Supplementary Figure 1. Chromatin signature profiling by mass spectrometry.
|
|
|
- Stanley George
- 5 years ago
- Views:
Transcription
1 Supplementary Information Supplementary Figure 1. Chromatin signature profiling by mass spectrometry. 115 cell lines from the CCLE collection were subjected to chromatin signature profiling by mass spectrometry. Each row corresponds to a histone H3 peptide with the specific combination of marks shown on the right. The value in each cell of the heatmap corresponds to the log2-fold change of the mark combination vs. the median for each row. Tissue origin, hematological subtypes and mutation status of EZH2 and NSD2 are indicated. Clusters A-F are marked for reference in the main text.
2
3 Supplementary Figure 2. Cell lines containing NSD2 E1099K mutation express both wild type and E1099K mutant NSD2 alleles. cdna from six NSD2 E1099K mutant expressing cell lines and three NSD2 wildtype cell lines was subjected to Sanger Sequencing and sequencing chromatograms are shown; left and right columns are chromatograms using forward and reverse sequencing primers respectively. cdna sequencing of all mutant cell lines showed that both the wild type and mutant NSD2 transcripts are expressed. Mutation caused nucleotide change of Gà A (red) at codon 1099 (underlined and marked by red arrow in the chromatograms) resulting amino acid change of Eà K.
4
5
6 Supplementary Figure 3. Structural model of NSD2 E1099K mutation Structural model of NSD2 sequence mutations, E1099 (magenta stick with black text) to K1099 (white stick with red text) and D1125 (magenta stick with black text) to N1125 (white stick with red text). They are predicted to alter binding between NSD2 (green cartoon) and the histone peptide (blue cartoon). Amino acids K1124 and C1183 are labeled and shown as green sticks. Residues that line the I-SET histone binding groove are labeled with grey text and shown as green sticks (C1102, I1106, M1119, I1127, and T1121). The residues S31 (blue stick) and K36 (blue stick) are shown to identify modeled locations of residues on the H3 peptide. Black dash highlights key interactions.
7 Supplementary Figure 4. Molecular chromatin signatures of KMS11-TKO engineered lines.
8 Supplementary Figure 5. NSD2 variants identified in primary ALL samples promote H3K36 dimethylation and transformation. A) Western blot of KMS11-TKO cells reconstituted with NSD2-wt, NSD2-CDM (catalytic dead mutation), NSD2-E1099K, NSD2-E1099K-CDM (harboring both E1099K and the catalytic inactive mutation), NSD2-D1125N, NSD2-MB4-2 (N-terminal truncation), or the empty vector. NSD2 MB4-2 is a structural variant identified in pediatric tumor sample (see Supplementary information, NSD2 structural variations in pediatric cancer. Cell lysates were blotted with the indicated antibodies. Histone H3 serves as loading control. B) Quantification of soft agar transformation (in presence of 5% serum) of KMS11-TKO cells reconstituted with the indicated NSD2 variants. Error bars indicate the s.d. of triplicate samples. C) Quantification of soft agar transformation of KMS11-TKO cells reconstituted with the indicated NSD2 variants in the presence of 10% serum. Two independent experiments were shown. Error bars indicate the s.d. of triplicate samples. This set of experiments was performed to evaluate if different serum concentration (5% vs 10%) might affect the outcome of soft agar transformation. The actual number of colonies differs under the different serum concentration. Overall, these data showed transforming activity of (NSD2 E1099K ~ NSD2 D1125N ~ MB4-2) > NSD2 wt > (vector control ~ NSD2 CDM ~ NSD2 E1099KCDM).
9
10 Supplementary Figure 6. NSD2 E1099K specifically alters H3K36 methylation and not generally affects other histone H3 modifications. a) Immunoblot of blot of H3K4me1/2/3 and H3K9me1/2/3 of KMS11-TKO cells transduced with the lentivirus expressing the indicated NSD2 variants. b) Mass spectrometry profiling of the indicated histone peptides in SEM cells following NSD2 knockdown. C) Immunoblot of H3K27me3 and H3 of the tumor samples as shown in Fig 4f.
11 Supplementary Figure 7. NSD2 E1099K promotes H3K36me2 in NIH3T3. Western blot of NSD2, H3K36me3, total H3 and γ-tubulin in NIH3T3 cells transfected with the indicated NSD2 variants. Results from two independent experiments were shown.
12 Supplementary Figure 8. An example of NSD2 mutant isoform detected by RNA-seq in the three ETV6-RUNX1 ALL cases with intra-genic 5 deletion. A) Junction reads from RNA-seq across the alternative exon 1 and exon 6 confirm a novel isoform resulting from intra-genic genomic deletion in sample ETV086. The junction reads spans the alternative exon 1 (encoded only by isoform 10 of NSD2, accession NM_ ) and exon 6 of NSD2. The alternative ATG is shown in black box. B) Mutant protein with truncated N-termini predicted from the mutant transcript.
13 Supplementary Figure 9. An example of inter-chromosomal re-arrangement resulting in fusion of the 5 UTR of ST6GAL1 with exon 1 of NSD2 Junction reads in RNA-seq that span 5 UTR of ST6GAL1 (left box, in blue) and the first coding exon of NSD2 (right box, in black) resulting from inter-chromosomal translocation in sample LGG039. The junction reads mapped to ST6GAL1 were marked as the soft-clipped reads by bwa (shown in light blue color in Bambino viewer at the left).
14 Supplementary Table 1. Cell lines screened by global histone profiling Cell Line Name Cluster Tissue of Origin Haematological Subtype t(4;14) translocation NSD2 variants EZH2 variants KARPAS422 A Heme/Lymph BCL Y641N Pfeiffer A Heme/Lymph BCL A677G SUDHL6 A Heme/Lymph BCL Y641N SUDHL4 A Heme/Lymph BCL Y641S RL A Heme/Lymph BCL Y641N WSUDLCL2 A Heme/Lymph BCL Y641F DB A Heme/Lymph BCL Y641N SUDHL10 A Heme/Lymph BCL Y641F 253JBV B Urinary A956P 5637 B Urinary SW1710 B Urinary KMBC2 B Urinary HEC108 B Endometrium K361Q RT4 B Urinary MOLM16 B Heme/Lymph AML L1236 B Heme/Lymph HL SCaBER B Urinary HT1376 B Urinary KHM1B B Heme/Lymph MM J82 B Urinary NCIH650 B Lung CAL29 B Urinary DEL B Heme/Lymph NS KMS20 B Heme/Lymph MM EB2 B Heme/Lymph BL KMS27 B Heme/Lymph MM UMUC3 B Urinary NCIH3255 B Lung BV173 B Heme/Lymph CML NB4 B Heme/Lymph AML T236A UBLC1 B Urinary QGP1 B Pancreas CMK115 B Heme/Lymph AML G945fs OCILY19 B Heme/Lymph BCL CMK B Heme/Lymph AML KMS12BM B Heme/Lymph MM MOLM6 B Heme/Lymph CML K361Q
15 MUTZ5 B Heme/Lymph ALL ML1 B Thyroid HDLM2 B Heme/Lymph HL SUPT11 B Heme/Lymph TCL PC3 B Prostate MonoMac1 B Heme/Lymph AML MonoMac6 B Heme/Lymph AML Nomo1 B Heme/Lymph AML Toledo B Heme/Lymph BCL GA10 B Breast KOPN8 B Heme/Lymph ALL KP4 B Pancreas Y696C TF1 B Heme/Lymph AML MDAMB157 B Breast OCIAML2 B Heme/Lymph AML NCIH1568 B Lung U937 B Heme/Lymph BCL UMUC1 C Urinary MOLM13 C Heme/Lymph AML JHUEM7 C Endometrium D672Y EJM C Heme/Lymph MM SUDHL5 C Heme/Lymph BCL KLE C Endometrium OCILY3 C Heme/Lymph BCL 647V C Urinary 639V C Urinary L428 C Heme/Lymph HL Y641S KMS26 D Heme/Lymph MM yes MM1S D Heme/Lymph MM E1099K KMS28BM D Heme/Lymph MM yes KMS34 D Heme/Lymph MM yes LP1 D Heme/Lymph MM yes OPM2 D Heme/Lymph MM yes KMS11 D Heme/Lymph MM yes RPMI8402 D Heme/Lymph ALL E1099K RS411 D Heme/Lymph ALL E1099K SEM D Heme/Lymph ALL E1099K MOLT13 D Heme/Lymph ALL E1099K RCHACV D Heme/Lymph ALL E1099K HPBALL D Heme/Lymph ALL E1099K
16 KMRC3 E Kidney G945fs IGR1 E Skin Y641N OVISE E Ovarian Set2 E Heme/Lymph ET MDAMB361 E Breast ZR7530 E Breast S619C LNCAP E Prostate THP1 E Heme/Lymph AML VMCUB1 E Urinary S804L TOV21G E Ovarian NAMALWA E Heme/Lymph BL KU1919 E Urinary HEL9217 E Heme/Lymph AML SKMEL30 E skin W624* BFTC905 E Urinary RT112 E Urinary L363 E Heme/Lymph MM T24 E Urinary NCIH1930 E Lung NCIH209 E Lung IGROV1 E Ovarian L33 E Pancreas MV411 E Heme/Lymph AML JJN3 E Heme/Lymph MM N268D SUIT2 E Pancreas SW780 E Urinary RERFLCAd2 E Lung TT E Oesophagus K361Q PL21 E Heme/Lymph AML ESS1 E Endometrium R685C JHOM1 E Ovarian KO52 E Heme/Lymph AML TALL1 F Heme/Lymph ALL I708M SW579 F Thyroid E1099K JMSU1 F Urinary G623D SKM1 F Heme/Lymph AML Y641C KE37 F Heme/Lymph ALL H279Y OCIAML5 F Heme/Lymph AML R685H Note: cell lines appear in the same order (left to right) as in Fig. 1 and Supplementary Fig. 1.
17 Supplementary Table 2. Summary of EZH2 status of 5 cell lines in cluster F by RNA-seq. Cell Line cdna_change Protein_Change t_alt_count t_ref_count JMSU1 c.1868g>a p.g623d KE37 c.835c>t p.h279y OCIAML5 c.2054g>a p.r685h SKM1 c.1922a>g p.y641c TALL1 c.1665c>a p.n555k TALL1 c.2124a>g p.i708m Set domain sequence alignment of five human methyltransferases performed by CLUSTALW. (*. : indicated identical residues, and residues showing different degrees of homology). Highlighted in yellow (G623), grey (Y641), green (R685) and red (I708) in the SET domain of EZH2. NSD2_ SETD2_ EZH2_ SUV39H1_ SETD7_ NSD2_ SETD2_ EZH2_ SUV39H1_ SETD7_ NSD2_ SETD2_ EZH2_ SUV39H1_ SETD7_ PETKIIKTD-GKGWGLVAKRDIRKGEFVNEYVGELIDEEECMARIKHAHE ADVEVILTE-KKGWGLRAAKDLPSNTFVLEYCGEVLDHKEFKARVKEYAR KHLLLAPSD-VAGWGIFIKDPVQKNEFISEYCGEIISQDEADRRGKVYD- YDLCIFRTDDGRGWGVRTLEKIRKNSFVMEYVGEIITSEEAERRGQIYDR GEGLFSKVAVGPNTVMSFYNGVRITHQEVDSRDWALNG * *: :..: * * :.* * NDITHFYMLTIDKDR-IIDAGPKGN-YSRFMNHSCQPNCETLKWTVN--- NKNIHYYFMALKNDE-IIDATQKGN-CSRFMNHSCEPNCETQKWTVN--- -KYMCSFLFNLNNDF-VVDATRKGN-KIRFANHSVNPNCYAKVMMVN--- QGATYLFDLDYVEDVYTVDAAYYGN-ISHFVNHSCDPNLQVYNVFIDNLD NTLSLDEETVIDVPEPYNHVSKYCASLGHKANHSFTPNCIYDMFVHPR--.. : *** ** -GDTRVGLFAVCDIPAGTELTFNYNLDCLGN -GQLRVGFFTTKLVPSGSELTFDYQFQRYGK -GDHRIGIFAKRAIQTGEELFFDYRYSQADA ERLPRIAFFATRTIRAGEELTFDYN FGPIKCIRTLRAVEADEELTVAYG : : : :. **. *
18 Supplementary Table 3. Summary of CCLE cell lines with NSD2 E1099K mutation. Cell lines HIST_SUBTYPE NSD2 protein change GENDER AGE_YRS sem acute_lymphoblastic_b_cell_leukaemia p.e1099k female 5 rs411 acute_lymphoblastic_b_cell_leukaemia p.e1099k female 32 rchacv acute_lymphoblastic_leukaemia p.e1099k female 8 hpball acute_lymphoblastic_t_cell_leukaemia p.e1099k male 14 rpmi8402 acute_lymphoblastic_t_cell_leukaemia p.e1099k female 16 molt13 acute_lymphoblastic_t_cell_leukaemia p.e1099k female 2 sw579 anaplastic_carcinoma p.e1099k male 59 mm1s plasma_cell_myeloma p.e1099k
19 Supplementary Table 4. Summary of colony growth in soft agar of 24 ALL cell lines.
20 Supplementary Table 5. Mutant Allele Fraction (MAF) of NSD2 somatic mutations in DNA and mrna samples of PCGP analyzed by WGS, custom capture and RNA-seq. MAF 0.1 was highlighted. The exon number of the four deletions were based on isoform 1 of NSD2 (accession NM_133330). * NGS read count in DNA combines data from WGS and custom capture when available ^MAF stands for mutant allele fraction, i.e. number of NGS reads with the mutant allele/number of all NGS at the site. Mutations with MAF <0.1 are matching normal sample of this tumor has approximately 20% contaminating tumor material.
21
22 Supplementary Table 6. Examples of synthetic peptides for many possible peptide/modification combinations on histone H3. Mass [m/z] Start time [min] End time [min] Normalized HCD Collision Energy (%) Charge State [z] Targeted Species* H3K4me2-L H3K4me2-H H3K27me2K36me2-L H3K4ac1-L H3K4me3-L H3K27me2K36me2-H H3K27me3K36me2-L+H3K27me2K36me3-L H3K4ac1-H H3K4me3-H H3K27me3K36me2-H+H3K27me2K36me3-H H3K27me3K36me3-L H3K4me0-L H3K27me3K36me3-H H3K4me0-H H3K4me1-L H3K4me1-H H3K9me2K14ac1-L H3K9me2K14ac1-H H3K9ac1K14ac1-L H3K9me3K14ac1-L H3K9me2K14ac0-L H3K9ac1K14ac1-H H3K9me3K14ac1-H H3K9me2K14ac0-H H3K9ac1K14ac0-L+H3K9me0K14ac1-L H3K9me3K14ac0-L H3K9ac1K14ac0-H+H3K9me0K14ac1-H H3K9me3K14ac0-H H3K9me0K14ac0-L H3K9me1K14ac1-L H3K27ac1K36me2-L H3K9me0K14ac0-H H3K9me1K14ac1-H
23 H3K9me1K14ac0-L H3K27ac1K36me2-H H3K27ac1K36me3-L H3K27me2K36me0-L H3K27me0K36me2-L H3(41-49)-L H3K9me1K14ac0-H H3K27me2K36me0-H H3K27me0K36me2-H H3K27ac1K36me3-H H3K27ac1K36me0-L H3K27me1K36me2-L H3K27me0K36me3-L H3K27me2K36me1-L H3K27me3K36me0-L H3K27ac1K36me0-H H3K27me1K36me2-H H3K27me3K36me0-H H3K27me2K36me1-H H3K27me0K36me3-H H3K27me0K36me0-L+H3K27ac1K36me1-L H3K27me1K36me3-L H3K27me3K36me1-L H3K9me2S10ph1K14ac1-L H3(41-49)-H H3K27ac1K36me1-H+H3K27me0K36me0-H H3K27me1K36me3-H H3K27me3K36me1-H H3K27me1K36me0-L H3K27me0K36me1-L H3.3K27me0K36me0-L H3K9me2S10ph1K14ac1-H H3K9ac1S10ph1K14ac1-L H3K9me3S10ph1K14ac1-L H3K9me2S10ph1K14ac0-L H3K27me1K36me0-H H3K27me0K36me1-H H3.3K27me0K36me0-H H3K27me1K36me1-L H3K18ac1K23ac1-L
24 H3K27me1K36me1-H H3K9ac1S10ph1K14ac1-H H3K9me3S10ph1K14ac1-H H3K9me2S10ph1K14ac0-H H3K9ac1S10ph1K14ac0-L+H3K9me0S10ph1K14ac1- L H3K9me3S10ph1K14ac0-L H3K18ac1K23ac1-H H3K18ac1K23ac0-L+H3K18ac0K23ac1-L H3K9ac1S10ph1K14ac0-H+H3K9me0S10ph1K14ac1- H H3K9me3S10ph1K14ac0-H H3K9me0S10ph1K14ac0-L H3K9me1S10ph1K14ac1-L H3K18ac1K23ac0-H+H3K18ac0K23ac1-H H3K18ac0K23ac0-L H3K9me1S10ph1K14ac1-H H3K9me0S10ph1K14ac0-H H3K9me1S10ph1K14ac0-L H3K18ac0K23ac0-H H3K9me1S10ph1K14ac0-H H3K18ub1K23ac0-L+H3K18ac0K23ub1-L H3K18ub1K23ac0-H+H3K18ac0K23ub1-H H3K56me2-L H3K56me2-H H3K56me0-L H3K56me0-H H3K56me1-L H3K56me1-H H3K79me2-L H3K79me2-H H3K79me0-L H3K79me0-H H3K79me1-L H3K79me1-H *The label of the species targeted refers to the resultant derivatized peptide containing the stated modification after preparation as described in the Methods section. In some cases one set of (m/z, time window) coordinates may cover multiple species. -L denotes light peptides derived from the CCLE cell line while -H denotes heavy peptides derived from the R 10 standard mix cell lines.
25 Supplementary Table 7. Peptides monitored by Global Chromatin Profiling with histone nomenclature and biochemical abbreviations Nomenclature Base Sequence Modified Sequence H3K4me0 TKQTAR (pr)-t(kpr)qtar H3K4me1 TKQTAR (pr)-t(kme1pr)qtar H3K4me2 TKQTAR (pr)-t(kme2)qtar H3K4ac1 TKQTAR (pr)-t(kac)qtar H3K9me0K14ac0 KSTGGKAPR (pr)-(kpr)stgg(kpr)apr H3K9me0K14ac1 KSTGGKAPR (pr)-(kpr)stgg(kac)apr H3K9me1K14ac0 KSTGGKAPR (pr)-(kme1pr)stgg(kpr)apr H3K9me1K14ac1 KSTGGKAPR (pr)-(kme1pr)stgg(kac)apr H3K9me2K14ac0 KSTGGKAPR (pr)-(kme2)stgg(kpr)apr H3K9me2K14ac1 KSTGGKAPR (pr)-(kme2)stgg(kac)apr H3K9me3K14ac0 KSTGGKAPR (pr)-(kme3)stgg(kpr)apr H3K9me3K14ac1 KSTGGKAPR (pr)-(kme3)stgg(kac)apr H3K9ac1K14ac0 KSTGGKAPR (pr)-(kac)stgg(kpr)apr H3K9ac1K14ac1 KSTGGKAPR (pr)-(kac)stgg(kac)apr H3K18ac0K23ac0 KQLATKAAR (pr)-(kpr)qlat(kpr)aar H3K18ac1K23ac0 KQLATKAAR (pr)-(kac)qlat(kpr)aar H3K18ac0K23ac1 KQLATKAAR (pr)-(kpr)qlat(kac)aar H3K18ac1K23ac1 KQLATKAAR (pr)-(kac)qlat(kac)aar H3K18ac0K23ubq1 KQLATKAAR (pr)-(kpr)qlat(kggpr)aar H3K27me0K36me0 KSAPATGGVKKPHR (pr)-(kpr)sapatggv(kpr)(kpr)phr H3K27me0K36me1 KSAPATGGVKKPHR (pr)-(kpr)sapatggv(kme1pr)(kpr)phr H3K27me0K36me2 KSAPATGGVKKPHR (pr)-(kpr)sapatggv(kme2)(kpr)phr H3K27me0K36me3 KSAPATGGVKKPHR (pr)-(kpr)sapatggv(kme3)(kpr)phr H3K27me1K36me0 KSAPATGGVKKPHR (pr)-(kme1pr)sapatggv(kpr)(kpr)phr H3K27me1K36me1 KSAPATGGVKKPHR (pr)- (Kme1pr)SAPATGGV(Kme1pr)(Kpr)PHR H3K27me1K36me2 KSAPATGGVKKPHR (pr)-(kme1pr)sapatggv(kme2)(kpr)phr H3K27me1K36me3 KSAPATGGVKKPHR (pr)-(kme1pr)sapatggv(kme3)(kpr)phr H3K27me2K36me0 KSAPATGGVKKPHR (pr)-(kme2)sapatggv(kpr)(kpr)phr H3K27me2K36me1 KSAPATGGVKKPHR (pr)-(kme2)sapatggv(kme1pr)(kpr)phr H3K27me2K36me2 KSAPATGGVKKPHR (pr)-(kme2)sapatggv(kme2)(kpr)phr H3K27me3K36me0 KSAPATGGVKKPHR (pr)-(kme3)sapatggv(kpr)(kpr)phr H3K27me3K36me1 KSAPATGGVKKPHR (pr)-(kme3)sapatggv(kme1pr)(kpr)phr H3K27ac1K36me0 KSAPATGGVKKPHR (pr)-(kac)sapatggv(kpr)(kpr)phr H3K27ac1K36me1 KSAPATGGVKKPHR (pr)-(kac)sapatggv(kme1pr)(kpr)phr H3K27ac1K36me2 KSAPATGGVKKPHR (pr)-(kac)sapatggv(kme2)(kpr)phr H3K27ac1K36me3 KSAPATGGVKKPHR (pr)-(kac)sapatggv(kme3)(kpr)phr H3.3K27me0K36me0 KSAPSTGGVKKPHR (pr)-(kpr)sapstggv(kpr)(kpr)phr H3Y41ph0 YRPGTVALR (pr)-yrpgtvalr
26 H3K56me0 YQKSTELLIR (pr)-yq(kpr)stellir H3K56me1 YQKSTELLIR (pr)-yq(kme1pr)stellir H3K79me0 EIAQDFKTDLR (pr)-eiaqdf(kpr)tdlr H3K79me1 EIAQDFKTDLR (pr)-eiaqdf(kme1pr)tdlr H3K79me2 EIAQDFKTDLR (pr)-eiaqdf(kme2)tdlr Abbreviations: Chemical derivatives: Pr Ac me1 me2 me3 proprionyl acetyl monomethyl dimethyl trimethyl Nomenclature not defined above: Ubq me0 = ac0 ubiquityl stub (GG) unmodified
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Information
 Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Figure 1: Normal vs. Tumor expression of PIM kinases mrna across 17 tissue types
 Supplementary Figure 1: Normal vs. Tumor expression of PIM kinases mrna across 17 tissue types Supplementary Figure 1 Legend: PIM1, PIM2 and PIM3 mrna expression Expression levels in cancer tissues derived
Supplementary Figure 1: Normal vs. Tumor expression of PIM kinases mrna across 17 tissue types Supplementary Figure 1 Legend: PIM1, PIM2 and PIM3 mrna expression Expression levels in cancer tissues derived
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells
 Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
Supplementary Information
 Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion
 Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Control shrna#9 shrna#12. shrna#12 CD14-PE CD14-PE
 a Control shrna#9 shrna#12 c Control shrna#9 shrna#12 e Control shrna#9 shrna#12 h 14 12 CFU-E BFU-E GEMM GM b Colony number 7 6 5 4 3 2 1 6 pm A pa pc CFU-E BFU-E GEMM GM pu pgm A p pg B d f CD11b-APC
a Control shrna#9 shrna#12 c Control shrna#9 shrna#12 e Control shrna#9 shrna#12 h 14 12 CFU-E BFU-E GEMM GM b Colony number 7 6 5 4 3 2 1 6 pm A pa pc CFU-E BFU-E GEMM GM pu pgm A p pg B d f CD11b-APC
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Supplementary Figure S1. Generation of LSL-EZH2 conditional transgenic mice.
 Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplemental Figure S1A Notch1
 Supplemental Figure S1A Notch1 erage) epth of Cove ormalized De Log1(No Notch exons Figure S1: A) Relative coverage of Notch1 and Notch 2 exons in HCC2218, HCC1187, MB157, MDA-MB157 cell lines. Blue color
Supplemental Figure S1A Notch1 erage) epth of Cove ormalized De Log1(No Notch exons Figure S1: A) Relative coverage of Notch1 and Notch 2 exons in HCC2218, HCC1187, MB157, MDA-MB157 cell lines. Blue color
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Information
 Supplementary Information Targeted Disruption of the EZH2/EED Complex Inhibits EZH2- dependent Cancer Woojin Kim 1,2,3, Gregory H. Bird 2,3,4, Tobias Neff 5, Guoji Guo 1,2,3, Marc A. Kerenyi 1,2,3, Loren
Supplementary Information Targeted Disruption of the EZH2/EED Complex Inhibits EZH2- dependent Cancer Woojin Kim 1,2,3, Gregory H. Bird 2,3,4, Tobias Neff 5, Guoji Guo 1,2,3, Marc A. Kerenyi 1,2,3, Loren
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
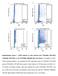 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Predictive PP1Ca binding region in BIG3 : 1,228 1,232aa (-KAVSF-) HEK293T cells *** *** *** KPL-3C cells - E E2 treatment time (h)
 Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Supplementary Figure 1
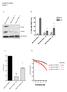 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/393/rs9/dc1 Supplementary Materials for Identification of potential drug targets for tuberous sclerosis complex by synthetic screens combining CRISPR-based knockouts
www.sciencesignaling.org/cgi/content/full/8/393/rs9/dc1 Supplementary Materials for Identification of potential drug targets for tuberous sclerosis complex by synthetic screens combining CRISPR-based knockouts
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing.
 Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical
 Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
ISOGENIC CELL LINES. ATCC No. Designation Mutation Parental Cell Line Disease Cancer Model
 THE ESSENTILS OF LIFE SCIENCE RESERCH GLOLLY DELIVERED ISOGENIC CELL LINES TCC CRISPR/CS9 GENE-EDITED ISOGENIC CELL LINES Clinically relevant cell models are critical for studies of molecular and cellular
THE ESSENTILS OF LIFE SCIENCE RESERCH GLOLLY DELIVERED ISOGENIC CELL LINES TCC CRISPR/CS9 GENE-EDITED ISOGENIC CELL LINES Clinically relevant cell models are critical for studies of molecular and cellular
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Supplementary Figure 1
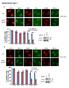 Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1
 Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
A. List of selected proteins with high SILAC (H/L) ratios identified in mass
 Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines.
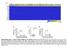 Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines. The scores summarize the global expression of the tissue
Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines. The scores summarize the global expression of the tissue
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
AP VP DLP H&E. p-akt DLP
 A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Generation of cetuximab-resistant cells and analysis of singlecell clones. Cetuximab-sensitive cells (LIM1215 and OXCO-2) were chronically treated with cetuximab
Supplementary Figure 1 Supplementary Figure 1: Generation of cetuximab-resistant cells and analysis of singlecell clones. Cetuximab-sensitive cells (LIM1215 and OXCO-2) were chronically treated with cetuximab
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Normal Skin. Tissue Samples and Melanoma Cell Lines. BRAF Mut. RAS Mut RAS WT /BRAF WT
 Supplemental Figure 1. MERTK gene expression in melanoma cell line panel from Cancer Cell Line Encyclopedia. A. Microarray analysis of melanoma cell lines from UNC collection grouped by oncogenic mutation.
Supplemental Figure 1. MERTK gene expression in melanoma cell line panel from Cancer Cell Line Encyclopedia. A. Microarray analysis of melanoma cell lines from UNC collection grouped by oncogenic mutation.
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
gliomas. Fetal brain expected who each low-
 Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
