Role of TAF4 in Transcriptional Activation by Rta of Epstein-Barr Virus
|
|
|
- Darlene Anthony
- 5 years ago
- Views:
Transcription
1 Role of TAF4 in Transcriptional Activation by Rta of Epstein-Barr Virus Ya-Chun Yang, Li-Kwan Chang* Department of Biochemical Science and Technology, College of Life Science, National Taiwan University, Taipei, Taiwan Abstract Epstein-Barr virus (EBV) expresses an immediate-early protein, Rta, to activate the transcription of EBV lytic genes. This protein usually binds to Rta-response elements or interacts with Sp1 or Zta via a mediator protein, MCAF1, to activate transcription. Rta is also known to interact with TBP and TFIIB to activate transcription. This study finds that Rta interacts with TAF4, a component of TFIID complex, in vitro and in vivo, and on the TATA sequence in the BcLF1 promoter. Rta also interacts with TAF4 and Sp1 on Sp1-binding sequences on TATA-less promoters, including those of BNLF1, BALF5, and the human androgen receptor. These interactions are important to the transcriptional activation of these genes by Rta since introducing TAF4 shrna substantially reduces the ability of Rta to activate these promoters. This investigation reveals how Rta interacts with TFIID to stimulate transcription. Citation: Yang Y-C, Chang L-K (2013) Role of TAF4 in Transcriptional Activation by Rta of Epstein-Barr Virus. PLoS ONE 8(1): e doi: / journal.pone Editor: Renaud Mahieux, Ecole Normale Supérieure de Lyon, France Received June 25, 2012; Accepted December 6, 2012; Published January 10, 2013 Copyright: ß 2013 Yang, Chang. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Funding: This research was supported by the National Health Research Institute of the Republic of China, Taiwan (contract number NHRI-EX BI) and the National Science Council of the Republic of China, Taiwan (contract number NSC B ). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Competing Interests: The authors have declared that no competing interests exist. * changlk@ntu.edu.tw Introduction Although Epstein-Barr virus is normally maintained under latent conditions in B lymphocytes after infection, the virus must enter a lytic cycle to produce infectious virions. In the immediateearly stage of the lytic cycle, EBV expresses Rta and Zta, which are encoded by BRLF1 and BZLF1, respectively, to activate the transcription of lytic genes [1 5]. As is commonly known, Rta binds to a defined sequence of Rta-response elements (RRE) in EBV lytic promoters, including those of BALF5, BMLF1 and BLLF1, to activate transcription [4,6 11]. However, Rta also activates transcription via a mechanism that is independent of RRE binding. For example, Rta interacts with Sp1 via an intermediary protein, MCAF1, to form a complex on Sp1-binding sites that autoregulates its own transcription and enhances the transcription that is regulated by Sp1 [12,13]. Rta also activates the transcription of BALF5 by interacting with USF and E2F instead of binding to an RRE [10]. Moreover, Rta modulates the phosphorylation of the p38, JNK and ERK signaling pathways, causing the binding of phosphorylated ATF2 to the ZII element in the BZLF1 promoter to activate Zta expression [14,15]. Rta also interacts with Oct-1 to enhance the transcription of BZLF1 [16] and with TSG101 to activate the expression of EBV late genes, including BcLF1, BDLF3, BILF2, BLLF1, and BLRF2 [17]. Our earlier study demonstrated that MCAF1 simultaneously interacts with Rta and Zta, allowing Rta and Zta to activate transcription synergistically [18]. Furthermore, Rta is conjugated with SUMO-1 via interaction with SUMO E2 and E3 ligases, including PIAS1, PIASxa, and PIASxb [19,20], increasing the transactivation activity of Rta. Additionally, LF2 regulates viral replication by binding to Rta and altering the subcellular localization of Rta [21]. Transcriptional activation in eukaryotes is a complex process that involves numerous transcriptional and chromatin-remodelling factors. The general transcription factor TFIID, which includes the TATA-binding protein (TBP) and 13 TBP-associated factors (TAFs), recognizes and binds to the TATA box. After the binding, a stepwise assembly of a transcriptional preinitiation complex (PIC) then follows by recruiting other general transcription factors, including TFIIA, TFIIB, TFIIE, TFIIF and TFIIH, and RNA polymerase II [22,23]. TFIID is known to initiate the TATAdependent transcription and also activates a promoter that lacks a TATA box by interacting with TAFs or other transcriptional activators, such as Sp1 [23 26]. Human TAF4 (TAFII130/135), a human homolog of Drosophila TAFII110, is considered a core subunit in TFIID due to its role in stabilizing the holo-tfiid complex [27]. TAF4 is a transcriptional coactivator that contains four glutamine-rich domains that interact with Sp1, CREB and huntingtin [28 30]. However, the preferential interaction of Sp1 with mutant huntingtin interferes with the functions of Sp1 and TAF4, resulting in the inhibition of binding of Sp1 to DNA, which is associated with the pathogenesis of Huntington s disease [30]. Additionally, TAF4 affects the transactivation activities of viral genes by interacting with a number of viral proteins. For instance, the overexpression of TAF4 stimulates the activation of transcription by the small T antigen to promote SV40 DNA replication [31]. TAF4 also interacts with human adenovirus E1A protein to inhibit transcription [32] and promotes the function of STAT2 in Type-I interferon (IFN)- stimulated transcription by forming a non-classical TBP-free acetyltransferase complex, which prevents the transcriptional shutoff of host genes during poliovirus infection [33]. TAF4 also activates the transcription from a TATA-less promoter that PLOS ONE 1 January 2013 Volume 8 Issue 1 e54075
2 contains a downstream core promoter element (DPE) [27]. The TATA-less promoters are involved in the transcription of many cellular and viral genes, including those that code for dihydrofolate reductase (DHFR), c-h-ras, adenosine deaminase, TGF-b, thymidylatesynthetase, human androgen receptor (har), SV-40 late gene products, and LMP1 and DNA polymerase (BALF5) of EBV [10,34 37]. These TATA-less promoters typically contain Sp1-binding sequences. Through the interaction between Sp1 and TAF4, other transcription factors and RNA polymerase II are recruited to initiate transcription from a promoter that lacks a TATA sequence [28,35]. Rta is a potent transcriptional factor that activates many viral and cellular genes through interaction with both TBP and TFIIB [38]. This study demonstrates that Rta enhances transcription via the interaction with TAF4. Methods Cell lines and EBV lytic induction P3HR1 (ATCC HTB-62), a Burkitt s lymphoma cell line that contains EBV [39], was cultured in RPMI 1640 medium containing 10% fetal calf serum. 293T cells (ATCC CRL-11268) were cultured in Dulbecco s modified Eagle s medium (DMEM) containing 10% fetal calf serum. P3HR1 cells were treated with 30 ng/ml 12-O-tetradecanoylphobol-13-acetate (TPA) and 3 mm sodium butyrate to activate the EBV lytic cycle [40,41]. Plasmids Plasmid pgex-4t1, which expresses GST, was purchased from Amersham Biosciences. Plasmid pbsk-taf4, which contains a TAF4 gene, was provided by Naoko Tanese [29]. Plasmid pgex- TAF4, which encodes GST-TAF4, was constructed by inserting a 3.9-kb EcoRI fragment, which contains the TAF4 gene, from pbsk-taf4 into the EcoRI site in pgex-4t3 (Amersham Biosciences) to generate pgex-taf4. Plasmid pegfp-taf4, which expresses TAF4 that is fused to GFP, was constructed by digesting pgex-taf4 with EcoRI and XhoI and then inserting the TAF4 DNA fragment into the EcoRI-SalI sites in pegfp-c1 (Clontech). Plasmid pegfp-taf4-nm, which encodes the region in TAF4 from amino acid 1 to 701, was constructed by digesting pegfp-taf4 with ClaI and XhoI, eluting the digested fragment and treating with the DNA polymerase I Klenow fragment, and then inserted into pegfp-c1. Plasmid pegfp-taf4-c, which expresses the C-terminal 246-amino acid region in TAF4 that is fused to GFP, was constructed by inserting a PCR-amplified TAF4 DNA fragment into the EcoRI-SalI sites in pegfp-c1. Plasmid pegfp-rta was provided by T. Y. Hsu [42]. Plasmids pcmv-r, pet-rta, pbmrf1-luc and pbmrf1-mrre were described elsewhere [18,19]. Plasmids that express deletion mutants of GFP- Rta, including GFP-N190, GFP-N191/415 and GFP-Rev, which encodes Rta regions from amino acids 1 190, and , respectively, were constructed by inserting a PCR-amplified BRLF1 fragment into the EcoRI-XhoI sites in pegfp-c1. Plasmids ptrl1-luc, pmtrl1-luc, pbalf5-luc and phar-luc were constructed by inserting PCR fragments that had been amplified using primer sets TRL1-H3 (59-CCCAAGCTTGAATGAG- GTGGCGGATT)/TRL1-X (59-CCGCTCGAGCCGCGGGG- GAAGGCCAC), TRL1-H3 (59-CCCAAGCTTGAATGAGGT- GGCGGATT)/mTRL1-X (59-CCGCTCGAGCCGCGGGG- GAAGGACCAGCTGCCTCC), BALF5(+170) (59-AGATGTC- GCAGCAGCCG)/BALF5(+16)(59-CCCAAGCTTCCACTGA- GGGTCCGGCC), and har(+140) (59-CCGCTCGAGTAG- CGCGCGGTGAGGGGA)/hAR(+57) (59-CCCAAGCTTCTC- TGGGCTTGCTCCGGA), respectively, into pgl2-basic (Promega) at the HindIII-XhoI sites. Plasmid pmbalf5-luc contains the same sequences as pbalf5-luc, but with the Sp1-binding sequence mutated from 59-CCGCCC to 59-AAGCCC. Plasmid pmhar-luc has the same sequences as is present in phar-luc, but 59-GGGGCGGG was changed to 59-GTGGGTTG. Plasmids pgl2-f23 was constructed by inserting a 23-bp double-strand DNA fragment that contains the region from nucleotide -38 to -16 in the BcLF1 promoter [43] into pgl2-basic. GST pull-down assay GST, GST-TAF4, and His-Rta were purified from E. coli BL21(DE3) using glutathione-sepharose 4B beads (Amersham Biosciences) according to a method described earlier [19]. Immunoprecipitation P3HR1 cells were treated or untreated with sodium butyrate and TPA for 24 hr to induce lytic cycle. A lysate was prepared using mripa buffer (50 mm Tris-HCl, ph 7.8, 150 mm NaCl, 5 mm EDTA, 0.5% Triton X-100, 0.5% Nonidet P-40) [13]. Monoclonal mouse anti-rta antibody (1:500 dilution) (Argene), or mouse anti-taf4 antibody (1:1000) (BD Biosciences) was added to the supernatant and incubated at 4uC for 1 hr. The normal mouse IgG (Santa Cruz Biotechnology) was used as a negative control. Protein G agarose beads (30 ml) (Oncogene) were then added to the lysate. After shaking for 1 hr at 4uC, the beads were collected by centrifugation and washed three times with mripa buffer. Proteins binding to the beads were eluted by adding 20 ml 2X electrophoresis sample buffer and analyzed by immunoblotting using anti-rta and anti-taf4 antibodies. Immunofluorescence analysis P3HR1 cells were transfected with pegfp-c1 or pegfp-taf4 and then treated with TPA and sodium butyrate to activate the EBV lytic cycle. After culturing for 24 hr, cells were harvested by centrifugation, plated on poly-l-lysine (Sigma)-coated coverslips, and fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 30 min. Immunostaining was performed using anti-rta monoclonal antibody. Cells were then treated with rhodamineconjugated goat anti-mouse IgG polyclonal antibody (DAKO). Nuclei were visualized by staining using 5 mg/ml of diamidino-2-phenylindole (DAPI). Finally, cells were examined under a Zeiss confocal laser-scanning microscope (Model LSM780). DNA affinity precipitation assay (DAPA) P3HR1 cells that had been treated with TPA and sodium butyrate were lysed in mripa buffer. The lysate was mixed with 300 ng 59-biotin end-labeled double-stranded probe, F23, mf23, TR-L1, mtr-l1, har or mhar in a binding buffer that contained 60 mm KCl, 12 mm HEPES, ph 7.9, 4 mm Tris- HCl, 5% glycerol, 0.5 mm EDTA, 1 mm DTT and 10 mg/ml each of leupeptin, aprotinin, and 4-(2-aminoethyl)-benzenesulfonyl fluoride. After it had been incubated under rotation for 1 hr at 4uC, DNA-protein complexes were captured using 100 ml Streptavidin MagneSphere Paramagnetic particles (Promega), incubated for 1 hr, and then washed five times in the binding buffer. Finally, precipitated DNA-protein complexes were mixed in 2X electrophoresis sample buffer and proteins were detected by immunoblotting. Probe F23 contains the sequence that covers the region from -38 to -16 in the BcLF1 promoter (59-GAATTAT- TAAACCGGGTGGCAGC). Probe mf23 includes the same sequence as probe F23 except that the 59-TATTAAA were changed to 59-TGTTGAA. TR-L1 contains the sequence between nucleotide and in the EBV genome (GenBank PLOS ONE 2 January 2013 Volume 8 Issue 1 e54075
3 accession number V01555) (59-CCGCGGGGGAAGGCCA- CGCCCCCTCCACTTTTTC). Probe mtr-l1, which contains the same sequence as in TR-L1 except for a mutated Sp1 site (59- CCGCGGGGGAAGGACCAGCTGCCTCCACTTTTTC), was used as a negative control. Probe har contained the nucleotide sequence from -32 to -63 in the human androgen receptor promoter (59-AGGAGGCCGGCCCGGTGGGGGCGGGAC- CCGAC) [44], and probe mhar contains a similar sequence as har except for the Sp1 site was mutated (59-AGGAGG- CCGGCCCGGTGGGTTCGGGACCCGAC). Chromatin immunoprecipitation (ChIP) assay ChIP assay was conducted with P3HR1 cells as described previously [45]. Briefly, P3HR1 cells ( ) were treated with TPA and sodium butyrate to induce the EBV lytic cycle or treated with DMSO as a control. Cells were mixed with 1% formaldehyde and incubated under shaking for 10 min to crosslink proteins and DNA. After sonication, crosslinked DNA-protein complexes were immunoprecipitated using anti-sp1 (Santa Cruz Biotechnology), anti-taf4, or anti-rta antibody. The presence of specific DNA fragments in the precipitates was detected by qpcr as described elsewhere [18]. Standard curves were generated using serial dilutions of input DNA (1000, 100, 10, 1 and 0.1%). The Ct value of each reaction was quantified against the standard curve. Primers used for amplifying the TR-L1 promoter were 59-AG- AATGAGGTGGCGGATTCA and 59-CGCCCCCTCCACTT- TTTC; the BALF5, 59-CGTGAGGGAAATAACCAGGATC and 59-TGGTTTCATCTCCTTCTTCAAAAAC; the har promoter, 59-GGAGGTGGGAAGGCAAGGA and 59-TGGAGAG- CAAATGCAACAGTTT; the BcLF1 promoter, 59-GCAGAG- GACCATATTTCGCAGTCTG and 59-GCTGCCACCCGGT- TTAATAATTC. Transient transfection assay Plasmids were purified from E. coli using a plasmid purification kit (Qiagen). P3HR1 cells ( ) were mixed with 5 mg plasmids in 300 ml cultured medium and transfected by electroporation at 975 mf and 0.2 V using a BTX ECM630 electroporator (BTX Instrument). 293T cells were transfected using TurboFect (Fermentas). Luciferase assay was monitored using a luminometer (Orion II; Berthod) as described elsewhere [46]. Each transfection experiment was performed at least three times, and each sample in the experiment was prepared in duplicate. RNAi The shrna of TAF4 was purchased from the National RNAi Core Facility, Genomic Research Center, Academia Sinica. Two plasmids (300 ng) that expressed TAF4 shrna (clone ID: TRCN and TRCN ) were transfected into 293T cells using TurboFect (Fermentas). Inhibition of TAF4 expression was examined at 48 hr after transfection. Results Interaction between Rta and TAF4 in vitro Since both Rta and TAF4 interact with Sp1 [28,30,35], this study investigated whether Rta interacts with TAF4 to promote transcription. GST-pulldown assay was performed using bacterially expressed GST-TAF4 fusion protein (Fig. 1, lane 8) to investigate the interaction between Rta and TAF4. Assay results revealed that Rta expressed in the P3HR1 cells after lytic induction was retained by GST-TAF4 that was bound to glutathione-sepharose beads but was not pulled down by GSTglutathione-Sepharose beads (Fig. 1, lanes 1 3). A similar assay was carried out by adding the E. coli BL21(DE3)(pET-Rta) lysate to GST- or GST-TAF4-glutathione-Sepharose beads. The results indicated that bacterially expressed His-Rta was retained by GST- TAF4-glutathione-Sepharose but not by GST-glutathione-Sepharose beads (Fig. 1, lanes 4 6), confirming the direct interaction between Rta and TAF4. Interaction between Rta and TAF4 in vivo Immunoprecipitation assay was conducted using the lysate from P3HR1 cells that had been treated with TPA and sodium butyrate for 24 hr to verify the interaction of Rta with TAF4 in vivo. The results reveal that Rta in the cell lysate was immunoprecipitated by anti-rta antibody (Fig. 2, lane 4) and coimmunoprecipitated with TAF4 by anti-taf4 antibody (Fig. 2, lane 3). TAF4 was also immunoprecipitated by anti-taf4 antibody (Fig. 2, lane 12) and coimmunoprecipitated with Rta by anti-rta antibody (Fig. 2, lane 11). Meanwhile, in a study that involved the lysate prepared from P3HR1 cells that had not been treated with TPA and sodium butyrate, immunoblotting revealed that Rta was not present in the lysate (Fig. 2, lane 5) and could not be immunoprecipitated by anti-rta or anti-igg antibody (Fig. 2, lanes 6, 8). Immunoblotting also revealed that TAF4 was present in the lysate (Fig. 2, lane 13) and was not immunoprecipitated by anti-igg (Fig. 2, lane 14). Moreover, although anti-taf4 antibody immunoprecipitated TAF4 in the lysate (Fig. 2, lane 16), did not coimmunoprecipitate Rta (Fig. 2, lane 7). Notably, a small amount of TAF4 was coimmunoprecipitated using anti-rta antibody (Fig. 2, lane 15), perhaps because of the presence of a low level of Rta in the lysate due to spontaneous EBV lytic activation. Moreover, the specificity of anti-rta antibody is significantly higher than that of anti-taf4 antibody. Therefore, the amount of latent lysate loaded to the gel for the detection of Rta (Fig. 2. lanes 1 8) was substantially less than that for the detection of TAF4 (Fig. 2, lanes 9 16), explaining why an Rta band was undetected in the experiment (Fig. 2, lanes 5 8). Furthermore, Rta in the lysates that were prepared from cells that were treated with TPA and sodium butyrate was coimmunoprecipitated by anti-taf4 antibody (Fig. 2, lane 3). Yet, we were unable to use the anti-taf4 antibody to coimmunoprecipitate Rta in the lysates that were prepared from cells not infected by EBV (Fig. 2, lane 7), showing that anti-rta antibody does not react to TAF4 nonspecifically. Colocalization of Rta and TAF4 in the nucleus A confocal microscopic work was performed to confirm the interaction between Rta and TAF4 in P3HR1 cells that were transfected with pegfp-taf4 and had been treated with sodium butyrate and TPA for 24 hr. The result showed that Rta colocalized with GFP-TAF4 as dots in the nucleus (Fig. 3H). Meanwhile, the cells transfected with an empty vector had GFP present diffusely in the entire cells and did not colocalize with Rta (Fig. 3C and 3D). Mapping the interaction domains in Rta and TAF4 To delineate the regions in TAF4 that interact with Rta, 293T cells were cotransfected with pcmv-r and plasmids that encode GFP-TAF4, GFP-TAF4-NM, and GFP-TAF4-C (Fig. 4A). An empty vector, pegfp-c1, that expressed GFP was used as a negative control. Immunoblotting using an anti-gfp antibody revealed the presence of GFP-TAF4 and GFP-TAF4-C in the lysate (Fig. 4B, lanes 2, 4). Immunoblotting using anti-rta antibody also revealed that Rta was coimmunoprecipitated with GFP-TAF4 and GFP-TAF4-C by anti-gfp antibody (Fig. 4B, lanes 7, 9). A parallel experiment revealed that although GFP and GFP-TAF4-NM were present in the lysate (Fig. 4B, lanes 1, 3), PLOS ONE 3 January 2013 Volume 8 Issue 1 e54075
4 Figure 1. Interaction between Rta and TAF4 in vitro. GST-TAF4 (lanes 3 and 6) or GST (lanes 2, 5) was added to the lysate prepared from P3HR1 cells that had been treated with TPA and sodium butyrate (lanes 1 3) or from E. coli BL21(DE3)(pET-Rta) (lanes 4 6). Proteins bound to GST-TAF4 were pulled down by glutathione-sepharose beads and analyzed by immunoblotting (IB) with anti-rta antibody. Lanes 1 and 4 were loaded with 5% of cell lysate. GST or GST-TAF4 bound to glutathione-sepharose beads were analyzed by immunoblotting (IB) with anti-gst antibody (lanes 7, 8). doi: /journal.pone g001 Rta was not coimmunoprecipitated with these proteins by anti- GFP antibody (Fig. 4B, lanes 6, 8). These results suggested that the region between amino acids 702 and 947 in TAF4 interacts with Rta. Additionally, deletions were generated in GFP-Rta (Fig. 4C) to identify the regions in Rta that interacted with TAF4. These GFP-Rta fusion proteins were detected in the lysate by immunoblotting using the anti-gfp antibody after transfection (Fig. 4D, lanes 1 5). Immunoblot analysis revealed that only GFP-Rta and GFP-N were coimmunoprecipitated with TAF4 by anti- TAF4 antibody (Fig. 4D, lanes 7, 9), indicating that the region between amino acids 191 and 415 in Rta interacts with TAF4. Activation of BcLF1 transcription by interaction of Rta with TAF4 Rta is known to activate a number of EBV late genes, including BcLF1 [17]. Previous studies have found that the BcLF1 gene is transcribed from a promoter that is 23 bp long, which includes a TATA box [43,47]. Hence, this study investigates whether Rta interacts with TAF4 to activate the transcription of BcLF1. Accordingly, DAPA was conducted using a biotin-labeled F23 DNA probe that contained the 23-bp sequence, from -38 to -16, in the BcLF1 promoter (Fig. 5A). The probe was added to a lysate that was prepared from P3HR1 cells that had been treated with sodium butyrate and TPA. Proteins that were bound to the probe were captured using streptavidin magnetic beads. Immunoblot analysis revealed the binding of TAF4 and Rta to the F23 probe. However, TAF4 and Rta did not bind to a mutant probe, mf23, which contains a mutated TATA sequence (Fig. 5A), indicating the binding of Rta and TAF4 to the TATA sequence in the BcLF1 promoter. Furthermore, ChIP analysis was performed to confirm the binding of TAF4 and Rta to the TATA box in the BcLF1 promoter in vivo. After P3HR1 cells were treated with sodium butyrate and TPA for 52 hr to induce the expression of EBV late genes, proteins and DNA in the cells were crosslinked with formaldehyde. qpcr with primers that were specific for the BcLF1 promoter sequence was subsequently performed. Both TAF4 and Rta were found to be bound to the BcLF1 promoter (Fig. 5B). The amounts of the BcLF1 promoter captured by anti- TAF4 and anti-rta antibodies was 3.6 and 7.7 times, respectively, the amounts that were immunoprecipitated by anti-igg antibody (Fig. 5B). Furthermore, only a background level of BcLF1 promoter DNA was immunoprecipitated by anti-igg, anti- TAF4, and anti-rta antibodies when P3HR1 cells were not Figure 2. Coimmunoprecipitation of Rta and TAF4. Interaction between TAF4 and Rta was examined using P3HR1 cells that were treated with DMSO (lanes 5 8; 13 16) or TPA and sodium butyrate (lanes 1 4; 9 12) for 24 hr. Proteins in the cell lysate were immunoprecipitated with anti-rta (lanes 4, 8, 11, 15), anti-taf4 (lanes 3, 7, 12, 16) or anti-igg (lanes 2, 6, 10, 14) antibody. Immunoprecipitated proteins were detected by immunoblotting with anti-rta (lanes 1 8) or anti-taf4 (lanes 9 16) antibody. Lanes 1, 5, 9 and 13 were loaded with 5% of the cell lysate. doi: /journal.pone g002 PLOS ONE 4 January 2013 Volume 8 Issue 1 e54075
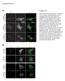
5 Figure 3. Indirect immunofluorescence analysis. P3HR1 cells were transfected with pegfp-c1 (A D) or pegfp-taf4 (E H) and then treated with sodium butyrate for 24 hr. Cells were incubated with monoclonal anti-rta antibody and observed under a confocal laser-scanning microscope. DAPI staining revealed the positions of nuclei (A and E). D and H are merged images. doi: /journal.pone g003 treated with sodium butyrate and TPA (Fig. 5B). These results indicate the binding of Rta and TAF4 to the BcLF1 promoter in vivo. Transient transfection assay was also carried out in 293T cells that were transfected with pgl2-f23, which contains the region from nucleotides -38 to -16 in the BcLF1 promoter to investigate whether TAF4 is involved in Rta-mediated BcLF1 transcription. After transfecting 293T cells with pcmv-r, the activity of the BcLF1 promoter increased by tenfold (Fig. 5C). Meanwhile, transfecting two plasmids that express the shrnas of TAF4 decreased the expression of TAF4 in the cells (Fig. 5D) and reduced by 70% the activity of the BcLF1 promoter that was activated by Rta (Fig. 5C). The expression of shrna did not influence the luciferase activity of pgl2-basic that was nonspecifically activated by Rta (Fig. 5C) [13,48]. These results suggested that the activation of BcLF1 transcription by Rta requires TAF4. Involvement of the Sp1-TAF4-Rta complex to Sp1- binding sites in TATA-less promoters of EBV This study investigates whether the transcriptional activation of the TR-L1 promoter of the EBV BNLF1 gene, which lacks a TATA box, by Rta [49] involves TAF4. A biotin-labeled TR-L1 DNA probe, which contains the region from -32 to -47 in the TR- L1 promoter (Fig. 6A), was added to a lysate that was prepared from P3HR1 cells that had been treated with sodium butyrate and TPA. Proteins that bound to the probe were then captured using streptavidin magnetic beads. Immunoblot analysis revealed the binding of Sp1, TAF4 and Rta to the probe (Fig. 6A). These proteins did not bind to a mutant probe, mtr-l1, whose sequence is identical to that of TR-L1, except for a mutated Sp1- binding sequence (Fig. 6A). After the binding of Sp1, TAF4, and Rta to the Sp1-binding site in the promoter in vitro was demonstrated, the binding of the complex to the Sp1 site in vivo was studied by ChIP analysis. Accordingly, P3HR1 cells were treated with sodium butyrate and TPA to induce EBV lytic activation and cells that were treated with DMSO were used as a control. Following the immunoprecipitation with anti-sp1, anti- TAF4 and anti-rta antibodies, qpcr analysis using primers that amplified the Sp1-binding sequences in the TR-L1 promoter revealed that these three antibodies immunoprecipitated the DNA fragments that contained the TR-L1 promoter. The amounts of promoter sequences that were captured by anti-sp1, anti-taf4 and anti-rta antibodies were 12.9, 3.4, and three times higher than that immunoprecipitated by anti-igg antibody (Fig. 6B). Additionally, when cells were treated with DMSO rather than TPA and sodium butyrate, only the background amount of TR-L1 promoter was amplified by qpcr (Fig. 6B). A similar result was obtained using the BALF5 promoter of EBV, which also lacks a TATA sequence, in P3HR1 cells that had been treated with sodium butyrate and TPA for 48 hr to induce the expression of Rta. qpcr analysis using primers that were specific for the Sp1- binding sites in the BALF5 promoter demonstrated that these three antibodies immunoprecipitated the BALF5 promoter sequences in amounts that were 6.5, 3.3, and 3.6 times that obtained using the anti-igg antibody, respectively (Fig. 6B). Meanwhile, background levels of the DNA fragment were captured by these antibodies when the cells were untreated with TPA and sodium butyrate (Fig. 6B). shrna that inhibits the expression of TAF4, was also used to demonstrate the participation of Rta and TAF4 in transcriptional activation of these two TATA-less promoters. Immunoblotting verified that TAF4 expression declined to an undetectable level after 293T cells were transfected with two plasmids that expressed shrnas of TAF4 (Fig. 6E). Thereafter, 293T cells were cotransfected with ptrl1- luc and pcmv-r in the presence of TAF4 shrna or control shrna. The results showed that introducing TAF4 shrna reduced the transcriptional activity of the TR-L1 promoter that was activated by Rta from 32-fold to tenfold (Fig. 6C). This study also used pmtrl1-luc, which contains a luc gene transcribed from a mutant TR-L1 promoter that lacks the Sp1 site. Despite of the mutation, Rta activated the promoter 18-fold; the expression of TAF4 shrnas further reduced the promoter activity to tenfold (Fig. 6C), revealing the involvement of TAF4 in Rta-mediated TRL1 transcription. A parallel experiment demonstrated that overexpressing Rta increased the luciferase activity of pbalf5-luc by a factor of 12 (Fig. 6C), but only by a factor of six after shrna had been expressed (Fig. 6C). When the reporter plasmid that contained a mutated Sp1 binding sites, pmbalf5-luc, was utilized, Rta increased the promoter activity by a factor of 5.6 (Fig. 6C). Silencing of TAF4 expression did not influence the activity of the mutant BALF5 promoter (Fig. 6C). These results suggest that TAF4 is important to the transcriptional activation of the TR-L1 and BALF5 promoters by Rta. PLOS ONE 5 January 2013 Volume 8 Issue 1 e54075
6 Figure 4. Mapping the interaction domains in TAF4 and Rta. (A) Plasmids that expressed deleted GFP-TAF4 were used to delineate the region in TAF4 that interacts with Rta. Numbers represent the positions of amino acids in TAF4. Q denotes the glutamine-rich regions. (B) 293T cells were cotransfected with pcmv-r and plasmids that expressed GFP fusion proteins including pegfp-taf4 (lanes 2, 7), pegfp-taf4-nm (lanes 3, 8), pegfp- TAF4-C (lanes 4 and 9) or pegfp-c1 (lanes 1, 6). Input lanes were loaded with 5% of the lysate (lanes 1 5). Proteins in the lysates were coimmunoprecipitated (IP) with anti-gfp antibody and analyzed by immunoblotting (IB) using anti-rta antibody (lanes 6 9). (C) Deletion mutants of PLOS ONE 6 January 2013 Volume 8 Issue 1 e54075
7 Rta were used to identify the region in Rta that interacts with TAF4. Numbers represent the positions of amino acids in Rta (D). Plasmids that expressed GFP-Rta (lanes 2, 7), GFP-N190 (lanes 3, 8), GFP-N (lanes 4, 9), GFP-Rev (lanes 5, 10) or GFP (lanes 1, 6) were transfected into 293T cells. The input lanes were loaded with 5% of the cell lysates and GFP-fusion proteins were detected using anti-gfp antibody (lanes 1 5). Proteins in the lysates were coimmunoprecipitated with anti-taf4 antibody and analyzed by immunoblotting using anti-gfp antibody (lanes 6 10). doi: /journal.pone g004 Rta is known to bind directly to RRE to affect transcription [4,6 11]. A transient transfection study was performed using the promoter pbmrf1-luc that contains an RRE [18] to investigate whether TAF4 also participates in RRE-dependent transcription. Accordingly, luciferase activity was monitored by cotransfecting 293T cells with pbmrf1-luc and pcmv-r. As expected, Rta activated the promoter activity of pbmrf1 23-fold (Fig. 6D). The increase in promoter activity that was exhibited by pbmrf1-luc was decreased to 12-fold after the expression of TAF4 shrna (Fig. 6D). When 293T cells were cotransfected with a reporter plasmid that contained a mutated RRE, pbmrf1-mrre, and pcmv-r, the luciferase activity of pbmrf1-mrre was eight Figure 5. Involvement of TAF4 in the activation of the BcLF1 promoter by Rta. (A) Biotin-labeled double-stranded F23 probes were added to a lysate prepared from P3HR1 cells that had been treated with TPA and sodium butyrate for 52 hr. Mutant probes mf23 that contains mutated sequences (asterisk) was used as negative controls. Proteins bound to the probes were captured by the streptavidin magnetic beads and detected by immunoblotting analysis using anti-taf4 and anti-rta antibodies. Input lanes were loaded with 5% of the cell lysate. DAPA: DNA affinity precipitation assay. (B) P3HR1 cells were treated with TPA and sodium butyrate for 52 hr. Formaldehyde-fixed DNA-protein complex was immunoprecipitated using anti-taf4 or anti-rta antibody. The reaction with added anti-igg antibody was used as a negative control. The binding of TAF4 and Rta to the BcLF1 promoter was investigated by qpcr. Error bar represents standard error. (C) 293T cells were cotransfected with pcmv-r and pgl2-f23 or a control vector pgl2-basic in the presence of control shrna (Ct-shRNA) (filled column) or TAF4 shrna (shtaf4) (empty column). Luciferase activity was detected at 48 hr after transfection. Each transfection experiment was performed three times, and each sample in the experiment was prepared in duplicate. (D) The effect of TAF4 shrnas was examined by immunoblotting with anti-taf4 and anti-a-tubulin antibodies. The p value from each experiment was analyzed statistically with the Student s t- test method. doi: /journal.pone g005 PLOS ONE 7 January 2013 Volume 8 Issue 1 e54075
8 Figure 6. Influence of TAF4 on the activation of EBV TATA-less promoters by Rta. (A) The TR-L1 reporter plasmid and sequences of biotinlabeled probes, TR-L1 and mtr-l1. The probes were added to a lysate prepared from P3HR1 cells that had been treated with TPA and sodium butyrate. Proteins bound to the probes were captured by streptavidin magnetic beads and detected by immunoblotting analysis using anti-sp1, anti- TAF4 and anti-rta antibodies. Input lanes were loaded with 5% of the cell lysate. DAPA: DNA-affinity precipitation assay. (B) P3HR1 cells that had been PLOS ONE 8 January 2013 Volume 8 Issue 1 e54075
9 treated with TPA and sodium butyrate for 48 hr (lytic) or treated with DMSO (latent) were fixed with formaldehyde and the DNA-protein complexes were immunoprecipitated using anti-sp1, anti-taf4 and anti-rta antibodies. The binding of Sp1, TAF4 and Rta to the TR-L1 and BALF5 promoters was investigated by qpcr. Error bar represents standard error. (C) 293T cells were cotransfected with pcmv-r and reporter plasmids, including ptrl1-luc and pbalf5-luc in the presence of control shrna (Ct-shRNA) (filled column) or TAF4 shrna (empty column). Luciferase activities were monitored at 48 hr after transfection. Each transfection experiment was performed at lease three times, and each sample in the experiment was prepared in duplicate. (D) A similar experiment in (C) was performed using pbmrf1-luc and pbmrf1-mrre-luc. (E) The effect of TAF4 shrnas on the expression of TAF4 was examined by immunoblotting using anti-taf4 and anti-a-tubulin antibodies. The p value from each experiment was analyzed statistically with the Student s t- test method. Luc: luciferase gene. doi: /journal.pone g006 times that of the negative control (Fig. 6D). Knockdown of TAF4 expression by shrna reduced the increase in activity of the promoter to threefold (Fig. 6D), showing that Rta-TAF4 complex does not need RRE to activate transcription. Function of TAF4 in the transcriptional activation of human androgen receptor (har) by Rta In this study, the promoter from human androgen receptor gene, which lacks a TATA box [44], was used to determine whether TAF4 is involved in Rta-activated transcription. The DAPA results indicated that Sp1, TAF4 and Rta were bound to the probe, but did not bind to the mutant probe that contained a mutated Sp1 site (Fig. 7A). ChIP assay revealed that when P3HR1 cells were treated with TPA and sodium butyrate, the amounts of Sp1, TAF4, and Rta bound to the har promoter were 4.2-, 3.7-, and 2.9-fold higher than those captured by anti-igg antibody (Fig. 7B). When cells were not treated with TPA and sodium butyrate, these antibodies captured only background amount of the promoter DNA (Fig. 7B). This study introduces TAF4 shrna into 293T cells to examine how TAF4 influences the transcription of the androgen receptor gene that is activated by Rta. Accordingly, ChIP assay was performed using 293T cells that were cotransfected with pcmv-r and TAF4 shrna or control shrna for 48 hr. qpcr analysis using primers to amplify the har promoter indicated that overexpressing Rta increased the amounts of Sp1, TAF4, and Rta bound to the promoter by factors of 3.2, 4, and 3.5, respectively (Fig. 7C). However, introducing TAF4 shrna reduced the levels of TAF4 and Rta that bound to the promoter from 4-fold to 3.3-fold and from 3.5-fold to 2.2-fold, respectively (Fig. 7C), but did not influence the binding of Sp1 to the promoter (Fig. 7C). A transient transfection experiment was conducted using a reporter plasmid phar-luc. Overexpressing Rta increased the luciferase activity of phar-luc 53-fold (Fig. 7D), but only 21-fold following the expression of the TAF4 shrna (Fig. 7D), The increase in Rta-activated activity of a mutant har promoter in pmhar-luc, which contains a mutated Sp1 site, declined from 13-fold to sixfold when TAF4 shrna was cotransfected (Fig. 7D). Discussion Rta is a transcription factor that is expressed by EBV in the immediate-early stage of the lytic cycle, to activate the transcription of EBV lytic genes by binding to RRE in promoters [11] or by forming a complex with Sp1 or Zta by interacting with MCAF1 [13,18]. Rta also activates TATA-less and GC-rich promoters of EBV, including BNLF1 and BALF5 [10,49]. Furthermore, Rta interacts with basal transcription factors, TBP and TFIIB, to activate transcription [38]. This study demonstrates that Rta interacts with a core component of TFIID, TAF4, to activate both TATA and TATA-less transcription, revealing how Rta interacts with the eukaryotic transcription machinery to activate transcription. Several lines of evidence indicate that TFIID binding to promoters is a rate-limiting step in transcription [22,23,50] and that transcription factors such as Sp1 or CREB recruits and stabilizes TFIID at the TATA sequence [28,51 53]. This study demonstrates that Rta interacts with TAF4 by GST-pulldown assay (Fig. 1), coimmunoprecipitation (Fig. 2), and confocal microscopy (Fig. 3H). The result shows that a small amount of TAF4 that was coimmunoprecipitated by anti-rta antibody (Fig. 2, lane 15). Because P3HR1 is known to express Rta in a cell population that is untreated with TPA and sodium butyrate due to spontaneous EBV lytic activation, coimmunoprecipitation of TAF4 by anti-rta antibody is likely attributed to the leaky expression of Rta. Additionally, the interaction involves the C- terminal domain in TAF4 and the region between amino acids 191 and 415 in Rta (Fig. 4). Previous studies have shown that the C-terminal region ( a.a.) of TAF4, which is conserved from yeast to humans, stabilizes and nucleates the holo-tfiid complex [27,54], suggesting that Rta promotes and stabilizes the assembly of the TFIID complex by interacting with the C-terminal region in TAF4. This study demonstrates that Rta cooperates with TAF4 to activate promoters that contain a TATA sequence. A previous study demonstrated that the activation by Rta of the transcription of BcLF1 through a 23-bp region, from nucleotide -38 to -16, which contains a TATA box, is sufficient for the promoter activity [43,47]. Hence, Rta probably interacts with basal transcription factors that bind to the TATA box, influencing the BcLF1 transcription. The DAPA and ChIP assay herein reveal that Rta and TAF4 form a complex on the BcLF1 promoter upon lytic induction (Fig. 5A and 5B). Transient transfection analysis also shows that introducing TAF4 shrna reduces the ability of Rta to activate the transcription of BcLF1 by 70% (Fig. 5C), indicating that Rta activates the transcription of BcLF1 by interacting with TAF4. However, this study cannot exclude the possible involvement of other EBV-encoded protein, such as BcRF1, which is a TBP-like protein that forms a complex on TATA motifs in the activation of late genes [55]. This study shows that the interaction of Rta and TAF4 activates the EBV promoters that lack a TATA sequence. Earlier studies have established that Rta transcriptionally activates two EBV promoters without a TATA sequence, the TR-L1 promoter of BNLF1 and the promoter of BALF5, after lytic activation [49,56]. Rta is demonstrated to interact with TAF4 and Sp1 and form a complex on the promoters of TR-L1 and BALF5 after EBV lytic induction (Fig. 6B). Additionally, TAF4 shrna substantially reduces the ability of Rta to activate these two promoters (Fig. 6C), revealing that the activation of EBV TATA-less promoters by Rta depends on the formation of the Sp1-TAF4-Rta complex and subsequent binding of this complex to the Sp1 site. Rta activates the TR-L1 promoter at a reduced level after the Sp1 site in the promoter is mutated (Fig. 6C), which may be attributed to the fact that the TR-L1 promoter contains high GC-rich sequences, such as the region from -3 to -14 [37,57], where Sp1 may also bind, allowing Rta to activate transcription through this sequence. This study shows that Rta considerably activates the har promoter, which has no TATA sequence (Fig. 7D). Mutation of the Sp1 sequence in the promoter reduces the luciferase activity PLOS ONE 9 January 2013 Volume 8 Issue 1 e54075
10 Figure 7. Role of TAF4 in the transcription of human androgen receptor by Rta. (A) An har reporter plasmid and sequences of biotinlabeled probes, har and mhar. The probes were added to a lysate prepared from P3HR1 cells that had been treated with TPA and sodium butyrate. Proteins bound to the probes were captured by streptavidin magnetic beads and detected by immunoblot analysis using anti-sp1, anti-taf4 and anti-rta antibodies. Input lanes were loaded with 5% of the cell lysate. DAPA: DNA-affinity precipitation assay. (B) P3HR1 cells that had been treated with TPA and sodium butyrate for 48 hr (lytic) or treated with DMSO (latent) were fixed with formaldehyde and the DNA-protein complexes were immunoprecipitated with anti-sp1, anti-taf4 and anti-rta antibodies. The binding of Sp1, TAF4 and Rta to the har promoter was investigated by qpcr. (C) 293T cells were cotransfected with pcmv-r and TAF4 shrna (empty column) or control shrna (filled column) for 48 hr. CHIP assay was subsequently performed. The reaction with added anti-igg antibody was used as a negative control. Error bar represents standard error. (D) 293T cells were cotransfected with pcmv-r and a reporter plasmid including phar-luc or pmhar-luc, in the presence of control shrna (Ct-shRNA) (filled column) or TAF4 shrna (empty column). Luciferase activities were monitored at 48 hr after transfection. Each transfection experiment was performed at lease three times, and each sample in the experiment was prepared in duplicate. The p value from each experiment was analyzed statistically with the Student s t- test method. Luc: luciferase gene. doi: /journal.pone g007 PLOS ONE 10 January 2013 Volume 8 Issue 1 e54075
11 that was activated by Rta (Fig. 7D), implying that Rta activates the har promoter via its Sp1-binding sequence. The DAPA and ChIP analysis reveal that Rta, TAF4, and Sp1 form a complex on the har promoter after lytic induction (Fig. 7A, 7B). Additionally, ChIP analysis revealed that the knockdown of TAF4 expression reduces the amount of Rta but not the amount of Sp1 on the promoter (Fig. 7C), indicating that Rta is recruited by TAF4 and then interacts with the Sp1-DNA complex. The fact that introducing TAF4 shrnas reduces the promoter activities and the amount of Rta-TAF4 that is bound to the promoter (Fig. 7C, 7D) also indicates that TAF4 is important to the activation of the har promoter by Rta. This study finds that Rta is capable of activating a mutant har promoter that lacks an Sp1-binding site although to a reduced extent (Fig. 7D), suggesting that Rta may affect har transcription through factors other than Sp1 and TAF4. Rta is known to activate many EBV lytic promoters by binding to RRE [4,6 11]. This study finds that after changing the RRE sequence in the BMRF1 promoter, the mutation does not completely eliminate the ability of Rta to activate the promoter. The increase in promoter activity was reduced from 23-fold to eightfold (Fig. 6D). Meanwhile, the knockdown of the expression of TAF4 reduces the ability of Rta to activate the transcription from eightfold to a threefold increase (Fig. 6D), showing that wildtype pbmrf1 and mutant pbmrf1 have similar levels when TAF4 shrna was cotransfected. This result implies that Rta- References 1. Chevallier-Greco A, Manet E, Chavrier P, Mosnier C, Daillie J, et al. (1986) Both Epstein-Barr virus (EBV)-encoded trans-acting factors, EB1 and EB2, are required to activate transcription from an EBV early promoter. EMBO J 5: Kenney S, Holley-Guthrie E, Mar EC, Smith M (1989) The Epstein-Barr virus BMLF1 promoter contains an enhancer element that is responsive to the BZLF1 and BRLF1 transactivators. J Virol 63: Holley-Guthrie EA, Quinlivan EB, Mar EC, Kenney S (1990) The Epstein-Barr virus (EBV) BMRF1 promoter for early antigen (EA-D) is regulated by the EBV transactivators, BRLF1 and BZLF1, in a cell-specific manner. J Virol 64: Quinlivan EB, Holley-Guthrie EA, Norris M, Gutsch D, Bachenheimer SL, et al. (1993) Direct BRLF1 binding is required for cooperative BZLF1/BRLF1 activation of the Epstein-Barr virus early promoter, BMRF1 [corrected and republished with original paging, article originally printed in Nucleic Acids Res 1993 Apr 25;21(8): ]. Nucleic Acids Res 21: Giot JF, Mikaelian I, Buisson M, Manet E, Joab I, et al. (1991) Transcriptional interference between the EBV transcription factors EB1 and R: both DNAbinding and activation domains of EB1 are required. Nucleic Acids Res 19: Kenney S, Holley-Guthrie E, Mar EC, Smith M (1989) The Epstein-Barr virus BMLF1 promoter contains an enhancer element that is responsive to the BZLF1 and BRLF1 transactivators. J Virol 63: Chevallier-Greco A, Gruffat H, Manet E, Calender A, Sergeant A (1989) The Epstein-Barr virus (EBV) DR enhancer contains two functionally different domains: domain A is constitutive and cell specific, domain B is transactivated by the EBV early protein R. J Virol 63: Feederle R, Kost M, Baumann M, Janz A, Drouet E, et al. (2000) The Epstein- Barr virus lytic program is controlled by the co-operative functions of two transactivators. EMBO J 19: Gruffat H, Duran N, Buisson M, Wild F, Buckland R, et al. (1992) Characterization of an R-binding site mediating the R-induced activation of the Epstein-Barr virus BMLF1 promoter. J Virol 66: Liu C, Sista ND, Pagano JS (1996) Activation of the Epstein-Barr virus DNA polymerase promoter by the BRLF1 immediate-early protein is mediated through USF and E2F. J Virol 70: Gruffat H, Manet E, Rigolet A, Sergeant A (1990) The enhancer factor R of Epstein-Barr virus (EBV) is a sequence-specific DNA binding protein. Nucleic Acids Res 18: Ragoczy T, Miller G (2001) Autostimulation of the Epstein-Barr virus BRLF1 promoter is mediated through consensus Sp1 and Sp3 binding sites. J Virol 75: Chang LK, Chung JY, Hong YR, Ichimura T, Nakao M, et al. (2005) Activation of Sp1-mediated transcription by Rta of Epstein-Barr virus via an interaction with MCAF1. Nucleic Acids Res 33: TAF4 complex does not need RRE to activate transcription. Rta and Zta are two transcription factors that critically influence the lytic development of EBV. The results of this study and other investigations [58 61] indicate that both transcription factors interact with TFIID, promoting transcription. However, the activation by Zta seems to be limited to a subset of EBV lytic promoters that contain ZRE and Zta does not activate a promoter without a TATA sequence [18]. However, Rta uses a different strategy, interacting with many transcription factors, including TAF4, Sp1, Zta, MCAF1, and RanBPM, to activate the transcription of both viral and cellular [13,18,62], verifying that Rta activates a broad spectrum of EBV genes [63]. These findings indicate that the two EBV immediate-early transcription factors, although both interact with basal transcription factors, act differently but synergistically to promote the lytic development of EBV. Acknowledgments Dr. Naoko Tanese is appreciated for providing a pbsk-taf4 plasmid. We also want to thank Yung-Chung Yang for his technical assistance. Author Contributions Conceived and designed the experiments: YCY LKC. Performed the experiments: YCY. Analyzed the data: YCY LKC. Contributed reagents/ materials/analysis tools: LKC. Wrote the paper: YCY LKC. 14. Lee YH, Chiu YF, Wang WH, Chang LK, Liu ST (2008) Activation of the ERK signal transduction pathway by Epstein-Barr virus immediate-early protein Rta. J Gen Virol 89: Adamson AL, Darr D, Holley-Guthrie E, Johnson RA, Mauser A, et al. (2000) Epstein-Barr virus immediate-early proteins BZLF1 and BRLF1 activate the ATF2 transcription factor by increasing the levels of phosphorylated p38 and c- Jun N-terminal kinases. J Virol 74: Robinson AR, Kwek SS, Hagemeier SR, Wille CK, Kenney SC (2011) Cellular transcription factor Oct-1 interacts with the Epstein-Barr virus BRLF1 protein to promote disruption of viral latency. J Virol 85: Chua HH, Lee HH, Chang SS, Lu CC, Yeh TH, et al. (2007) Role of the TSG101 gene in Epstein-Barr virus late gene transcription. J Virol 81: Chang LK, Chuang JY, Nakao M, Liu ST (2010) MCAF1 and synergistic activation of the transcription of Epstein-Barr virus lytic genes by Rta and Zta. Nucleic Acids Res 38: Chang LK, Lee YH, Cheng TS, Hong YR, Lu PJ, et al. (2004) Post-translational modification of Rta of Epstein-Barr virus by SUMO-1. J Biol Chem 279: Liu ST, Wang WH, Hong YR, Chuang JY, Lu PJ, et al. (2006) Sumoylation of Rta of Epstein-Barr virus is preferentially enhanced by PIASxbeta. Virus Res 119: Heilmann AM, Calderwood MA, Johannsen E (2010) Epstein-Barr virus LF2 protein regulates viral replication by altering Rta subcellular localization. J Virol 84: Albright SR, Tjian R (2000) TAFs revisited: more data reveal new twists and confirm old ideas. Gene 242: Cler E, Papai G, Schultz P, Davidson I (2009) Recent advances in understanding the structure and function of general transcription factor TFIID. Cell Mol Life Sci 66: Muller F, Tora L (2004) The multicoloured world of promoter recognition complexes. EMBO J 23: Vorobyeva NE, Soshnikova NV, Nikolenko JV, Kuzmina JL, Nabirochkina EN, et al. (2009) Transcription coactivator SAYP combines chromatin remodeler Brahma and transcription initiation factor TFIID into a single supercomplex. Proc Natl Acad Sci U S A 106: Herbig E, Warfield L, Fish L, Fishburn J, Knutson BA, et al. (2010) Mechanism of mediator recruitment by tandem Gcn4 activation domains and three Gal11 activator-binding domains. Mol Cell Biol 30: Wright KJ, Marr MT 2nd, Tjian R (2006) TAF4 nucleates a core subcomplex of TFIID and mediates activated transcription from a TATA-less promoter. Proc Natl Acad Sci U S A 103: Saluja D, Vassallo MF, Tanese N (1998) Distinct subdomains of human TAFII130 are required for interactions with glutamine-rich transcriptional activators. Mol Cell Biol 18: PLOS ONE 11 January 2013 Volume 8 Issue 1 e54075
Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators
 /, 2017, Vol. 8, (No. 53), pp: 91425-91444 Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators Roni Sarkar 1 and Subhash C. Verma 1 1 Department of Microbiology
/, 2017, Vol. 8, (No. 53), pp: 91425-91444 Egr-1 regulates RTA transcription through a cooperative involvement of transcriptional regulators Roni Sarkar 1 and Subhash C. Verma 1 1 Department of Microbiology
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection
 Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Brd4 Coactivates Transcriptional Activation of NF- B via Specific Binding to Acetylated RelA
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
Epstein-Barr Virus Transcription Activator Rta Upregulates Decoy Receptor 3 Expression by Binding to Its Promoter
 JOURNAL OF VIROLOGY, May 2007, p. 4837 4847 Vol. 81, No. 9 0022-538X/07/$08.00 0 doi:10.1128/jvi.02448-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Epstein-Barr Virus Transcription
JOURNAL OF VIROLOGY, May 2007, p. 4837 4847 Vol. 81, No. 9 0022-538X/07/$08.00 0 doi:10.1128/jvi.02448-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Epstein-Barr Virus Transcription
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Epstein-Barr Virus BZLF1 Protein Binds to Mitotic Chromosomes
 JOURNAL OF VIROLOGY, June 2005, p. 7899 7904 Vol. 79, No. 12 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.12.7899 7904.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Epstein-Barr
JOURNAL OF VIROLOGY, June 2005, p. 7899 7904 Vol. 79, No. 12 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.12.7899 7904.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Epstein-Barr
Epstein Barr Virus (EBV) Late Gene Transcription Depends on the Assembly of a Viral Specific Pre-Initiation Complex
 JVI Accepts, published online ahead of print on 27 August 2014 J. Virol. doi:10.1128/jvi.02139-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14
JVI Accepts, published online ahead of print on 27 August 2014 J. Virol. doi:10.1128/jvi.02139-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14
Activation of Gene Expression by Human Herpes Virus 6
 Activation of Gene Expression by Human Herpes Virus 6 M. E. M. Campbell and S. McCorkindale 1 Introduction Human herpes virus type 6 (HHV-6) was first detected by Salahuddin et al. [6] and has been isolated
Activation of Gene Expression by Human Herpes Virus 6 M. E. M. Campbell and S. McCorkindale 1 Introduction Human herpes virus type 6 (HHV-6) was first detected by Salahuddin et al. [6] and has been isolated
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplemental Information
 Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Information
 Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
~Lentivirus production~
 ~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
X/01/$ DOI: /JVI Copyright 2001, American Society for Microbiology. All Rights Reserved.
 JOURNAL OF VIROLOGY, July 2001, p. 6135 6142 Vol. 75, No. 13 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.13.6135 6142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Epstein-Barr
JOURNAL OF VIROLOGY, July 2001, p. 6135 6142 Vol. 75, No. 13 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.13.6135 6142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Epstein-Barr
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Tyrosine 112 of Latent Membrane Protein 2A Is Essential for Protein Tyrosine Kinase Loading and Regulation of Epstein-Barr Virus Latency
 JOURNAL OF VIROLOGY, Oct. 1998, p. 7796 7806 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Tyrosine 112 of Latent Membrane Protein 2A Is
JOURNAL OF VIROLOGY, Oct. 1998, p. 7796 7806 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Tyrosine 112 of Latent Membrane Protein 2A Is
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
The Epstein-Barr virus BcRF1 gene product is a TBP-like protein with an essential role in late gene expression.
 JVI Accepts, published online ahead of print on 28 March 2012 J. Virol. doi:10.1128/jvi.00159-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 The Epstein-Barr virus BcRF1
JVI Accepts, published online ahead of print on 28 March 2012 J. Virol. doi:10.1128/jvi.00159-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 The Epstein-Barr virus BcRF1
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
Doctoral Degree Program in Marine Biotechnology, College of Marine Sciences, Doctoral Degree Program in Marine Biotechnology, Academia Sinica, Taipei,
 Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
The K-bZIP Protein from Kaposi s Sarcoma-Associated Herpesvirus Interacts with p53 and Represses Its Transcriptional Activity
 JOURNAL OF VIROLOGY, Dec. 2000, p. 11977 11982 Vol. 74, No. 24 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The K-bZIP Protein from Kaposi s Sarcoma-Associated
JOURNAL OF VIROLOGY, Dec. 2000, p. 11977 11982 Vol. 74, No. 24 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The K-bZIP Protein from Kaposi s Sarcoma-Associated
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Received 25 November 2003/Accepted 7 April 2004
 JOURNAL OF VIROLOGY, Aug. 2004, p. 8615 8629 Vol. 78, No. 16 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.16.8615 8629.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Kaposi s
JOURNAL OF VIROLOGY, Aug. 2004, p. 8615 8629 Vol. 78, No. 16 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.16.8615 8629.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Kaposi s
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
supplementary information
 Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Received 6 July 2010/Accepted 1 November 2010
 JOURNAL OF VIROLOGY, Jan. 2011, p. 784 794 Vol. 85, No. 2 0022-538X/11/$12.00 doi:10.1128/jvi.01397-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. The Nuclear and Adherent Junction
JOURNAL OF VIROLOGY, Jan. 2011, p. 784 794 Vol. 85, No. 2 0022-538X/11/$12.00 doi:10.1128/jvi.01397-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. The Nuclear and Adherent Junction
of Nebraska - Lincoln
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 12-2001 Identification of a Cellular Protein That Interacts and Synergizes
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 12-2001 Identification of a Cellular Protein That Interacts and Synergizes
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Neither the activation nor suppression of the MAPK pathway affects the ASK1/Vif interaction. (a, b) HEK293 cells were cotransfected with plasmids encoding
Supplementary Figure 1 Supplementary Figure 1. Neither the activation nor suppression of the MAPK pathway affects the ASK1/Vif interaction. (a, b) HEK293 cells were cotransfected with plasmids encoding
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Epstein-Barr Virus (EBV) SM Protein Induces and Recruits Cellular Sp110b To Stabilize mrnas and Enhance EBV Lytic Gene Expression
 JOURNAL OF VIROLOGY, Sept. 2004, p. 9412 9422 Vol. 78, No. 17 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.17.9412 9422.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Epstein-Barr
JOURNAL OF VIROLOGY, Sept. 2004, p. 9412 9422 Vol. 78, No. 17 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.17.9412 9422.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Epstein-Barr
Supplements. Figure S1. B Phalloidin Alexa488
 Supplements A, DMSO, PP2, PP3 Crk-myc Figure S1. (A) Src kinase activity is necessary for recruitment of Crk to Nephrin cytoplasmic domain. Human podocytes expressing /7-NephrinCD () were treated with
Supplements A, DMSO, PP2, PP3 Crk-myc Figure S1. (A) Src kinase activity is necessary for recruitment of Crk to Nephrin cytoplasmic domain. Human podocytes expressing /7-NephrinCD () were treated with
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting. protein3) regulate autophagy and mitophagy in renal tubular cells in. acute kidney injury
 Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting protein3) regulate autophagy and mitophagy in renal tubular cells in acute kidney injury by Masayuki Ishihara 1, Madoka Urushido 2, Kazu Hamada
Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting protein3) regulate autophagy and mitophagy in renal tubular cells in acute kidney injury by Masayuki Ishihara 1, Madoka Urushido 2, Kazu Hamada
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element
 Research Article ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element Evelyn Chukwurah, Indhira Handy and Rekha C. Patel Department
Research Article ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element Evelyn Chukwurah, Indhira Handy and Rekha C. Patel Department
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
Chromatin IP (Isw2) Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles.
 Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
p = formed with HCI-001 p = Relative # of blood vessels that formed with HCI-002 Control Bevacizumab + 17AAG Bevacizumab 17AAG
 A.. Relative # of ECs associated with HCI-001 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 ol b p < 0.001 Relative # of blood vessels that formed with HCI-001 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 l b p = 0.002 Control IHC:
A.. Relative # of ECs associated with HCI-001 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 ol b p < 0.001 Relative # of blood vessels that formed with HCI-001 1.4 1.2 1.0 0.8 0.6 0.4 0.2 0.0 l b p = 0.002 Control IHC:
SUPPLEMENT. Materials and methods
 SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
Supplementary Figure 1
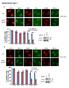 Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
 The Nuclear Chaperone Nucleophosmin Escorts an Epstein-Barr Virus Nuclear Antigen to Establish Transcriptional Cascades for Latent Infection in Human B Cells The Harvard community has made this article
The Nuclear Chaperone Nucleophosmin Escorts an Epstein-Barr Virus Nuclear Antigen to Establish Transcriptional Cascades for Latent Infection in Human B Cells The Harvard community has made this article
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format:
 Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Impact of hyper-o-glcnacylation on apoptosis and NF-κB activity SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
Recruitment of HBO1 Histone Acetylase and Blocks
 Molecular Cell, Volume 44 Supplemental Information JNK1 Phosphorylation of Cdt1 Inhibits Recruitment of HO1 Histone cetylase and locks Replication Licensing in Response to Stress enoit Miotto and Kevin
Molecular Cell, Volume 44 Supplemental Information JNK1 Phosphorylation of Cdt1 Inhibits Recruitment of HO1 Histone cetylase and locks Replication Licensing in Response to Stress enoit Miotto and Kevin
Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication
 Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
of Nebraska - Lincoln
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for February 2008 Mutual Inhibition between Kaposi s Sarcoma- Associated Herpesvirus
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for February 2008 Mutual Inhibition between Kaposi s Sarcoma- Associated Herpesvirus
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
Edinburgh Research Explorer
 Edinburgh Research Explorer Rta of murine gammaherpesvirus 68 reactivates the complete lytic cycle from latency Citation for published version: Wu, TT, Usherwood, EJ, Stewart, JP, Nash, AA & Sun, R 2000,
Edinburgh Research Explorer Rta of murine gammaherpesvirus 68 reactivates the complete lytic cycle from latency Citation for published version: Wu, TT, Usherwood, EJ, Stewart, JP, Nash, AA & Sun, R 2000,
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the
 Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi: /jvi
 JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi:10.1128/jvi.00379-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 19 May 2010 J. Virol. doi:10.1128/jvi.00379-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Supplementary Figure 1
 Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3
 Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
/06/$15.00/0 Molecular Endocrinology 20(9): Copyright 2006 by The Endocrine Society doi: /me
 0888-8809/06/$15.00/0 Molecular Endocrinology 20(9):2062 2079 Printed in U.S.A. Copyright 2006 by The Endocrine Society doi: 10.1210/me.2005-0316 Androgens, Progestins, and Glucocorticoids Induce Follicle-Stimulating
0888-8809/06/$15.00/0 Molecular Endocrinology 20(9):2062 2079 Printed in U.S.A. Copyright 2006 by The Endocrine Society doi: 10.1210/me.2005-0316 Androgens, Progestins, and Glucocorticoids Induce Follicle-Stimulating
Supplementary Information
 Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts
 Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Autoregulation of DNA Binding and Protein Stability of Kaposi s Sarcoma-Associated Herpesvirus ORF50 Protein
 JOURNAL OF VIROLOGY, Oct. 2004, p. 10657 10673 Vol. 78, No. 19 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.19.10657 10673.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Autoregulation
JOURNAL OF VIROLOGY, Oct. 2004, p. 10657 10673 Vol. 78, No. 19 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.19.10657 10673.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Autoregulation
Interferon Regulatory Factor 7 Is Negatively Regulated by the Epstein-Barr Virus Immediate-Early Gene, BZLF-1
 JOURNAL OF VIROLOGY, Aug. 2005, p. 10040 10052 Vol. 79, No. 15 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.15.10040 10052.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Interferon
JOURNAL OF VIROLOGY, Aug. 2005, p. 10040 10052 Vol. 79, No. 15 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.15.10040 10052.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Interferon
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet
 Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
The C-Terminal Region but Not the Arg-X-Pro Repeat of Epstein-Barr Virus Protein EB2 Is Required for Its Effect on RNA Splicing and Transport
 JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
