Supplemental Data. Candat et al. Plant Cell (2014) /tpc Cytosol. Nucleus. Mitochondria. Plastid. Peroxisome. Endomembrane system
|
|
|
- Avice Morrison
- 6 years ago
- Views:
Transcription
1 Cytosol Nucleus PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT Mitochondria Plastid Peroxisome Endomembrane system PSORT MultiLoc YLoc SubLoc TargetP Predotar PProwler MITOPRED BaCelLo WoLF PSORT PSORT MultiLoc YLoc TargetP Predotar PProwler BaCelLo WoLF PSORT PSORT MultiLoc YLoc PProwler BaCelLo WoLF PSORT PSORT MultiLoc YLoc SubLoc TargetP Predotar PProwler BaCelLo WoLF PSORT Probability in % < Supplemental Figure 1. Heatmap of subcellular localization predictions of the 51 Arabidopsis LEA proteins. The heatmap illustrates probability over 50 % of LEA protein targeting to various cellular compartments given by the different software indicated on the right. LEA proteins are numbered from 1 to 51, and the color coding represents the targeting probability below 50 % (yellow) or between 50 to 100% (red). The two qualitative predictors BaCelLo and WoLF PSORT were included with black square indicating a positive prediction.
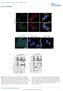
2 LEA-GFP NLS-RFP Merge GFP-LEA Supplemental Figure 2. Transient expression of cytosolic LEA protein in Arabidopsis protoplasts. Representative examples of LEA-GFP and GFP-LEA fusion proteins localized either in the cytosol only or both in the cytosol and the nucleus were co-expressed with a RFP nuclear marker in Arabidopsis mesophyll protoplasts. Numbers refer to the corresponding LEA proteins. Green, GFP; Red, RFP; Blue, chlorophyll. Bar = 10 µm.
3 LEA-GFP NLS-RFP Merge GFP-LEA Supplemental Figure 2. (continued).
4 LEA-GFP NLS-RFP Merge GFP-LEA Supplemental Figure 2. (continued).
5 LEA-GFP NLS-RFP Merge GFP-LEA Supplemental Figure 2. (continued).
6 LEA-GFP NLS-RFP Merge GFP-LEA Supplemental Figure 2. (continued).
7 LEA-GFP NLS-RFP Merge GFP-LEA 51 Supplemental Figure 2. (continued).
8 Supplemental Figure 3. Diffusion of cytosolic LEA-GFP proteins in the nucleus. The graph shows the distribution of LEA protein fusions between cytosolic only (C) or cytosolic-nuclear (C+N) compartments as a function of their molecular mass. Each box encloses 50% of the data with the median value of the variable displayed as a line. The mean is indicated as a dashed line. The top and bottom of the box mark the limits of ± 25% of the variable population. The lines extending from the top and bottom of each box mark the minimum and maximum values within the data set that fall within an acceptable range.
9 GFP Supplemental Figure 4. Western blot analysis of LEA protein expressed in protoplasts. Total proteins extracted from protoplasts expressing LEA-GFP fusions were separated by SDS-PAGE and transferred on a nylon membrane. LEA-GFP were immuno-detected with a specific anti-gfp antibody. Each LEA protein is indicated by its number. The molecular mass in the scale is in kda. Lanes between black bars (2, 4, 5, 21, 26, 30, 36, 38, 41, 47 and 49) were obtained from a second western blot analysis and were merged in the figure for more clarity. Arrows indicate the bands of interest.
10 Supplemental Figure 5. Cytosolic localization of GFP-LEA fusions for proteins localized in mitochondria or plastid when expressed as LEA-GFP fusions (see Figures 3 and 4). Representative images of GFP-LEA fusion proteins expressed in wild type Arabidopsis mesophyll protoplasts. Numbers refer to different LEA proteins for which LEA-GFP fusions were localized in plastid or mitochondria. Green, GFP; Blue, chlorophyll. Bar = 10 µm.
11 LEA-GFP mt-rfp Merge Supplemental Figure 6. Leaf cells from Arabidopsis transgenic lines expressing LEA proteins targeted to mitochondria or dual targeted to mitochondria and plastid. Representative images of LEA-GFP fusion proteins expressed in transgenic lines expressing a RFP mitochondrial marker. In the merged images fro LEA42 and 48, chlorophyll autofluorescence is also shown (in blue). Numbers refer to different LEA proteins. Green, GFP; Red, RFP; Blue, chlorophyll. Bar = 10 µm.
12 LEA-RFP Golgi-GFP Merge 13 Supplemental Figure 7. LEA13-RFP does not co-localize with Golgi in Arabidopsis protoplasts. LEA13-RFP fusion protein was expressed in Arabidopsis mesophyll protoplasts carrying a Golgi marker (GFP). Bar = 10 µm.
13 LEA-RFP ER-YFP Merge LEA-RFP Perox-YFP Merge 9 Supplemental Figure 8. Arabidopsis leaf cells from transgenic lines expressing LEA proteins targeted to the secretory pathway or the pexophagosome. Representative images of LEA-RFP fusion proteins expressed in transgenic lines expressing a YFP ER marker (ER-YFP) or a YFP peroxisome marker (Perox-YFP). Numbers refer to different LEA proteins. Green, YFP; Red, RFP. Bar = 10 µm.
14 Supplemental Figure 9. Transient expression in Arabidopsis protoplasts of GFP-LEA fusions for proteins which are targeted to the secretory pathway as LEA-GFP fusions. Representative images of GFP-LEA fusion proteins expressed in wild type Arabidopsis mesophyll protoplasts. Numbers refer to different LEA proteins for which LEA-GFP fusions were targeted to the secretory pathway. Green, GFP; Blue, chlorophyll. Bar = 10 µm.
15 LEA-RFP Golgi-GFP Merge 21 Supplemental Figure 10. Vacuolar and Golgi localization of LEA 21 in Arabidopsis protoplasts. LEA21-RFP fusion protein was expressed in Arabidopsis mesophyll protoplasts carrying a Golgi marker (GFP). Bar = 10 µm.
16 LEA21-GFP LEA22-GFP Supplemental Figure 11. Localization of LEA 21-GFP and LEA22-GFP in Arabidopsis protoplasts. GFP fusion protein was expressed in wild type Arabidopsis mesophyll protoplasts. In the case of LEA22-GFP, the fluorescence signal was very weak. Bar = 10 µm.
17 Supplemental Figure 12. LEA11 and LEA12 sequence alignment. Sequences were aligned using the MultAlin software ( using default parameters. The targeting peptide cleavage site of LEA11 is indicated with a black arrow.
18 ER (6) PProwler False positive 0 6 MultiLoc 0 5 Yloc 0 5 TargetP 0 5 Predotar 0 5 PSORT 1 3 SubLoc 2 2 Mitochondria (5) PProwler False positive 2 4 Yloc 2 4 TargetP 3 PSORT 1 2 MITOPRED 1 2 Predotar 1 1 MultiLoc 0 SubLoc 6 Plastid (6) PProwler False positive 0 5 TargetP 2 5 Predotar 0 3 Yloc 1 3 PSORT 2 2 MultiLoc 2 Correct prediction Correct prediction Correct prediction Cytosol (36) False positive Correct prediction Yloc 1 PSORT 2 14 SubLoc 3 7 MultiLoc Supplemental Figure 13. Estimation of prediction software efficiency. The bar graph shows the number of proteins with correct or false predictions by the different software. For each compartment, the total number of experimentally identified proteins is indicated between brackets. Correct predictions were achieved when the experimental subcellular localization matched with the prediction. False positives correspond to erroneous subcellular compartment attribution with a strong prediction.
19 LEA-RFP ER-GFP Merge 7 29 Supplemental Figure 14. Cytosolic localization of LEA7 and LEA29 in Arabidopsis protoplasts. LEA-RFP fusion proteins were expressed in Arabidopsis mesophyll protoplasts carrying a ER marker (GFP). Bar = 10 µm.
20 LEA# Loc ID Cell culture Flowers Leaves Pollen Roots Seedling shoots Seeds Siliques 1 C At1g C At1g C At1g C At1g C At1g C At1g C At1g C At1g Pxp At1g C At1g P At2g P At2g ER At2g C At2g C At2g C At2g C At2g C At2g C At2g C At2g V At2g V At2g P At2g P At2g C At2g C At2g C At2g C At3g C At3g ER At3g C At3g C At3g C At3g C At3g C At3g C At3g M At3g M At4g C At4g C At4g M At4g MP At4g ER At4g C At4g C At4g C At5g C At5g MP At5g C At5g C At5g C At5g Supplemental Figure 15. Proteomic quantification of LEA proteins in different organs of Arabidopsis. Proteomic data were obtained from the pep2pro database ( and the subcellular localizations were added from our experimental observations: C, cytosol ; P, plastid ; M, mitochondria ; MP, mitochondria and plastid ; ER, Endoplasmic Reticulum ; V, vacuole ; Pxp, pexophagosome. Values are expressed as Log2 of spectral counts and normalized (single hits were previously removed). Results were color coded as a function of values from yellow to red with Excel.
21 a b Supplemental Figure 16. LEA2 and LEA38 sequence alignment. Sequences were aligned using the MultAlin software ( using default parameters. The targeting peptide cleavage sites of LEA38 shown as black arrows were predicted using the MitoProt (a) and TargetP (b) software.
22 PsLEAM ( ) Mitochondria LEA36 ( ) Cytosol LEA9 ( ) Pexophagosome LEA11( ) Plastid LEA30 ( ) ER LEA42 (93-128) Mitochondria-Plastid Supplemental Figure 17. Modeling of class A α-helix motif of LEA proteins. Helical projections of α-helices were obtained using the HeliQuest webserver ( ). Each wheel was obtained with a 36 amino acids window. Residues are color coded with blue for positively (K, R) and red (D, E) for positively charged residues. Non polar residues are shown in yellow or grey, and others in purple, light blue or light pink colors. The arrow shows the hydrophobic moment.
23 LEA23 (61-96) Plastid LEA23 ( ) Plastid LEA24 (61-96) Plastid LEA24 (87-122) Plastid Supplemental Figure 18. Modeling of class A-like α-helix motif of LEA23 and LEA24. Helical projections of α-helices were obtained using the HeliQuest webserver ( ). Each wheel was obtained with a 36 amino acids window. Residues are color coded with blue for positively (K, R) and red (D, E) for positively charged residues. Non polar residues are shown in yellow or grey, and others in purple, light blue or light pink colors. The arrow shows the hydrophobic moment.
24 Supplemental Table 1. Theoretical and apparent molecular masses of LEA and fusion proteins. LEA protein Molecular weight (kda) Molecular weight with GFP (kda) Molecular weight with GFP deduced from Western blot (kda)
25 Supplemental Table 2. Quantitative data about transiently transformed protoplasts. Exp, independent transformation; FP, fluorescent protoplast number protoplasts Exp 1 Exp 2 Exp 3 Exp 4 Exp 5 LEA number of Protopl. Protopl. Protopl. Protopl. Protopl. Total independant FP FP FP FP FP observed observed observed observed observed FP transformation
26 Supplemental Table 3. Arabidopsis LEA templates used for PCR amplification. LEA# Accession number (TAIR) Template Origin 1 At1g01470 cdna Arabidopsis seeds MPIMP 2 At1g02820 pentr/sd/d-topo (+CDS) MPIMP 3 At1g03120 cdna Arabidopsis seeds MPIMP 4 At1g20440 pentr/sd/d-topo (+CDS) MPIMP 5 At1g20450 puni51 (U24382) ABRC 6 At1g32560 pentr/sd/d-topo (+CDS) MPIMP 7 At1g52690 pentr/sd/d-topo (+CDS) MPIMP 8 At1g54410 pentr/sd/d-topo (+CDS) MPIMP 9 At1g72100 pentr/sd/d-topo (+CDS) MPIMP 10 At1g76180 pentr/sd/d-topo (+CDS) MPIMP 11 At2g03740 pentr/sd/d-topo (+CDS) MPIMP 12 At2g03850 pentr/sd/d-topo (+CDS) MPIMP 13 At2g18340 cdna Arabidopsis seeds MPIMP 14 At2g21490 pentr/sd/d-topo (+CDS) MPIMP 15 At2g23110 pentr/sd/d-topo (+CDS) MPIMP 16 At2g23120 pentr/sd/d-topo (+CDS) MPIMP 17 At2g33690 puni51 (U63047) ABRC 18 At2g35300 cdna Arabidopsis seeds MPIMP 19 At2g36640 cdna Arabidopsis seeds MPIMP 20 At2g40170 pentr/sd/d-topo (+CDS) MPIMP 21 At2g41260 puc57 (synthetic gene) GenScript 22 At2g41280 pentr/sd/d-topo (+CDS) MPIMP 23 At2g42530 pentr/sd/d-topo (+CDS) MPIMP 24 At2g42540 pentr/sd/d-topo (+CDS) MPIMP 25 At2g42560 pentr/sd/d-topo (+CDS) MPIMP 26 At2g44060 puni51 (U83572) ABRC 27 At2g46140 pentr/sd/d-topo (+CDS) MPIMP 28 At3g02480 pentr/sd/d-topo (+CDS) MPIMP 29 At3g15670 pentr/sd/d-topo (+CDS) MPIMP 30 At3g17520 pentr/sd/d-topo (+CDS) MPIMP 31 At3g22490 pentr/sd/d-topo (+CDS) MPIMP 32 At3g22500 pentr/sd/d-topo (+CDS) MPIMP 33 At3g50970 pentr/sd/d-topo (+CDS) MPIMP 34 At3g50980 cdna Arabidopsis seeds MPIMP 35 At3g51810 pentr/sd/d-topo (+CDS) MPIMP 36 At3g53040 pentr/sd/d-topo (+CDS) MPIMP 37 At3g53770 pentr221 (pentr221-at3g53770) ABRC 38 At4g02380 pentr/sd/d-topo (+CDS) MPIMP 39 At4g13230 cdna Arabidopsis bud MPIMP 40 At4g13560 pentr/sd/d-topo (+CDS) MPIMP 41 At4g15910 puni51 (U23419) ABRC 42 At4g21020 puni51 (U20045) ABRC 43 At4g36600 puni51 (S63786) ABRC 44 At4g38410 pentr/sd/d-topo (+CDS) MPIMP 45 At4g39130 cdna Arabidopsis seeds MPIMP 46 At5g06760 pentr/sd/d-topo (+CDS) MPIMP 47 At5g27980 puc57 (synthetic gene) GenScript 48 At5g44310 puni51 (U66244) ABRC 49 At5g53260 puc57 (synthetic gene) GenScript 50 At5g53270 cdna Arabidopsis seeds MPIMP 51 At5g66400 pentr/sd/d-topo (+CDS) MPIMP
27 Supplemental Table 4. Primers used for the construction of LEA gene fusions and markers. Sequence Name Forward primer Length Name Reverse primer Length LEA1 LEA2 LEA3 LEA4 LEA5 LEA6 LEA7 LEA8 LEA9 LEA10 LEA11 LEA12 LEA13 LEA14 LEA15 LEA16 LEA17 LEA18 LEA19 LEA20 LEA21 LEA22 LEA23 LEA24 LEA25 LEA26 LEA27 LEA28 LEA29 LEA30 LEA31 LEA32 LEA33 LEA34 LEA35 LEA36 FL1 CACCATGGCGAGCTTGCTAGAT 22 RL1 GAAGAAATCTTTAAAAGTAGG 21 FnoL1 CACCGCGAGCTTGCTAGATAAA 22 RsL1 TCAGAAGAAATCTTTAAAAGT 21 FL2 CACCATGGCTCGTTCTCTCGCT 22 RL2 TTGCTTGTTGTTCAAGAG 18 FnoL2 CACCGCTCGTTCTCTCGCTAAC 22 RsL2 TTATTGCTTGTTGTTCAA 18 FL3 CACCATGGCACAGCATCAGCATTC 24 RL3 GAGTTGTTGATTGAGCCT 18 FnoL3 CACCGCACAGCATCAGCATTCTCC 24 RsL3 CTAGAGTTGTTGATTGAGC 19 FL4 CACCATGGCTGAGGAGTACAAG 22 RL4 ATCATCAGACTCTTTTTCTTTC 22 FnoL4 CACCGCTGAGGAGTACAAGAAC 22 RsL4 TTAATCATCAGACTCTTTTTCT 22 FL5 CACCATGGCAGAAGAGTACAAGAAC 25 RL5 ATCAGACACTTTTTCTTTCTTC 22 FnoL5 CACCGCAGAAGAGTACAAGAACACC 25 RsL5 TTAATCAGACACTTTTTCTTTC 22 FL6 CACCATGCAATCGGCGAAACAG 22 RL6 GTAGTGATGATGATTATGATGTCC 24 FnoL6 CACCCAATCGGCGAAACAGAAG 22 RsL6 TTAGTAGTGATGATGATTATGATG 24 FL7 CACCATGGCGTCTCATCAAGAA 22 RL7 CTTCCTCTGTGTCTCACG 18 FnoL7 CACCGCGTCTCATCAAGAACAG 22 RsL7 TCACTTCCTCTGTGTCTC 18 FL8 CACCATGGCAGGACTCATCAAC 22 RL8 ATCGCTGTCGCTGTCACT 18 FnoL8 CACCGCAGGACTCATCAACAAG 22 RsL8 TTAATCGCTGTCGCTGTC 18 FL9 CACCATGACGAATCTTTTGGCC 22 RL9 ACAAGCGGCGCCGAGTCTCTG 21 FnoL9 CACCACGAATCTTTTGGCCTTG 22 RsL9 TCAACAAGCGGCGCCGAGTCT 21 FL10 CACCATGGCTGAGGAAATCAAG 22 RL10 TTCTTTATCTTTCTTCTCCTCC 22 FnoL10 CACCGCTGAGGAAATCAAGAAT 22 RsL10 TTATTCTTTATCTTTCTTCTCC 22 FL11 CACCATGTCGATCTCCGGAGCT 22 RL11 TGCATCAGTTTTTGGAGGC 19 FnoL11 CACCTCGATCTCCGGAGCTGTG 22 RsL11 CTATGCATCAGTTTTTGGA 19 FL12 CACCATGGCGATGTCGATCTCC 22 RL12 AGTTTTTGGAGGCATTATAGC 21 FnoL12 CACCGCGATGTCGATCTCCGGA 22 RsL12 TTAAGTTTTTGGAGGCATTAT 21 FL13 CACCATGATGGAGAGAAGAAGAAC 24 RL13 GAGCTCAGCATAACGGTC 18 FnoL13 CACCATGGAGAGAAGAAGAACGG 23 RsL13 TTAGAGCTCAGCATAACG 18 FL14 CACCATGGCGGATTTGAGGGAC 22 RL14 TGGGTGGTTGTGGTTATG 18 FnoL14 CACCGCGGATTTGAGGGACGAA 22 RsL14 TCATGGGTGGTTGTGGTT 18 FL15 CACCATGGAGGATCAGAAAAAGCC 24 RL15 CGGAACGCCCTGACGGTT 18 FnoL15 CACCGAGGATCAGAAAAAGCCACC 24 RsL15 TCACGGAACGCCCTGACG 18 FL16 CACCATGGAGGCCGGGAAAACA 22 RL16 CGGAGCTTTCGCATCGGT 18 FnoL16 CACCGAGGCCGGGAAAACACCA 22 RsL16 TCACGGAGCTTTCGCATC 18 FL17 CACCATGTCGAAGAGTGAAGAGAA 24 RL17 CTTCTTAGCTTTTTGGTTAGCG 22 FnoL17 CACCTCGAAGAGTGAAGAGAAACA 24 RsL17 TCACTTCTTAGCTTTTTGGTTA 22 FL18 CACCATGCAGTCGGCGAAGGAA 22 RL18 GATCTGTCCCGGCGGGTA 18 FnoL18 CACCCAGTCGGCGAAGGAAAAG 22 RsL18 TTAGATCTGTCCCGGCGG 18 FL19 CACCATGGCGTCAGACAAACAAAA 24 RL19 CAGCTTTCCCTTATCTTTCC 20 FnoL19 CACCGCGTCAGACAAACAAAAGGC 24 RsL19 TCACAGCTTTCCCTTATCTT 20 FL20 CACCATGGCGTCTCAACAAGAG 22 RL20 GGTCTTGGTCCTGAATTTGG 20 FnoL20 CACCGCGTCTCAACAAGAGAAG 22 RsL20 TTAGGTCTTGGTCCTGAATT 20 FL21 CACCATGGGAAACCTCAAGTCTC 23 RL21 TGGCTTAGCTTGTTGGGC 18 FnoL21 CACCGGAAACCTCAAGTCTCTCG 23 RsL21 TCATGGCTTAGCTTGTTGG 19 FL22 CACCATGGGAAACCTCATGTCT 22 RL22 TGGCTTAGCTTCTTCCTT 18 FnoL22 CACCGGAAACCTCATGTCTCTC 22 RsL22 TCATGGCTTAGCTTCTTCC 19 FL23 CACCATGGCGATGTCTTTATCAGG 24 RL23 GGACTTTGTGGCATTCTTAG 20 FnoL23 CACCGCGATGTCTTTATCAGGAGC 24 RsL23 TCAGGACTTTGTGGCATTCT 20 FL24 CACCATGGCGATGTCTTTCTCAGG 24 RL24 CTTTGTGGCATCCTTAGC 18 FnoL24 CACCGCGATGTCTTTCTCAGGAGC 24 RsL24 CTACTTTGTGGCATCCTT 18 FL25 CACCATGGCGTCAGAGCAAGCA 22 RL25 ACGTTGTCCATGTTCCCG 18 FnoL25 CACCGCGTCAGAGCAAGCAAGG 22 RsL25 TCAACGTTGTCCATGTTC 18 FL26 CACCATGTCGACATCTGAGGATAA 24 RL26 TTCCTCATCGTCGTCATC 18 FnoL26 CACCTCGACATCTGAGGATAAACC 24 RsL26 TTATTCCTCATCGTCGTC 18 FL27 CACCATGGCATCAGCGGATGAA 22 RL27 AAAGAAGTCGCGAAGGGA 18 FnoL27 CACCGCATCAGCGGATGAAAAG 22 RsL27 TTAAAAGAAGTCGCGAAG 18 FL28 CACCATGGACAACAAGCAAAACGC 24 RL28 GTGGCTTTTGTTCATGCC 18 FnoL28 CACCGACAACAAGCAAAACGCGAG 24 RsL28 TTAGTGGCTTTTGTTCAT 18 FL29 CACCATGGCATCCAACCAACAG 22 RL29 CTTCCTCTGATAAGTCTGATGA 22 FnoL29 CACCGCATCCAACCAACAGAGC 22 RsL29 TCACTTCCTCTGATAAGTCTGA 22 FL30 CACCATGGGGTTAGAGAGGAAA 22 RL30 GAGCTCAGCATCATCGTC 18 FnoL30 CACCGGGTTAGAGAGGAAAGTG 22 RsL30 TCAGAGCTCAGCATCATC 18 FL31 CACCATGAGTCAAGAAGAACAACC 24 RL31 TATATCAGCTCTCTCGTTAAGC 22 FnoL31 CACCAGTCAAGAAGAACAACCAAA 24 RsL31 TCATATATCAGCTCTCTCGTTA 22 FL32 CACCATGAGCCAAGAGCAACCA 22 RL32 TATATCAACTCTCTCGTTAAGC 22 FnoL32 CACCAGCCAAGAGCAACCAAGG 22 RsL32 TCATATATCAACTCTCTCGTTA 22 FL33 CACCATGAATTCTCACCAGAATCA 24 RL33 GTGATGACCACCGGGAAG 18 FnoL33 CACCAATTCTCACCAGAATCAAAC 24 RsL33 CTAGTGATGACCACCGGG 18 FL34 CACCATGGAGTCTTACCAAAACC 23 RL34 ATGATGACCACCCGGAAG 18 FnoL34 CACCGAGTCTTACCAAAACCAG 22 RsL34 CTAATGATGACCACCCGG 18 FL35 CACCATGGCGTCAAAGCAACTG 22 RL35 CTTGTTGGTGAACTTTGACTC 21 FnoL35 CACCGCGTCAAAGCAACTGAGC 22 RsL35 TCACTTGTTGGTGAACTTTGA 21 FL36 CACCATGGCATCAGGACAACGG 22 RL36 CAGCTTTTTCTCTCCCAC 18 FnoL36 CACCGCATCAGGACAACGGGAG 22 RsL36 CTACAGCTTTTTCTCTCC 18
28 LEA37 FL37 CACCATGTCTCAATCACTTTTCAA 24 RL37 GCTCACTACGTACGTCTTTTG 21 FnoL37 CACCTCTCAATCACTTTTCAATCT 24 RsL37 TTAGCTCACTACGTACGTCTT 21 LEA38 FL38 CACCATGGCTCGTTCTATCTCTAA 24 RL38 CTGCTTGTTGTTCAAGAGAGC 21 FnoL38 CACCGCTCGTTCTATCTCTAACGT 24 RsL38 TCACTGCTTGTTGTTCAAGAG 21 LEA39 FL39 CACCATGACAAGCTTCGCGGTC 22 RL39 TTTGAGGTTCTTGGTGTTC 19 FnoL39 CACCACAAGCTTCGCGGTCGTTG 23 RsL39 TTATTTGAGGTTCTTGGTG 19 LEA40 FL40 CACCATGTCGCAACAACAATTC 22 RL40 TTTCTTCTCGTTCATGCC 18 FnoL40 CACCTCGCAACAACAATTCAAC 22 RsL40 TCATTTCTTCTCGTTCAT 18 LEA41 FL41 CACCATGGCCGCTCGTTCACTC 22 RL41 GAAAGACTTTGCTTTGTTTTTC 22 FnoL41 CACCGCCGCTCGTTCACTCTCC 22 RsL41 TCAGAAAGACTTTGCTTTGTTT 22 LEA42 FL42 CACCATGGCGGCCATGCAACTAAC 24 RL42 GTTAAATGTTATGAAATCGTCC 22 FnoL42 CACCGCGGCCATGCAACTAACAAG 24 RsL42 TCAGTTAAATGTTATGAAATCGTC 24 LEA43 FL43 CACCATGATGCTTACGACGGTG 22 RL43 AAGCTCAGCGCTACGGTC 18 FnoL43 CACCATGCTTACGACGGTGGTG 22 RsL43 CTAAAGCTCAGCGCTACG 18 LEA44 FL44 CACCATGGCGGATCATCCTCGT 22 RL44 GGTTTCCTTCTTCTTCTCATC 21 FnoL44 CACCGCGGATCATCCTCGTTCT 22 RsL44 CTAGGTTTCCTTCTTCTTCTC 21 LEA45 FL45 CACCATGGCGGATCTGAAAGAC 22 RL45 AAGATCATTATGGTGGCC 18 FnoL45 CACCGCGGATCTGAAAGACGAA 22 RsL45 TCAAAGATCATTATGGTG 18 LEA46 FL46 CACCATGCAGTCGATGAAAGAA 22 RL46 TCCAGTATATCCCCCGCC 18 FnoL46 CACCCAGTCGATGAAAGAAACA 22 RsL46 TTATCCAGTATATCCCCC 18 LEA47 FL47 CACCATGAGCGAAGAACAGCTG 22 RL47 TTTGGACTGATTGATCCG 18 FnoL47 CACCAGCGAAGAACAGCTGCAG 22 RsL47 TCATTTGGACTGATTGAT 18 LEA48 FL48 CACCATGGCGGCTATGCAGTTAACG 25 RL48 GAACCTCTTGAAATCATCATCC 22 FnoL48 CACCGCGGCTATGCAGTTAACGAG 24 RsL48 TTAGAACCTCTTGAAATCATCATC 24 LEA49 FL49 CACCATGGGTTCATCAAAAGATAG 24 RL49 AAGAGACGATGGATCGTG 18 FnoL49 CACCGGTTCATCAAAAGATAGTGC 24 RsL49 TCAAAGAGACGATGGATCG 19 LEA50 FL50 CACCATGATGTTCGGGTTCGGC 22 RL50 AAGAGATACATTGCATGG 18 FnoL50 CACCATGTTCGGGTTCGGCCTT 22 RsL50 TCAAAGAGATACATTGCA 18 LEA51 FL51 CACCATGGCGTCTTACCAGAAC 22 RL51 ACGGCCACCACCGGGAAG 18 FnoL51 CACCGCGTCTTACCAGAACCGT 22 RsL51 TTAACGGCCACCACCGGG 18 U16034 (At3g10920) MSD1-F CACCATGGCGATTCGTTGTGTAGC 24 MSD1-R GTTGTTTTCCTTCTCATAAACCTC 24 U12392 (At4g05180) PsbQ-F CACCATGGCTCAAGCAGTGACTTCG 25 PsbQ-R ACCGAGCTTGGCAAGAACATTGTTC 25 U13389 (At5g38430) RBCS1B-F CACCATGGCTTCCTCTATGCTCT 23 RBCS1B-R AGCATCAGTGAAGCTTGG 18 p2fgw7 FpGX GCGAAACCCTATAAGAACC 19 RpGX ACCACTACCAGCAGAACA 18 p2gwf7 FpXG GTGGTGCAGATGAACTTCAG 20 RpXG GCACAATCCCACTATCCTTC 20 RevEGFP TTACTTGTACAGCTCGTCCAT 21 p2gwr7 RFP-F CACCATGGAGGGCTCCGTGAACGG 24 RevRFP TTAGGCGCCGGTGGAGTGGC 20 NLS-F CACCATGCCACCAAAAAAGAAAAGAAAGGTT 31 NLS-R AACCTTTCTTTTCTTTTTTGGTGGCAT 27 PDHA1 (NM_000284) PDHE1-F ATGAGGAAGATGCTCGCC 18 PDHE1t-R CCTGGTGAGCACTGTTGTGA 20
Supplemental Data. Wang et al. (2013). Plant Cell /tpc
 Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplemental Data. Wu et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
Open Flower. Juvenile leaf Flowerbud. Carpel 35 NA NA NA NA 61 NA 95 NA NA 15 NA 41 3 NA
 PaxDB Root Juvenile leaf Flowerbud Open Flower Carpel Mature Pollen Silique Seed Sec3a Sec3b Sec5a Sec5b Sec6 Sec8 Sec10a/b Sec15a Sec15b Exo84a Exo84b Exo84c Exo70A1 Exo70A2 Exo70A3 49 47 8 75 104 79
PaxDB Root Juvenile leaf Flowerbud Open Flower Carpel Mature Pollen Silique Seed Sec3a Sec3b Sec5a Sec5b Sec6 Sec8 Sec10a/b Sec15a Sec15b Exo84a Exo84b Exo84c Exo70A1 Exo70A2 Exo70A3 49 47 8 75 104 79
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering
 nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
nature methods Organelle-specific, rapid induction of molecular activities and membrane tethering Toru Komatsu, Igor Kukelyansky, J Michael McCaffery, Tasuku Ueno, Lidenys C Varela & Takanari Inoue Supplementary
Cell wall components:
 Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. The Cell Wall The primary cell wall is capable of rapid expansion during
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. The Cell Wall The primary cell wall is capable of rapid expansion during
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure.
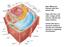 Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. Animal cells have a lysosome (related to vacuole) and centrioles (function
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. Animal cells have a lysosome (related to vacuole) and centrioles (function
SUPPLEMENTARY INFORMATION. Supplementary Figures
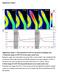 SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Practice Exam 2 MCBII
 1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
Supplementary Figures
 Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Supporting Information
 Supporting Information Lee et al. 10.1073/pnas.0910950106 Fig. S1. Fe (A), Zn (B), Cu (C), and Mn (D) concentrations in flag leaves from WT, osnas3-1, and OsNAS3-antisense (AN-2) plants. Each measurement
Supporting Information Lee et al. 10.1073/pnas.0910950106 Fig. S1. Fe (A), Zn (B), Cu (C), and Mn (D) concentrations in flag leaves from WT, osnas3-1, and OsNAS3-antisense (AN-2) plants. Each measurement
Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70.
 Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70. peptide MH+ charge XCorr score Delta Cn score Protein ID KNAVVTVPAYFNDSQRQ 1680.83 2 2.22 0.37
Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70. peptide MH+ charge XCorr score Delta Cn score Protein ID KNAVVTVPAYFNDSQRQ 1680.83 2 2.22 0.37
Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human
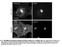 Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human LPP3wt-GFP, fixed and stained for GM130 (A) or Golgi97
Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human LPP3wt-GFP, fixed and stained for GM130 (A) or Golgi97
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
A Tour of the Cell Lecture 2, Part 1 Fall 2008
 Cell Theory 1 A Tour of the Cell Lecture 2, Part 1 Fall 2008 Cells are the basic unit of structure and function The lowest level of structure that can perform all activities required for life Reproduction
Cell Theory 1 A Tour of the Cell Lecture 2, Part 1 Fall 2008 Cells are the basic unit of structure and function The lowest level of structure that can perform all activities required for life Reproduction
Cells Alive- Internet Lesson
 Name Date Period Cells Alive Webquest Cells Alive- Internet Lesson URL: www.cellsalive.com Objective: You will look at computer models of cells, learn the functions and the descriptions of the cells and
Name Date Period Cells Alive Webquest Cells Alive- Internet Lesson URL: www.cellsalive.com Objective: You will look at computer models of cells, learn the functions and the descriptions of the cells and
Lecture 6 9/17 Dr. Hirsh Organization of Cells, continued
 Cell structure of Eukaryotic cells Lecture 6 9/17 Dr. Hirsh Organization of Cells, continued Lots of double-membraned organelles Existence of an Endo-membrane system separation of areas of cell, transport
Cell structure of Eukaryotic cells Lecture 6 9/17 Dr. Hirsh Organization of Cells, continued Lots of double-membraned organelles Existence of an Endo-membrane system separation of areas of cell, transport
Supplemental Data. Hao et al. (2014). Plant Cell /tpc
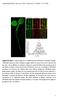 Supplemental Figure 1. Confocal Images and VA-TIRFM Analysis of GFP-RbohD in Arabidopsis Seedlings. (A) RbohD expression in whole Arabidopsis seedlings. RbohD was expressed in the leaves, hypocotyl, and
Supplemental Figure 1. Confocal Images and VA-TIRFM Analysis of GFP-RbohD in Arabidopsis Seedlings. (A) RbohD expression in whole Arabidopsis seedlings. RbohD was expressed in the leaves, hypocotyl, and
Supplementary information
 Supplementary information Supplementary Figure 1: Components of Arabidopsis tricarboxylic acid (TCA) cycle. Schematic summary of the TCA cycle and the enzymes related to the reactions. The large text and
Supplementary information Supplementary Figure 1: Components of Arabidopsis tricarboxylic acid (TCA) cycle. Schematic summary of the TCA cycle and the enzymes related to the reactions. The large text and
Supplementary Information
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2018 Supplementary Information Spying on Protein Interactions in Living Cells with Reconstituted Scarlet
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2018 Supplementary Information Spying on Protein Interactions in Living Cells with Reconstituted Scarlet
Expression constructs
 Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Supplemental Information. Figures. Figure S1
 Supplemental Information Figures Figure S1 Identification of JAGGER T-DNA insertions. A. Positions of T-DNA and Ds insertions in JAGGER are indicated by inverted triangles, the grey box represents the
Supplemental Information Figures Figure S1 Identification of JAGGER T-DNA insertions. A. Positions of T-DNA and Ds insertions in JAGGER are indicated by inverted triangles, the grey box represents the
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Supplementary Figure S1
 Supplementary Figure S1 Supplementary Figure S1. PARP localization patterns using GFP-PARP and PARP-specific antibody libraries GFP-PARP localization in non-fixed (A) and formaldehyde fixed (B) GFP-PARPx
Supplementary Figure S1 Supplementary Figure S1. PARP localization patterns using GFP-PARP and PARP-specific antibody libraries GFP-PARP localization in non-fixed (A) and formaldehyde fixed (B) GFP-PARPx
crossmark Ca V subunits interact with the voltage-gated calcium channel
 crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Cytology II Study of Cells
 Cytology II Study of Cells Biology 20 Cellular Basis of Life 1. Basic unit of Life 2. Composed of one or more cells 3. Arises from pre-existing cells Asexual (Mitosis)/Sexual (Meiosis) 4. Surrounded by
Cytology II Study of Cells Biology 20 Cellular Basis of Life 1. Basic unit of Life 2. Composed of one or more cells 3. Arises from pre-existing cells Asexual (Mitosis)/Sexual (Meiosis) 4. Surrounded by
General Biology. The Fundamental Unit of Life The Cell. All organisms are made of cells The cell is the simplest collection of matter that can live
 General Biology Course No: BNG2003 Credits: 3.00 3. A Tour of the Cell Prof. Dr. Klaus Heese The Fundamental Unit of Life The Cell All organisms are made of cells The cell is the simplest collection of
General Biology Course No: BNG2003 Credits: 3.00 3. A Tour of the Cell Prof. Dr. Klaus Heese The Fundamental Unit of Life The Cell All organisms are made of cells The cell is the simplest collection of
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb2822 a MTC02 FAO cells EEA1 b +/+ MEFs /DAPI -/- MEFs /DAPI -/- MEFs //DAPI c HEK 293 cells WCE N M C P AKT TBC1D7 Lamin A/C EEA1 VDAC d HeLa cells WCE N M C P AKT Lamin A/C EEA1 VDAC Figure
DOI:.38/ncb2822 a MTC02 FAO cells EEA1 b +/+ MEFs /DAPI -/- MEFs /DAPI -/- MEFs //DAPI c HEK 293 cells WCE N M C P AKT TBC1D7 Lamin A/C EEA1 VDAC d HeLa cells WCE N M C P AKT Lamin A/C EEA1 VDAC Figure
Figure legends of supplementary figures
 Figure legends of supplementary figures Figure 1. Phenotypic analysis of rice early flowering1 () plants and enhanced expression of floral identity genes in.. Leaf emergence of,, and plants with complementary
Figure legends of supplementary figures Figure 1. Phenotypic analysis of rice early flowering1 () plants and enhanced expression of floral identity genes in.. Leaf emergence of,, and plants with complementary
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
University of Groningen
 University of Groningen Mechanisms of Hemagglutinin Targeted Influenza Virus Neutralization Brandenburg, Boerries; Koudstaal, Wouter; Goudsmit, Jaap; Klaren, Vincent; Tang, Chan; Bujny, Miriam V.; Korse,
University of Groningen Mechanisms of Hemagglutinin Targeted Influenza Virus Neutralization Brandenburg, Boerries; Koudstaal, Wouter; Goudsmit, Jaap; Klaren, Vincent; Tang, Chan; Bujny, Miriam V.; Korse,
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Supplementary Figure 1.
 Supplementary Figure 1. Visualization of endoplasmic reticulum-mitochondria interaction by in situ proximity ligation assay. A) Illustration of targeted proteins in mitochondria (M), endoplasmic reticulum
Supplementary Figure 1. Visualization of endoplasmic reticulum-mitochondria interaction by in situ proximity ligation assay. A) Illustration of targeted proteins in mitochondria (M), endoplasmic reticulum
A Tour of the Cell. reference: Chapter 6. Reference: Chapter 2
 A Tour of the Cell reference: Chapter 6 Reference: Chapter 2 Monkey Fibroblast Cells stained with fluorescent dyes to show the nucleus (blue) and cytoskeleton (yellow and red fibers), image courtesy of
A Tour of the Cell reference: Chapter 6 Reference: Chapter 2 Monkey Fibroblast Cells stained with fluorescent dyes to show the nucleus (blue) and cytoskeleton (yellow and red fibers), image courtesy of
Rapid blue-light mediated induction of protein interactions in living cells
 Nature Methods Rapid blue-light mediated induction of protein interactions in living cells Matthew J Kennedy, Robert M Hughes, Leslie A Peteya, Joel W Schwartz, Michael D Ehlers & Chandra L Tucker Supplementary
Nature Methods Rapid blue-light mediated induction of protein interactions in living cells Matthew J Kennedy, Robert M Hughes, Leslie A Peteya, Joel W Schwartz, Michael D Ehlers & Chandra L Tucker Supplementary
A. Major parts 1. Nucleus 2. Cytoplasm a. Contain organelles (see below) 3. Plasma membrane (To be discussed in Cellular Transport Lecture)
 Lecture 5: Cellular Biology I. Cell Theory Concepts: 1. Cells are the functional and structural units of living organisms 2. The activity of an organism is dependent on both the individual and collective
Lecture 5: Cellular Biology I. Cell Theory Concepts: 1. Cells are the functional and structural units of living organisms 2. The activity of an organism is dependent on both the individual and collective
A Tour of the Cell. reference: Chapter 6. Reference: Chapter 2
 A Tour of the Cell reference: Chapter 6 Reference: Chapter 2 Monkey Fibroblast Cells stained with fluorescent dyes to show the nucleus (blue) and cytoskeleton (yellow and red fibers), image courtesy of
A Tour of the Cell reference: Chapter 6 Reference: Chapter 2 Monkey Fibroblast Cells stained with fluorescent dyes to show the nucleus (blue) and cytoskeleton (yellow and red fibers), image courtesy of
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation
 Lecture 3 I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation Ref: MBoC (5th Edition), Alberts Johnson Lewis Raff Roberts Walter Chapter 10 Membrane Structure Chapter 12 Intracellular
Lecture 3 I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation Ref: MBoC (5th Edition), Alberts Johnson Lewis Raff Roberts Walter Chapter 10 Membrane Structure Chapter 12 Intracellular
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2294 Figure S1 Localization and function of cell wall polysaccharides in root hair cells. (a) Spinning-disk confocal sections of seven day-old A. thaliana seedlings stained with 0.1% S4B
DOI: 10.1038/ncb2294 Figure S1 Localization and function of cell wall polysaccharides in root hair cells. (a) Spinning-disk confocal sections of seven day-old A. thaliana seedlings stained with 0.1% S4B
Essential Cell Biology
 Alberts Bray Hopkin Johnson Lewis Raff Roberts Walter Essential Cell Biology FOURTH EDITION Chapter 15 Intracellular Compartments and Protein Transport Copyright Garland Science 2014 CHAPTER CONTENTS MEMBRANE-ENCLOSED
Alberts Bray Hopkin Johnson Lewis Raff Roberts Walter Essential Cell Biology FOURTH EDITION Chapter 15 Intracellular Compartments and Protein Transport Copyright Garland Science 2014 CHAPTER CONTENTS MEMBRANE-ENCLOSED
Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 )
 770 771 Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 ) Human CXCL1 GCGCCCAAACCGAAGTCATA ATGGGGGATGCAGGATTGAG PF4 CCCCACTGCCCAACTGATAG TTCTTGTACAGCGGGGCTTG
770 771 Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 ) Human CXCL1 GCGCCCAAACCGAAGTCATA ATGGGGGATGCAGGATTGAG PF4 CCCCACTGCCCAACTGATAG TTCTTGTACAGCGGGGCTTG
Problem Set 5, 7.06, Spring of 13
 Problem Set 5, 7.06, Spring 2003 1 of 13 1. In order to please your demanding thesis advisor, you've completed an extensive fractionation and biochemical purification of proteins localized to the mitochondria,
Problem Set 5, 7.06, Spring 2003 1 of 13 1. In order to please your demanding thesis advisor, you've completed an extensive fractionation and biochemical purification of proteins localized to the mitochondria,
A. Generation and characterization of Ras-expressing autophagycompetent
 Supplemental Material Supplemental Figure Legends Fig. S1 A. Generation and characterization of Ras-expressing autophagycompetent and -deficient cell lines. HA-tagged H-ras V12 was stably expressed in
Supplemental Material Supplemental Figure Legends Fig. S1 A. Generation and characterization of Ras-expressing autophagycompetent and -deficient cell lines. HA-tagged H-ras V12 was stably expressed in
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
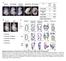 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Supplemental Figures:
 Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
A Tour of the cell. 2- Eukaryotic cells have internal membranes that compartmentalize their functions
 A Tour of the cell 1- To study cells, biologists use microscopes and the tools of biochemistry 2- Eukaryotic cells have internal membranes that compartmentalize their functions 3- The eukaryotic cell s
A Tour of the cell 1- To study cells, biologists use microscopes and the tools of biochemistry 2- Eukaryotic cells have internal membranes that compartmentalize their functions 3- The eukaryotic cell s
The Destination for Single-Pass Membrane Proteins Is Influenced Markedly by the Length of the Hydrophobic Domain
 The Plant Cell, Vol. 14, 1077 1092, May 2002, www.plantcell.org 2002 American Society of Plant Biologists The Destination for Single-Pass Membrane Proteins Is Influenced Markedly by the Length of the Hydrophobic
The Plant Cell, Vol. 14, 1077 1092, May 2002, www.plantcell.org 2002 American Society of Plant Biologists The Destination for Single-Pass Membrane Proteins Is Influenced Markedly by the Length of the Hydrophobic
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
A Tour of the Cell. Chapter 7
 A Tour of the Cell Chapter 7 Cytology: Study of Cells Light Microscopes uses light & a set of lenses Magnification ratio of object s image size to its real size Resolution measures the clarity of the image
A Tour of the Cell Chapter 7 Cytology: Study of Cells Light Microscopes uses light & a set of lenses Magnification ratio of object s image size to its real size Resolution measures the clarity of the image
Comparison of Membrane Targeting Strategies for the Accumulation of the Human Immunodeficiency Virus p24 Protein in Transgenic Tobacco
 Int. J. Mol. Sci. 2013, 14, 13241-13265; doi:10.3390/ijms140713241 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Comparison of Membrane Targeting
Int. J. Mol. Sci. 2013, 14, 13241-13265; doi:10.3390/ijms140713241 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Comparison of Membrane Targeting
Supplementary Figure 1 Validation of Per2 deletion in neuronal cells in N Per2 -/- mice. (a) Western blot from liver extracts of mice held under ad
 Supplementary Figure 1 Validation of Per2 deletion in neuronal cells in N Per2 -/- mice. (a) Western blot from liver extracts of mice held under ad libitum conditions detecting PER2 protein in brain and
Supplementary Figure 1 Validation of Per2 deletion in neuronal cells in N Per2 -/- mice. (a) Western blot from liver extracts of mice held under ad libitum conditions detecting PER2 protein in brain and
A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus
 Supplementary figures for: A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus officinalis Daisuke Tsugama, Kohei Matsuyama, Mayui Ide, Masato Hayashi, Kaien Fujino, and
Supplementary figures for: A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus officinalis Daisuke Tsugama, Kohei Matsuyama, Mayui Ide, Masato Hayashi, Kaien Fujino, and
Chapter 9 - Biological Membranes. Membranes form a semi-permeable boundary between a cell and its environment.
 Chapter 9 - Biological Membranes www.gsbs.utmb.edu/ microbook/ch037.htmmycoplasma Membranes form a semi-permeable boundary between a cell and its environment. Membranes also permit subcellular organization
Chapter 9 - Biological Membranes www.gsbs.utmb.edu/ microbook/ch037.htmmycoplasma Membranes form a semi-permeable boundary between a cell and its environment. Membranes also permit subcellular organization
Plant Cell, Development & Ultrastructure
 PCDU Lecture Plant Cell, Development & Ultrastructure Plant Cell Biology Labs Download at: http://goo.gl/111tha Microtubules 13 Protofilamente 25nm Plant MT Cytoskeleton Animal MT cytoskeleton Preprophase
PCDU Lecture Plant Cell, Development & Ultrastructure Plant Cell Biology Labs Download at: http://goo.gl/111tha Microtubules 13 Protofilamente 25nm Plant MT Cytoskeleton Animal MT cytoskeleton Preprophase
(Please activate your clickers)
 Organelles in prokaryotes and eukaryotes! Plasma membrane! Cell wall (in prokaryotes, plants, fungi, some protists)! Ribosomes! Nucleoid/nucleus! Mitochondria! Plastids (Please activate your clickers)
Organelles in prokaryotes and eukaryotes! Plasma membrane! Cell wall (in prokaryotes, plants, fungi, some protists)! Ribosomes! Nucleoid/nucleus! Mitochondria! Plastids (Please activate your clickers)
BIOSC 041. v Today s lecture. v Today s lab. v Note- Monday is a holiday good time to do some reading!
 BIOSC 041 v Today s lecture Review questions Chapter 6, Cells More review questions v Today s lab Quick review of lab safety The Scientific Method start thinking about which environments you might want
BIOSC 041 v Today s lecture Review questions Chapter 6, Cells More review questions v Today s lab Quick review of lab safety The Scientific Method start thinking about which environments you might want
Name: Per/row: Cell Structure and Function Practice: Use Ch 4 in Mader Biology
 Cell Structure and Function Practice: Use Ch 4 in Mader Biology Name: Per/row: 1. Write the name of the cell part in the box next to its description/function. Cell membrane Centrioles Chloroplast Chromatin
Cell Structure and Function Practice: Use Ch 4 in Mader Biology Name: Per/row: 1. Write the name of the cell part in the box next to its description/function. Cell membrane Centrioles Chloroplast Chromatin
Cell Structure and Organelles SBI4U 2016/10/14
 Cell Structure and Organelles SBI4U 2016/10/14 Inside the cell These are generalizations, not rules! Everything inside the cell membrane besides the nucleus is called the cytoplasm; The liquid is known
Cell Structure and Organelles SBI4U 2016/10/14 Inside the cell These are generalizations, not rules! Everything inside the cell membrane besides the nucleus is called the cytoplasm; The liquid is known
A Tour of the Cell. Chapter 6. Biology. Edited by Shawn Lester. Inner Life of Cell. Eighth Edition Neil Campbell and Jane Reece
 Chapter 6 A Tour of the Cell Inner Life of Cell Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin
Chapter 6 A Tour of the Cell Inner Life of Cell Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Thursday, October 16 th
 Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
The Cell. The building blocks of life
 The Cell The building blocks of life Learning Goals I can describe the cell theory. I can differentiate between a prokaryotic and eukaryotic cell. I can describe the similarities and differences between
The Cell The building blocks of life Learning Goals I can describe the cell theory. I can differentiate between a prokaryotic and eukaryotic cell. I can describe the similarities and differences between
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Problem Set #5 4/3/ Spring 02
 Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
The Fundamental Unit of Life The Cell. General Biology. All organisms are made of cells. The cell is the simplest collection of matter that can live
 Course No: BNG2003 Credits: 3.00 3. A Tour of the Cell General Biology The Fundamental Unit of Life The Cell All organisms are made of cells The cell is the simplest collection of matter that can live
Course No: BNG2003 Credits: 3.00 3. A Tour of the Cell General Biology The Fundamental Unit of Life The Cell All organisms are made of cells The cell is the simplest collection of matter that can live
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Homework Hanson section MCB Course, Fall 2014
 Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
Figure S1 Expression of AHL gene family members in diploid (Ler Col) and triploid (Ler
 Supplemental material Supplemental figure legends Figure S Expression of AHL gene family members in diploid (Ler ) and triploid (Ler osd) seeds. AHLs from clade B are labelled with (I), and AHLs from clade
Supplemental material Supplemental figure legends Figure S Expression of AHL gene family members in diploid (Ler ) and triploid (Ler osd) seeds. AHLs from clade B are labelled with (I), and AHLs from clade
Ch. 3 CELLS AND TISSUES. Copyright 2010 Pearson Education, Inc.
 Ch. 3 CELLS AND TISSUES Generalized Cell All cells: Human cells have three basic parts: Plasma membrane flexible outer boundary Cytoplasm intracellular fluid containing organelles Nucleus control center
Ch. 3 CELLS AND TISSUES Generalized Cell All cells: Human cells have three basic parts: Plasma membrane flexible outer boundary Cytoplasm intracellular fluid containing organelles Nucleus control center
Supplemental Data. Beck et al. (2010). Plant Cell /tpc
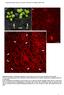 Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Draw and Complete the Chart.
 Draw and Complete the Chart. In this True/False Activity; you and your partner will discuss the question, record your response (both), and share your answer with the class. Be prepared to justify your
Draw and Complete the Chart. In this True/False Activity; you and your partner will discuss the question, record your response (both), and share your answer with the class. Be prepared to justify your
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Cytosol the fluid Cytoplasm cell interior, everything outside the nucleus but within the cell membrane, includes the organelles, cytosol, and
 Cell Organelles Plasma Membrane comprised of a phospholipid bilayer and embedded proteins Outer surface has oligosaccharides separates the cells s contents from its surroundings Cytosol the fluid Cytoplasm
Cell Organelles Plasma Membrane comprised of a phospholipid bilayer and embedded proteins Outer surface has oligosaccharides separates the cells s contents from its surroundings Cytosol the fluid Cytoplasm
Structure and Function of Cells
 Structure and Function of Cells Learning Outcomes Explain the cell theory Explain why cell size is usually very small Describe the Fluid Mosaic Model of membranes Describe similarities and differences
Structure and Function of Cells Learning Outcomes Explain the cell theory Explain why cell size is usually very small Describe the Fluid Mosaic Model of membranes Describe similarities and differences
Molecular Cell Biology 5068 In Class Exam 1 October 3, 2013
 Molecular Cell Biology 5068 In Class Exam 1 October 3, 2013 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your number
Molecular Cell Biology 5068 In Class Exam 1 October 3, 2013 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your number
Supplemental Information. Spatial Auxin Signaling. Controls Leaf Flattening in Arabidopsis
 Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Endomembrane system 11/1/2018. Endomembrane System. Direct physical continuity. Transfer of membrane segments as vesicles. Outer Nuclear envelope
 Endomembrane system Endomembrane System Outer Nuclear envelope Direct physical continuity Transfer of membrane segments as vesicles Endoplasmic reticulum BUT membranes are not identical in structure and
Endomembrane system Endomembrane System Outer Nuclear envelope Direct physical continuity Transfer of membrane segments as vesicles Endoplasmic reticulum BUT membranes are not identical in structure and
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Cell Theory states that: 1. All living things are made of cells 2. Cells are the basic unit of structure and function in living things 3. New cells are produced from
Cells Variation and Function of Cells Cell Theory states that: 1. All living things are made of cells 2. Cells are the basic unit of structure and function in living things 3. New cells are produced from
EGFR shrna A: CCGGCGCAAGTGTAAGAAGTGCGAACTCGAGTTCGCACTTCTTACACTTGCG TTTTTG. EGFR shrna B: CCGGAGAATGTGGAATACCTAAGGCTCGAGCCTTAGGTATTCCACATTCTCTT TTTG
 Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Nucleic acids. Nucleic acids are information-rich polymers of nucleotides
 Nucleic acids Nucleic acids are information-rich polymers of nucleotides DNA and RNA Serve as the blueprints for proteins and thus control the life of a cell RNA and DNA are made up of very similar nucleotides.
Nucleic acids Nucleic acids are information-rich polymers of nucleotides DNA and RNA Serve as the blueprints for proteins and thus control the life of a cell RNA and DNA are made up of very similar nucleotides.
AP Biology
 Tour of the Cell (1) 2007-2008 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Cell Size Why organelles? Specialized structures - specialized
Tour of the Cell (1) 2007-2008 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Cell Size Why organelles? Specialized structures - specialized
Early scientists who observed cells made detailed sketches of what they saw.
 Early scientists who observed cells made detailed sketches of what they saw. Early scientists who observed cells made detailed sketches of what they saw. CORK Early scientists who observed cells made detailed
Early scientists who observed cells made detailed sketches of what they saw. Early scientists who observed cells made detailed sketches of what they saw. CORK Early scientists who observed cells made detailed
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Main Functions maintain homeostasis
 The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Supporting Information
 Supporting Information Fig. S1. Overexpression of Rpr causes progenitor cell death. (A) TUNEL assay of control intestines. No progenitor cell death could be observed, except that some ECs are undergoing
Supporting Information Fig. S1. Overexpression of Rpr causes progenitor cell death. (A) TUNEL assay of control intestines. No progenitor cell death could be observed, except that some ECs are undergoing
Human height. Length of some nerve and muscle cells. Chicken egg. Frog egg. Most plant and animal cells Nucleus Most bacteria Mitochondrion
 10 m 1 m 0.1 m 1 cm Human height Length of some nerve and muscle cells Chicken egg Unaided eye 1 mm Frog egg 100 µm 10 µm 1 µm 100 nm 10 nm Most plant and animal cells Nucleus Most bacteria Mitochondrion
10 m 1 m 0.1 m 1 cm Human height Length of some nerve and muscle cells Chicken egg Unaided eye 1 mm Frog egg 100 µm 10 µm 1 µm 100 nm 10 nm Most plant and animal cells Nucleus Most bacteria Mitochondrion
Cell Structure and and Function Chapter 4
 Cell Structure and Function Chapter 4 Robert Hooke (1635-1703) 1703) Discovered cells by studying the cork layer of bark from an oak tree. Found cells when studied tree stems, roots, and leaves. Antony
Cell Structure and Function Chapter 4 Robert Hooke (1635-1703) 1703) Discovered cells by studying the cork layer of bark from an oak tree. Found cells when studied tree stems, roots, and leaves. Antony
A TOUR OF THE CELL 10/1/2012
 A TOUR OF THE CELL Chapter 6 KEY CONCEPTS: Eukaryotic cells have internal membranes that compartmentalize their functions The eukaryotic cell s genetic instructions are housed in the nucleus and carried
A TOUR OF THE CELL Chapter 6 KEY CONCEPTS: Eukaryotic cells have internal membranes that compartmentalize their functions The eukaryotic cell s genetic instructions are housed in the nucleus and carried
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
293T cells were transfected with indicated expression vectors and the whole-cell extracts were subjected
 SUPPLEMENTARY INFORMATION Supplementary Figure 1. Formation of a complex between Slo1 and CRL4A CRBN E3 ligase. (a) HEK 293T cells were transfected with indicated expression vectors and the whole-cell
SUPPLEMENTARY INFORMATION Supplementary Figure 1. Formation of a complex between Slo1 and CRL4A CRBN E3 ligase. (a) HEK 293T cells were transfected with indicated expression vectors and the whole-cell
10/5/2015. Cell Size. Relative Rate of Reaction
 The Cell Biology 102 Fundamental unit of life Smallest unit that displays all the basic elements of life Lecture 5: Cells Cell Theory 1. All living things are made of one or more cells Cell Theory 2. The
The Cell Biology 102 Fundamental unit of life Smallest unit that displays all the basic elements of life Lecture 5: Cells Cell Theory 1. All living things are made of one or more cells Cell Theory 2. The
