Spatio-genomic heterogeneity within localized, multi-focal prostate cancer
|
|
|
- Bernadette Casey
- 5 years ago
- Views:
Transcription
1 Spatio-genomic heterogeneity within localized, multi-focal prostate cancer Paul C. Boutros,1,2,3, Michael Fraser *,4, Nicholas J. Harding *,1, Richard de Borja *,1, Dominique Trudel *,5, Emilie Lalonde 1,2, Alice Meng 3, Pablo H. Hennings-Yeomans 1, Andrew McPherson 6, Veronica Y. Sabelnykova 1, Amin Zia 1, Natalie S. Fox 1,2, Julie Livingstone 1, Yu-Jia Shiah 1, Jianxin Wang 1, Timothy A. Beck 1, Cherry L. Have 5, Taryne Chong 1, Michelle Sam 1, Jeremy Johns 1, Lee Timms 1, Nicholas Buchner 1, Ada Wong 1, John D. Watson 1, Trent T. Simmons 1, Christine P ng 1, Gaetano Zafarana 4, Francis Nguyen 1, Xuemei Luo 1, Kenneth C. Chu 1, Stephenie D. Prokopec 1, Jenna Sykes 7, Alan Dal Pra 8, Alejandro Berlin 8, Andrew M. Brown 1, Michelle A. Chan- Seng-Yue 1, Fouad Yousif 1, Robert E. Denroche 1, Lauren C. Chong 1, Gregory M. Chen 1, Esther Jung 1, Clement Fung 1, Maud H.W. Starmans 1,9, Hanbo Chen 1, Shaylan K. Govind 1, James Hawley 1, Alister D Costa 1, Melania Pintilie 7, Daryl Waggott 1, Faraz Hach 6, Philippe Lambin 9, Lakshmi Muthuswamy 1,2, Colin S. Cooper 10,11, Rosalind Eeles 10,12, David E. Neal 13,14, Bernard Tetu 15, Cenk Sahinalp 6, Lincoln D. Stein 1, Neil Fleshner 16, Sohrab P. Shah 17,18,19, Colin C. Collins 20,21, Thomas J. Hudson 1,2, John D. McPherson 1,2, Theodorus van der Kwast 5, Robert G. Bristow 2,4,8, *These authors contributed equally to this work Corresponding authors: Paul.Boutros@oicr.on.ca and Rob.Bristow@rmp.uhn.on.ca 1 Ontario Institute for Cancer Research, Toronto, Ontario, Canada 2 Department of Medical Biophysics, University of Toronto, Toronto, Ontario, Canada 3 Department of Pharmacology & Toxicology, University of Toronto, Toronto, Ontario, Canada 4 Ontario Cancer Institute, Princess Margaret Cancer Centre/University Health Network, Toronto, Ontario, Canada 5 Department of Pathology and Laboratory Medicine, Toronto General Hospital/University Health Network, Toronto, Ontario, Canada 6 School of Computing Science, Simon Fraser University, Burnaby, British Columbia, Canada 7 Department of Biostatistics, Princess Margaret Cancer Centre/University Health Network, Toronto, Ontario, Canada 8 Department of Radiation Oncology, University of Toronto, Toronto, Ontario, Canada 9 Department of Radiotherapy, Maastricht University, Maastricht, The Netherlands 10 Division of Genetics and Epidemiology, The Institute Of Cancer Research, Sutton, UK Page 1 of 28
2 11 - Department of Biological Sciences and School of Medicine, University of East Anglia, Norwich, UK 12 - Royal Marsden NHS Foundation Trust, London and Sutton, UK 13 - Urological Research Laboratory, Cancer Research UK Cambridge Research Institute, Cambridge, UK 14 - Department of Surgical Oncology, University of Cambridge, Addenbrooke's Hospital, Cambridge, UK 15 - Department of Pathology, Laval University, Quebec City, Quebec, Canada 16 Division of Urology, Princess Margaret Cancer Centre/University Health Network, Toronto, Ontario, Canada 17 Department of Pathology, University of British Columbia, Vancouver, British Columbia, Canada 18 Department of Computer Science, University of British Columbia, British Columbia, Canada 19 British Columbia Cancer Agency Research Centre, Vancouver, British Columbia, Canada 20 Department of Urologic Sciences, University of British Columbia, Vancouver, British Columbia, Canada 21 Laboratory for Advanced Genome Analysis, Vancouver Prostate Centre, Vancouver, British Columbia, Canada Page 2 of 28
3 Supplementary Figures Supplementary Figure 1: Association of CNAs with Gleason Score We compared tumours with a Gleason Score of 3+4 (n=48) to those with a Gleason Score of 4+3 (n=27). There was no difference in median PGA (A), in the medium number of CNAs called (B) or in the frequencies of aberration of any individual gene s copy-number profile as shown by a volcano plot (C) and a histogram of unadjusted p-values (D). Page 3 of 28
4 Supplementary Figure 2: Association of CNAs with T-Category We compared tumours with a T-category T2a (n=48) to those with a T-category of T2b or T2c (n=27). There was no difference in median PGA (A), in the medium number of CNAs called (B) or in the frequencies of aberration of any individual gene s copy-number profile as shown by a volcano plot (C) and a histogram of unadjusted p-values (D). Page 4 of 28
5 Supplementary Figure 3: Association of CNAs with Pre-Treatment PSA We compared tumours with a pre-treatment PSA of 10 ng/ml or less (n=59) to those with a pretreatment PSA above 10 ng/ml (n=15 ). There was no difference in median PGA (A), in the medium number of CNAs called (B) or in the frequencies of aberration of any individual gene s copynumber profile as shown by a volcano plot (C) and a histogram of unadjusted p-values (D). Page 5 of 28
6 Supplementary Figure 4: MYCL1 Minimally Amplified Region MYCL1 amplified regions are shown for a representative subset of specimens. Lines indicate the region of copy number gain for the listed specimen. CPCG-0238 showed a larger ~1.1Mb amplification. CPCG-0189 showed the smallest gain, encompassing the minimally amplified region of ~8.5 kbp shown by the gray box. The median gain across all specimens was ~22 kbp (range 8.5kb 1.1 Mb). Page 6 of 28
7 Supplementary Figure 5: qpcr Validation of MYCL1 Amplification Custom TaqMan probes were designed to flank the MYCL1 locus on 1p34.3, across a region of 2 Mbp (+/- 1 Mbp on either side of the MYCL1 gene). Page 7 of 28
8 Supplementary Figure 6: Fluorescence in situ Hybridization (FISH) Analysis FISH was performed using probes directed to the MYCL1 locus on 1p34.3 (green) or to the centromeric region of chromosome 1 (CEP1; red). Results show analysis of benign prostate (FFPE) from a pathologically-verified, cancer-free cystoprostatectomy (A), showing 2 copies of MYCL1 and CEP1, or two FFPE prostate cancer specimens (CPCG-0364 and CPCG-0334), which were taken from a region of the index lesion immediately adjacent to the frozen specimens used for molecular analysis. Page 8 of 28
9 Supplementary Figure 7: Differential Gene Expression in MYCL1 Amplified Tumours We used mrna abundance microarrays to probe the downstream consequences of MYCL1 amplification. (A) shows large enrichment of differential expression in MYCL1-amplified tumours relative to non-mycl1 amplified tumours; (B) highlights some of the specific genes involved in a volcano plot; (C) and (D) shows similar differential abundance between MYCL1-amplified tumours and TP53-deleted tumours lacking MYCL1 amplification (note small sample sizes in this sub-group analysis lead to reduced statistical power). Page 9 of 28
10 Supplementary Figure 8: MYCL1 Clinical Correlates MYCL1 CNA status was not associated with pre-treatment PSA or T-category (A), but was inversely correlated to patient age and weakly associated with T2E events (B). All statistical analyses used Pearson s correlation or the point-biserial correlation. Page 10 of 28
11 Supplementary Figure 9: Patient-to-Cell-Line Transcriptome Profiles Two prostate cancer cell-lines (22RV1s and LNCAPs) had transfection of four separate MYCL1 isoforms, followed by mrna abundance profiling. The dotmaps on the left highlights genes showing statistical significance (q-value for the fold-change < 0.10) and large effect-sizes ( log 2 FoldChange > 1.5) in at least one cell-line (where both cell-lines meet these criteria, the larger effect-size is shown). The background shading gives the multiple-testing adjusted p-value, while the foreground dot-size and colour give the fold-change relative to empty-vector control. For each of these genes, we then compared their abundance in a subset of 16 MYCL1-wildtype and 8 MYCL1-amplified primary tumours that had whole-transcriptome profiling performed. The central heatmap gives abundance levels in each sample, while the red/white annotation bar at the top gives MYCL1 copy-number aberration status. Finally the two barplots on the right give the fold-change and p-value for each gene. In total 8/19 genes show p < 0.10 (relative to 2 expected by chance alone) even in this small patient cohort. Page 11 of 28
12 Supplementary Figure 10: Low-Input NGS Protocol Development of low-input library preparation Protocol. Table A displays the quality control metrics for three CPCG0184 samples as an example of the criteria used to evaluate library sequencing results. With the exception of # Lanes Sequenced and Sample Coverage, the values displayed are the average of the lane level metrics along with their standard deviation in brackets. PF Yield is the total number of pass filter bases available for alignment. Error Rate is the percentage of aligned bases which are not concordant with the reference. Reads/Start Point is a measure of library complexity it is the total number of reads divided by the number of unique mapping locations for those reads. Finally, Sample Coverage is the total filtered and collapsed coverage of the genome available for this sample from all lanes combined. Figure B shows a histogram of the insert distribution for an example lane from each of the three samples listed in Table A. Insert distribution is analogous to DNA fragment size; it is defined as the distance from the leftmost aligned base to the rightmost aligned based of a pair of reads. Page 12 of 28
13 Supplementary Figure 11: Genomic Rearrangement Profile of Index Lesions (A) The number of copy-number aberrations (top panel), the proportion of the genome altered (middle panel) and the number of genomic rearrangements of different types (bottom panel) for the index lesion of each patient, as determined by SNP microarray and whole-genome sequencing. (B) The genomic rearrangement profile of the index lesion of each sample across the genome, with the top frequency bar indicating the proportion of samples that had an aberration in a specific 1 Mbp bin of the genome. The covariates note the presence of genomic rearrangements in three large public datasets (Baca et al., Weischenfeldt et al. and Berger et al.). Page 13 of 28
14 Supplementary Figure 12: OncoScan SNV Validation Germline SNV calls from the OncoScan SNP array were compared to predictions from those made by WGS in blood or frozen tumour specimens (right panel) or in FFPE specimens (left panel). In each case, almost all samples showed over 90% concordance. Page 14 of 28
15 Supplementary Figure 13: Validation Re-Sequencing Coverage A heatmap of the coverage (log-scale) for each sample subjected to deep amplicon resequencing on an IonTorrent PGM. Each column corresponds to a sample (top covariates) and each row to an individual amplicon in the validation experiment. Page 15 of 28
16 Supplementary Figure 14: Germline Call Overlap For each tumour that had multiple regions sequenced, we compared the set of germline SNV calls between regions using Venn diagrams (A-D). For each tumour the vast majority of germline calls were common to all regions, providing a lower-bound on the error-rate of our calling pipeline. Page 16 of 28
17 Supplementary Figure 15: Intra-Focal Heterogeneity of CPCG0099 Visualization of mutation pattern over two foci of a Gleason 7 prostate tumour (CPCG0099). Multiple slices were taken for pathological examination, with both H&E and ERG staining shown. Note that no positional information is available for the bottom sample. Tissue from each region was subject to WGS, and common SNVs and CNAs are shown in the central panel. The right panel shows a subset of selected targetable mutations. Page 17 of 28
18 Supplementary Figure 16: Intra-Focal Heterogeneity of CPCG0102 Visualization of mutation pattern over three foci of a Gleason 7 prostate tumour (CPCG0102). Multiple slices were taken for pathological examination, with both H&E and ERG staining shown. Note that no positional information is available for the bottom sample. Tissue from each region was subject to WGS, and common SNVs and CNAs are shown in the central panel. The right panel shows a subset of selected targetable mutations. Page 18 of 28
19 Supplementary Figure 17: Intra-Focal Heterogeneity of CPCG0183 Visualization of mutation pattern over four foci of a Gleason 7 prostate tumour (CPCG0183). Multiple slices were taken for pathological examination, with both H&E and ERG staining shown. Note that no positional information is available for the bottom sample. Tissue from each region was subject to WGS, and common SNVs and CNAs are shown in the central panel. The right panel shows a subset of selected targetable mutations. Page 19 of 28
20 Supplementary Figure 18: Intra-Focal Heterogeneity of CPCG0184 Visualization of mutation pattern over five foci of a Gleason 7 prostate tumour (CPCG0184). Multiple slices were taken for pathological examination, with both H&E and ERG staining shown. Note that no positional information is available for the bottom sample. Tissue from each region was subject to WGS, and common SNVs and CNAs are shown in the central panel. The right panel shows a subset of selected targetable mutations. Page 20 of 28
21 Supplementary Figure 19: Phylogenetic Modeling Approach An overview of the phylogenetic modeling strategy, involving tree generation (A and B), followed by use of the blood-based reference sample as an outlier (C), forcing the tree to be ultrametric (i.e. every branch tip has equal distance to the root) (D) and finally removing the reference sample (E). Page 21 of 28
22 Supplementary Figure 20: CNA-Based Phylogeny of CPCG0099 log 2 Ratio plots for CPCG0099 of relative tumour/normal coverage across the entire genome (blue) with red lines indicating segmented CN status. Regions are clustered using phylogenetic methods. Page 22 of 28
23 Supplementary Figure 21: CNA-Based Phylogeny of CPCG0102 log 2 Ratio plots for CPCG0102 of relative tumour/normal coverage across the entire genome (blue) with red lines indicating segmented CN status. Regions are clustered using phylogenetic methods. Page 23 of 28
24 Supplementary Figure 22: CNA-Based Phylogeny of CPCG0183 log 2 Ratio plots for CPCG0183 of relative tumour/normal coverage across the entire genome (blue) with red lines indicating segmented CN status. Regions are clustered using phylogenetic methods. Numbers on the tree give bootstrap confidence in the branching patterns. Page 24 of 28
25 Supplementary Figure 23: CNA-Based Phylogeny of CPCG0184 log 2 Ratio plots for CPCG0184 of relative tumour/normal coverage across the entire genome (blue) with red lines indicating segmented CN status. Regions are clustered using phylogenetic methods. Numbers on the tree give bootstrap confidence in the branching patterns. Page 25 of 28
26 Supplementary Figure 24: Batch-Effects Removal in mrna Profiling Batch-effect removal in mrna data. Prior to batch-effect removal the two experimental batches cluster closely together (A), while after they show distinct patterns of mrna abundance (B). Page 26 of 28
27 Supplementary Table Legends Supplementary Table 1: Gene-Wise CNA Profiles for All Patients For each sample that received OncoScan SNP array interrogation of copy-number aberrations (n=75) this table gives for each gene whether it is amplified (1), deleted (-1) or unchanged (0). Additionally each gene is annotated with the Ensembl gene and transcript IDs, the chromosome, the starting and ending base-pair and the gene-symbol from both HUGO and HGNC. Supplementary Table 2: Regions of Recurrent CNAs GISTIC analysis of copy-number array data identified regions of recurrent copy-number alteration (rows). The columns give the name for each region, its chromosomal location (both arm and precise coordinates and probes involved) and statistical support (q-values, and amplitude estimates). For each patient, a coding of 0 (no event) vs. 1/2 (event) is given. Supplementary Table 3: GISTIC Genes Genes identified in recurrent GISTIC peaks are listed, along with their individual locations, Cytobands, q-values and gene-symbols. Supplementary Table 4: Validation of MYCL1 and MYC Amplification We performed quantitative PCR using probes directed to the putatively amplified regions of either MYCL1 or MYC, using a probe directed against RPPH1 (RNaseP H) as a control gene. Overall validation rates are shown. Supplementary Table 5: Summary of Flanking qpcr We performed qpcr analysis using the indicated probes, which flank the MYCL1 locus (which encompasses the probe shown in yellow) over a region of ~2 Mbp. NCI-H510A non-small cell lung cancer cells were used a positive control for MYCL1 amplification, since these cells contain a ~2.9 Mbp amplification of chromosome 1p, including the entire region covered by these probes. PC3 prostate cancer cells were used as a negative control. Supplementary Table 6: Genomic Instability Associated with MYC Family Gain For each Myc family member, we assessed the mean, median and standard-deviation of PGA and the total number of CNAs detected. Supplementary Table 7: Differential CNAs Associated with MYCL1 Amplification For each gene, we compared its frequency of CNA in MYCL1-amplified tumours vs. that in MYCamplified tumours. This table genes the gene ID (both Ensemble gene and transcript) along with genesymbols and genomic location. It lists the frequency of occurrence in MYCL1-amplified tumours, in MYC-amplified, the p-value from a proportion test, and the multiple-testing adjusted q-value. Page 27 of 28
28 Supplementary Table 8: MYCL1-Associated Transcriptome Dysregulation Comparison of tumours harbouring MYCL1-amplifications (n = 8) to those without (n=16) identified 294 genes showing differential abundance (q < 0.05, Bayesian-moderated t-test, see Methods). A list of gene-symbols for these genes is given here. Supplementary Table 9: Patient Annotation Key clinical information about each patient, including age at treatment, diagnostic Gleason Score, clinical T-category, Biochemical Recurrence status and ERG fusion status. Supplementary Table 10: Tumour Cellularity Analysis For each tumour sample subject to whole-genome sequencing tumour cellularity was assessed both by a urological pathologist (CellularityPath) and by the Qpure algorithm executed on SNP microarray data (CellularityQpure). Supplementary Table 11: Sequencing Statistics Overview of whole-genome sequencing. For each tumour and region, the collapsed coverage for blood (replicated for each region) and tumour are given, along with the input material type for the tumour sequencing, and the number of SNVs (of various functional categories), CNAs and Genomic Rearrangements. The number of somatic events in FFPE samples is elevated, likely due to artifacts of the FFPE procedure. Supplementary Table 12: All Genomic Rearrangements All detected Genomic Rearrangements, along with their chromosomal positions and a categorization of the rearrangement type, genes involved, and the score output from the destruct algorithm are given. Supplementary Table 13: Functional SNVs All detected functional somatic SNVs, along with their genomic locations, base-change and their status in each sequenced tumour region. Supplementary Table 14: WGA Effects Comparison of samples with and without WGA amplification based on the identity of SNPs detected by the OncoScan microarray platform. Supplementary Table 15: Pathway Analysis of MYCL1-Associated mrna Differences The GOEAST tool was used to assess functional enrichment amongst genes showing different mrna abundance between MYCL1-amplified and MYCL1-neutral tumours. Supplementary Table 16: Effects of WGA on SNP-Array Performance Comparison of concordance of SNP calls between matched WGA and non-wga specimens on the OncoScan array platform. Page 28 of 28
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Spatial genomic heterogeneity within localized, multifocal prostate cancer
 Spatial genomic heterogeneity within localized, multifocal prostate cancer npg 5 Nature America, Inc. All rights reserved. Paul C Boutros 3, Michael Fraser 4,3, Nicholas J Harding,3, Richard de Borja,3,
Spatial genomic heterogeneity within localized, multifocal prostate cancer npg 5 Nature America, Inc. All rights reserved. Paul C Boutros 3, Michael Fraser 4,3, Nicholas J Harding,3, Richard de Borja,3,
Improving Care for Prostate Cancer
 News Briefing Improving Care for Prostate Cancer Wednesday, Sept. 25, 2013 8:15 a.m. Colleen A.F. Lawton, MD, FASTRO Chairman, ASTRO Genomic Instability in Common Fragile Sites (CFSs) is Associated with
News Briefing Improving Care for Prostate Cancer Wednesday, Sept. 25, 2013 8:15 a.m. Colleen A.F. Lawton, MD, FASTRO Chairman, ASTRO Genomic Instability in Common Fragile Sites (CFSs) is Associated with
Supplementary Figure 1 Overall study design
 Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Assessment of Breast Cancer with Borderline HER2 Status Using MIP Microarray
 Assessment of Breast Cancer with Borderline HER2 Status Using MIP Microarray Hui Chen, Aysegul A Sahin, Xinyan Lu, Lei Huo, Rajesh R Singh, Ronald Abraham, Shumaila Virani, Bal Mukund Mishra, Russell Broaddus,
Assessment of Breast Cancer with Borderline HER2 Status Using MIP Microarray Hui Chen, Aysegul A Sahin, Xinyan Lu, Lei Huo, Rajesh R Singh, Ronald Abraham, Shumaila Virani, Bal Mukund Mishra, Russell Broaddus,
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies
 Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Supplemental Figure legends
 Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Interactive analysis and quality assessment of single-cell copy-number variations
 Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
CNV Detection and Interpretation in Genomic Data
 CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
Detection of copy number variations in PCR-enriched targeted sequencing data
 Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff
 5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff National molecular screening of patients with lung cancer for a national trial of multiple novel agents. 2000 NSCLC patients/year (late
5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff National molecular screening of patients with lung cancer for a national trial of multiple novel agents. 2000 NSCLC patients/year (late
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols.
 Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Nature Genetics: doi: /ng Supplementary Figure 1. Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms.
 Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Copy number and somatic mutations drive tumors
 Detection of copy number alterations, ploidy and loss of heterozygosity across the genome in FFPE specimens Utility for diagnosis and treatment with comparison to FISH-based and as a complement to sequencing
Detection of copy number alterations, ploidy and loss of heterozygosity across the genome in FFPE specimens Utility for diagnosis and treatment with comparison to FISH-based and as a complement to sequencing
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation
 NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
The common colorectal cancer predisposition SNP rs at chromosome 8q24 confers potential to enhanced Wnt signaling
 SUPPLEMENTARY INFORMATION The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling Sari Tuupanen 1, Mikko Turunen 2, Rainer Lehtonen 1, Outi
SUPPLEMENTARY INFORMATION The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling Sari Tuupanen 1, Mikko Turunen 2, Rainer Lehtonen 1, Outi
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Transform genomic data into real-life results
 CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University of Cambridge, Clara East, The Old Schools, Cambridge CB2 1TN
 AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
MPS for translocations
 MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
NGS ONCOPANELS: FDA S PERSPECTIVE
 NGS ONCOPANELS: FDA S PERSPECTIVE CBA Workshop: Biomarker and Application in Drug Development August 11, 2018 Rockville, MD You Li, Ph.D. Division of Molecular Genetics and Pathology Food and Drug Administration
NGS ONCOPANELS: FDA S PERSPECTIVE CBA Workshop: Biomarker and Application in Drug Development August 11, 2018 Rockville, MD You Li, Ph.D. Division of Molecular Genetics and Pathology Food and Drug Administration
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
gliomas. Fetal brain expected who each low-
 Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Nature Genetics: doi: /ng Supplementary Figure 1. Details of sequencing analysis.
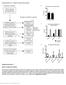 Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
Below, we included the point-to-point response to the comments of both reviewers.
 To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
 AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
AD Award Number: W81XWH-12-1-0480 TITLE: Molecular Characterization of Indolent Prostate Cancer PRINCIPAL INVESTIGATOR: Jun Luo, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins University, Baltimore, MD
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients
 The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics
 ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET
 Supplemental Figure Legends Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET Multiple unsupervised analyses were performed on human HT-1v4 expression array (Illumina) data from
Supplemental Figure Legends Figure 1 Gene expression profiles define 3 molecular sub-groups of CNS-PNET Multiple unsupervised analyses were performed on human HT-1v4 expression array (Illumina) data from
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer
 Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
NGS IN ONCOLOGY: FDA S PERSPECTIVE
 NGS IN ONCOLOGY: FDA S PERSPECTIVE ASQ Biomed/Biotech SIG Event April 26, 2018 Gaithersburg, MD You Li, Ph.D. Division of Molecular Genetics and Pathology Food and Drug Administration (FDA) Center for
NGS IN ONCOLOGY: FDA S PERSPECTIVE ASQ Biomed/Biotech SIG Event April 26, 2018 Gaithersburg, MD You Li, Ph.D. Division of Molecular Genetics and Pathology Food and Drug Administration (FDA) Center for
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Molecular Heterogeneity of High Gleason Prostate Cancer
 Molecular Heterogeneity of High Gleason Prostate Cancer Aliccia Bollig-Fischer, PhD Assistant Professor Department of Oncology Karmanos Cancer Institute Wayne State University Detroit, Michigan, USA Disclosure
Molecular Heterogeneity of High Gleason Prostate Cancer Aliccia Bollig-Fischer, PhD Assistant Professor Department of Oncology Karmanos Cancer Institute Wayne State University Detroit, Michigan, USA Disclosure
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST
 INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Hematopathology Service Memorial Sloan Kettering Cancer Center, New York
 SH2017-0334 t(14;18) Negative Follicular Lymphoma with 1p36 abnormality associated with In Situ Follicular Neoplasia with t(14;18) translocation Pallavi Khattar MD, Jennifer Maerki MD, Alexander Chan MD,
SH2017-0334 t(14;18) Negative Follicular Lymphoma with 1p36 abnormality associated with In Situ Follicular Neoplasia with t(14;18) translocation Pallavi Khattar MD, Jennifer Maerki MD, Alexander Chan MD,
Deploying the full transcriptome using RNA sequencing. Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven
 Deploying the full transcriptome using RNA sequencing Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven Roadmap Biogazelle the power of RNA reasons to study non-coding RNA
Deploying the full transcriptome using RNA sequencing Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven Roadmap Biogazelle the power of RNA reasons to study non-coding RNA
Children, Toronto, Ontario, Canada. Department of Laboratory Medicine and Pathobiology Hospital for Sick Children, Toronto, Ontario, Canada, M5G 1X8
 Supplementary Information for Clinically Relevant Copy Number Variations Detected In Cerebral Palsy Maryam Oskoui 1, *, Matthew J. Gazzellone 2,3, *, Bhooma Thiruvahindrapuram 2,3, Mehdi Zarrei 2,3, John
Supplementary Information for Clinically Relevant Copy Number Variations Detected In Cerebral Palsy Maryam Oskoui 1, *, Matthew J. Gazzellone 2,3, *, Bhooma Thiruvahindrapuram 2,3, Mehdi Zarrei 2,3, John
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
