR+T hepatoblastoma cells. The cells were treated with TGF-β1 for 2 h. (D) TGF-β1 increases β2sp/smad3 binding activities on human and mouse PAI-1 promoter. Genomic DNA was isolated from normal hepatocytes and wild type MEFs. ChIP assays were performed and enrichment of β2sp or Smad3 transcripts was measured by quantitative PCR for (B) to (D). (E) TGF-β increases CTCF binding activities on the sites ( GTGCGCCCCCTTTCGTTAT - 278, but not on CCCTC or on CCCTC ) on mouse TERT promoter region. ChIP assays were performed to detect the binding activity of CTCF on the mouse TERT promoter region. Genomic DNA was isolated from wild type MEFs treated with TGF-β1. (F) Binding ability of CTCF on htert promoter region is β2sp dependent. Normal human hepatocytes or SNU398 or BWS KvDMR+T cells were treated with TGF-β1 for 2 h and ChIP assays were performed. Enrichment of CTCF transcripts was measured by quantitative PCR. Error bars are shown as standard deviations. Each result shown is representative of three independent experiments (B-F) *: P < 0.001, one-way ANOVA with post-hoc Bonferroni test. 2
21 Supplemental Figure 10. Increased levels of IGF2 in livers and tumors in Sptbn1 +/ /Smad3 +/ mice. (A) Increased mrna expressions of IGF2 in Sptbn1 +/ /Smad3 +/, Smad3 +/, and Sptbn1 +/ mouse livers. (B) Increased mrna expressions of c-myc, TERT, IGF2, ITFG2 and MMP9 in mouse tumor cell lines. 4 cell lines were established from individual tumors (three hepatocarcinomas and one lymphoma) in Sptbn1 +/ /Smad3 +/ mice. Results are the average of three independent experiments and are presented as mean ± SD (A and B). *: P < 0.001, versus wild type mouse hepatocytes or wild type MEFs, one-way ANOVA with post-hoc Bonferroni test. 21
22 Supplemental Figure 11. Increased mrna expression levels of stemness genes in MEFs and BWS cell lines. Q-PCR was performed to detect gene expression. Error bars are shown as standard deviations. Results are the average of three independent experiments and are presented as mean ± SD. ǂ: P < 0.05, #: P < 0.01, *: P < 0.001, versus wild type MEFs (Student s t-test) or human normal fibroblasts (one-way ANOVA with post-hoc Bonferroni test). 22
23 Supplemental Table 1. Classification of tumors in Sptbn1 +/- /Smad3 +/-, Sptbn1 +/- and Smad3 +/- mice. Site Tumor Categories Sptbn1 +/- /Smad3 +/- Sptbn1 +/- Smad3 +/- No. of mice Incidence (%) No. of mice Incidence (%) No. of mice Incidence (%) Small Intestine Adenocarcinoma Liver Hepatocellular Carcinoma Pancreas Adenocarcinoma Head & Neck Squamous Cell Carcinoma Lung Adenocarcinoma Colon Adenocarcinoma Thymus Thymoma Mediastinal Mass Sarcoma Spleen Lymphoma Kidney Clear Cell Carcinoma Adrenal Adrenocortical Carcinoma Skin Squamous Cell Carcinoma Breast Adenoma Lacrimal Gland Adenocarcinoma Optic Nerve Glioma Total Sptbn1 +/- /Smad3 +/- mice, n=15; total mice Sptbn1 +/- /Smad3 +/- with tumors, n=11; mice with multiple tumors, n=9. Total Sptbn1 +/- mice n=43; total mice Sptbn1 +/- with tumors, n=19; mice with multiple tumors, n=4. Total Smad3 +/- mice n=30; total mice Smad3 +/- with tumors, n=3; mice with multiple tumors n=0.
24 Supplemental Table 2. Sporadic tumor development in Sptbn1 +/- /Smad3 +/-, Sptbn1 +/- and Smad3 +/- mice. Genotype Mouse number Gender Thymoma Head & Neck cancer Glioma Lacrimal gland cancer Skin cancer Adrenocortic al carcinoma Pancreati c cancer Kidney cancer Sarcoma Breast cancer Colon adenoca rcinoma Small Intestine cancer Lymphoma Lung cancer Hepatocellular carcinoma Sptbn1 +/ / Smad3 +/ *: Mice without tumor. 1 F F M M M F M F F M + 11 M * F:2 M:2 Sptbn1 +/ 1 M F M M + 5 M + 7 F + 7 F + 8 F + 9 F + 10 M + 11 F + 12 M + 13 F + 14 M + 15 M + 16 M + 17 F M + 19 F * F;10 M: 14 Smad3 +/ 1 M + 2 F + 3 F * F:14 M:13
25 Supplemental Table 3. mutations of Smad3 and β2sp identified on COSMIC and LIHC TCGA. COSMIC Gene Name Sample ID SMAD p.a349s c.1045g>t SMAD p.v331i c.991g>a AA CDS Primary Tissue Histology Status Autonomic ganglia Neuroblastoma Central nervous system Primitive neuroectodermal tumourmedulloblastoma SMAD p.r268c c.802c>t Endometrium Carcinoma SMAD p.r295w c.883c>t Endometrium Carcinoma SMAD p.v331i c.991g>a Endometrium Carcinoma SMAD p.s425y c.1274c>a Endometrium Carcinoma Haematopoietic SMAD p.y226fs*1 c.676_677insa and lymphoid Lymphoid neoplasm SMAD p.r90h c.269g>a Large intestine Carcinoma SMAD p.e178d c.534g>t Large intestine Carcinoma SMAD p.y237c c.710a>g Large intestine Carcinoma SMAD p.n276i c.827a>t Large intestine Carcinoma SMAD p.r287w c.859c>t Large intestine Carcinoma SMAD p.r368q c.1103g>a Large intestine Carcinoma SMAD p.r373h c.1118g>a Large intestine Carcinoma SMAD p.g379a c.1136g>c Large intestine Carcinoma SMAD p.p393l c.1178c>t Large intestine Carcinoma SMAD p.p393l c.1178c>t Large intestine Carcinoma SMAD p.r142h c.425g>a Liver Carcinoma SMAD p.e239v c.716a>t Lung Carcinoma SMAD p.r243c c.727c>t Lung Carcinoma SMAD p.p253s c.757c>t Lung Carcinoma SMAD p.r268h c.803g>a Lung Carcinoma SMAD p.e149g c.446a>g Oesophagus Carcinoma SMAD p.s423g c.1267a>g Ovary Carcinoma SMAD p.s422f c.1265c>t Pancreas NS SMAD p.a329s c.985g>t Prostate Carcinoma Malignant SMAD p.r373c c.1117c>t Skin melanoma Previously Reported
26 SMAD p.s425c c.1274c>g Upper aerodigestive tract Carcinoma SPTBN p.a682v c.2045c>t Breast Carcinoma SPTBN p.h1760p c.5279a>c Breast Carcinoma SPTBN p.e1646k c.4936g>a Breast Carcinoma SPTBN p.r42w c.124c>t Breast Carcinoma SPTBN p.r107* c.319c>t Breast Carcinoma SPTBN p.a2326t c.6976g>a Endometrium Carcinoma SPTBN p.r1258g c.3772a>g Endometrium Carcinoma SPTBN p.d1813n c.5437g>a Endometrium Carcinoma SPTBN p.e475k c.1423g>a Endometrium Carcinoma SPTBN p.a395t c.1183g>a Endometrium Carcinoma SPTBN p.v1946i c.5836g>a Endometrium Carcinoma SPTBN p.f2236i c.6706t>a Endometrium Carcinoma Haematopoietic SPTBN p.g246s c.736g>a and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.l1973i c.5917c>a and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.t2320m c.6959c>t and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.k497r c.1490a>g and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.k401t c.1202a>c and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.n1226s c.3677a>g and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.v1860m c.5578g>a and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.r681w c.2041c>t and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.f694y c.2081t>a and lymphoid Lymphoid neoplasm Haematopoietic SPTBN p.r681w c.2041c>t and lymphoid Lymphoid neoplasm SPTBN p.d161y c.481g>t Large intestine Carcinoma SPTBN p.l2084s c.6251t>c Large intestine Carcinoma SPTBN p.h1634r c.4901a>g Large intestine Carcinoma SPTBN p.r384w c.1150c>t Large intestine Carcinoma SPTBN p.a2240t c.6718g>a Large intestine Carcinoma SPTBN p.r913h c.2738g>a Large intestine Carcinoma SPTBN p.r1375q c.4124g>a Large intestine Carcinoma SPTBN p.w970r c.2908t>c Large intestine Carcinoma Previously Reported
27 SPTBN p.q1440h c.4320a>c Large intestine Carcinoma SPTBN p.r1739q c.5216g>a Large intestine Carcinoma SPTBN p.n1527i c.4580a>t Large intestine Carcinoma SPTBN p.l1948f c.5842c>t Large intestine Carcinoma SPTBN p.e622a c.1865a>c Large intestine Carcinoma SPTBN p.e1533* c.4597g>t Large intestine Carcinoma SPTBN p.r107* c.319c>t Large intestine Carcinoma SPTBN p.s2245a c.6733t>g Liver Carcinoma SPTBN p.l1355v c.4063c>g Liver Carcinoma p.k593fs*1 c.1777_1781delaagt SPTBN T Lung Carcinoma SPTBN p.i1636m c.4908c>g Ovary Carcinoma SPTBN p.l962f c.2884c>t Ovary Carcinoma SPTBN p.l1314* c.3941t>a Ovary Carcinoma SPTBN p.r144w c.430c>t Pancreas Carcinoma SPTBN p.m133t c.398t>c Prostate Carcinoma SPTBN p.m526i c.1578g>a Skin Carcinoma SPTBN p.r972q c.2915g>a Skin Carcinoma Malignant SPTBN p.r1915c c.5743c>t Skin melanoma Malignant SPTBN p.r1915c c.5743c>t Skin melanoma Previously Reported Previously Reported LIHC TCGA Gene Name SMAD3 SMAD3 SMAD3 SMAD3 SMAD3 Tumor_Sample_ Barcode AA Change Chrom Change Primary Tissue Variant Status TCGA-DD-A11C-01A- Missense 11D-A12Z-10 p.f376l c.t1126c Liver TCGA-DD-A1EL-01A- Missense 11D-A p.l42f c.c124t Liver TCGA-DD-A39Y-01A- 11D-A20W-10 p.t392fs c.1175delc Liver Frame_Shift_Del TCGA-G3-A6UC-01A- Missense 21D-A33K-10 p.g25v c.g74t Liver TCGA-RC-A6M4-01A- Missense 11D-A32G-10 p.k333e c.a997g Liver SPTBN1 SPTBN1 SPTBN1 SPTBN1 TCGA-BW-A5NP-01A- 11D-A27I-10 p.n32y c.a94t Liver TCGA-DD-A119-01A- 11D-A12Z-10 p.e33d c.g99t Liver TCGA-DD-A3A1-01A- 11D-A20W-10 p.l388r c.t1163g Liver TCGA-FV-A3R2-01A- 11D-A22F-10 p.e489k c.g1465a Liver Missense Missense Missense Missense SPTBN1 TCGA-G3-A25Y-01A- 11D-A16V-10 p.g562c c.g1684t Liver Missense
28 SPTBN1 SPTBN1 SPTBN1 SPTBN1 SPTBN1 SPTBN1 SPTBN1 TCGA-FV-A4ZQ-01A- 11D-A25V-10 p.n1106s c.a3317g Liver TCGA-DD-A1EH-01A- 11D-A12Z-10 p.r1141w c.c3421t Liver TCGA-QA-A7B7-01A- 11D-A32G-10 p.a1220v c.c3659t Liver Missense Missense Missense TCGA-BC-A112-01A- 11D-A12Z-10 NA c a>g Liver Splice_Site TCGA-UB-A7MB-01A- 11D-A33Q-10 p.a1645e c.c4934a Liver TCGA-DD-A4NG-01A- 11D-A27I-10 p.l1783r c.t5348g Liver TCGA-EP-A26S-01A- 11D-A16V-10 p.w653r c.t1957c Liver Missense Missense Missense MD Anderson Gene Name Tumor_Sample_ Barcode AA Change Chrom Change Primary Tissue Variant Classification Status SPTBN1 9194MDACC p.d1089y c.g3265t Liver Missense_
29 Supplemental Table 4. Mutual exclusivity of co-occurrence of SPTBN1 with SMAD2, SMAD3 or SMAD4 in TCGA dataset.* Co-occur Cancer type Cohort Gene A Gene B p-value Log Odds Ratio Association SPTBN1+SMAD2 Colorectal Adenocarcinoma Genentech SPTBN1 SMAD significant tendency towards co-occurrence TCGA SPTBN1 SMAD association towards co-occurrence Stomach Adenocarcinoma TCGA SPTBN1 SMAD association towards co-occurrence Uterine Corpus Endometrioid Carcinoma TCGA SPTBN1 SMAD association towards co-occurrence SPTBN1+SMAD3 Cervical Squamous Cell Carcinoma TCGA SPTBN1 SMAD >3 significant tendency towards co-occurrence Kidney Chromophobe TCGA SPTBN1 SMAD >3 significant tendency towards co-occurrence Colorectal Adenocarcinoma TCGA SPTBN1 SMAD association towards co-occurrence Head and Neck Squamous Cell Carcinoma TCGA SPTBN1 SMAD >3 association towards co-occurrence Nasopharyngeal Carcinoma Singapore SPTBN1 SMAD >3 significant tendency towards co-occurrence Uterine Corpus Endometrioid Carcinoma TCGA SPTBN1 SMAD association towards co-occurrence SPTBN1+SMAD4 Cervical Squamous Cell Carcinoma TCGA SPTBN1 SMAD <-3 association towards co-occurrence Colorectal Adenocarcinoma Genentech SPTBN1 SMAD association towards co-occurrence Stomach Adenocarcinoma Pfizer/UHK SPTBN1 SMAD association towards co-occurrence Uterine Corpus Endometrioid Carcinoma TCGA SPTBN1 SMAD association towards co-occurrence *: Analyzed by cbio portal complements existing tools ( : Derived from Fisher Exact Text. : p-value < 0.05: significant tendency towards co-occurrence. Log Odds Ratio > 0: association towards co-occurrence; Log Odds Ratio 0: association towards mutual exclusivity.
30 Supplemental Table 5. Top 5 predicted activation and inactivation diseases/functions in MEFs. Sptbn1 +/ /Smad3 +/ z-score Sptbn +/ z-score Sptbn1 / z-score Smad3 +/ z-score Activation: Cell movement Organism death Hypertrophy Organism death Leukocyte migration 3.46 Hypertrophy of heart Size of infarct Synthesis of polysaccharide Migration of cells Cardiac fibrosis Size of lesion Metabolism of carbohydrate Cell movement of leukocytes Abnormal morphology of body cavity Lesion formation Blood pressure Chemotaxis of cells Quantity of vitamin 1.98 Heart Disease Multiple congenital anomalies Inactivation: Organism death Infection of mammalia Activation of blood cells Activation of leukocytes Parasitic infection Activation of cells Activation of myeloid Inflammation of organ cells Adhesion of blood Nephritis cells Damage of connective tissue Size of body Accumulation of Metabolism of phagocytes terpenoid Damage of cartilage tissue Analgesia Accumulation of myeloid cells Antinociception Cell movement of macrophages Infiltration of cells
31 Supplemental Table 6. Top 5 predicted activation and inactivation diseases/functions in BWS cell lines. CDKN1C+ z-score KvDMR+ z-score KvDMR- z-score Activation: Organism death Fertility Size of body Growth failure Proliferation of embryonic cells Proliferation of embryonic cells Anemia Synthesis of DNA Lymphangiogenesis Heart disease Cell surface receptor linked signal transduction Cell surface receptor linked signal transduction Tumorigenesis of malignant tumor Size of body 1.93 Synthesis of DNA Inactivation: Size of body Organism death Organism death Cell movement Morphology of digestive system Perinatal death Viral infection Adhesion of blood cells Damage of lung Cell viability Perinatal death Adhesion of blood cells Cell survival Congenital anomaly of musculoskeletal system Hypoplasia
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Information
 Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Figure 1
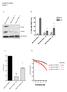 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the
 Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous
 Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous LRP5 in intact adult mouse ventricular myocytes (AMVMs)
Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous LRP5 in intact adult mouse ventricular myocytes (AMVMs)
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with
 Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung
 Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
Imprinting. Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821
 Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation
 Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Supplementary Figure S1 Supplementary Figure S2
 Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplementary Materials and Methods
 DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
(A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and a
 Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser
 Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Genetic Alteration Panels
 TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure 1
 Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth.
 Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt
 Supplementary Information CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt Jae Hyang Lim, Hirofumi Jono, Kensei Komatsu, Chang-Hoon Woo, Jiyun Lee, Masanori Miyata,
Supplementary Information CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt Jae Hyang Lim, Hirofumi Jono, Kensei Komatsu, Chang-Hoon Woo, Jiyun Lee, Masanori Miyata,
(A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939 (5 µm)
 Supplementary Figure Legends Figure S1. Tankyrase inhibition suppresses cell proliferation in an axin/β-catenin independent manner. (A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939
Supplementary Figure Legends Figure S1. Tankyrase inhibition suppresses cell proliferation in an axin/β-catenin independent manner. (A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Supplemental Information. Integrated Genomic Analysis of the Ubiquitin. Pathway across Cancer Types
 Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated
 Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated with the H3K27me3 mark in chondrocytes (see Table S1,
Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated with the H3K27me3 mark in chondrocytes (see Table S1,
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts
 Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III
 Supplementary Materials: Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III patient samples. Genomic DNA samples extracted from punch biopsies from either FFPE or frozen tumor
Supplementary Materials: Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III patient samples. Genomic DNA samples extracted from punch biopsies from either FFPE or frozen tumor
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Nature Genetics: doi: /ng Supplementary Figure 1. Alternative splicing events in the 5K panel.
 Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
SUPPLEMENTARY FIGURES AND TABLE
 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
The subcortical maternal complex controls symmetric division of mouse zygotes by
 The subcortical maternal complex controls symmetric division of mouse zygotes by regulating F-actin dynamics Xing-Jiang Yu 1,2, Zhaohong Yi 1, Zheng Gao 1,2, Dan-dan Qin 1,2, Yanhua Zhai 1, Xue Chen 1,
The subcortical maternal complex controls symmetric division of mouse zygotes by regulating F-actin dynamics Xing-Jiang Yu 1,2, Zhaohong Yi 1, Zheng Gao 1,2, Dan-dan Qin 1,2, Yanhua Zhai 1, Xue Chen 1,
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1. Deletion of Smad3 prevents B16F10 melanoma invasion and metastasis in a mouse s.c. tumor model.
 A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β
 Supplementary Figures Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β and LPS. Non-parenchymal liver cells were isolated and treated with or without TGF-β
Supplementary Figures Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β and LPS. Non-parenchymal liver cells were isolated and treated with or without TGF-β
AP VP DLP H&E. p-akt DLP
 A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
microrna-200b and microrna-200c promote colorectal cancer cell proliferation via
 Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells
 Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Increased ABHD5 expression in human colon cancer associated macrophages. (a) Murine peritoneal macrophages were treated with regular culture medium (Ctrl) or
Supplementary Figures Supplementary Figure 1 Increased ABHD5 expression in human colon cancer associated macrophages. (a) Murine peritoneal macrophages were treated with regular culture medium (Ctrl) or
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis
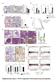 Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Belgian study into von Willebrand Disease (B-Will): First results
 Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
HIF-inducible mir-191 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment.
 HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
Nature Immunology doi: /ni.3268
 Supplementary Figure 1 Loss of Mst1 and Mst2 increases susceptibility to bacterial sepsis. (a) H&E staining of colon and kidney sections from wild type and Mst1 -/- Mst2 fl/fl Vav-Cre mice. Scale bar,
Supplementary Figure 1 Loss of Mst1 and Mst2 increases susceptibility to bacterial sepsis. (a) H&E staining of colon and kidney sections from wild type and Mst1 -/- Mst2 fl/fl Vav-Cre mice. Scale bar,
Supplementary Information
 Supplementary Information Supplementary Figure 1. Western blotting with ERβ antibodies Full blots corresponding to Fig. 2, along with replicated experiments at different time points, different batches,
Supplementary Information Supplementary Figure 1. Western blotting with ERβ antibodies Full blots corresponding to Fig. 2, along with replicated experiments at different time points, different batches,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
