Nature Structural & Molecular Biology: doi: /nsmb.2448
|
|
|
- Gwenda Wilcox
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. Chromatin immunoprecipitation (ChIP) for Smad1, Zfx and E2f1 on histone gene promoters. From left to right: ChIP for Smad1 on H1 and H4 promoters, Zfx on H1 promoter, E2f1 on H1, H2a, H2b, H3 and H4 promoters. Next three bars and right-most bars show no enrichment for these factors on Serpin promoter and on an intergenic region, both of which were used as negative controls.
2 Supplementary Figure 2. Fold enrichment of E2f4 in different tissues. 1.7-fold in adipocyte (P = 1.3x10-3), 1.9-fold in liver (P = 1.8x10-4), 3.1-fold in Myoblast (P < ), 3.9-fold in ESCs (P < ), 5.4-fold in B-cell lymphoma (P < ) and 7.5-fold in erythroleukemia (P < ).
3 Supplementary Figure 3. Read density of p300 on chromosome 13 in ESC, forebrain, midbrain and limb bud.
4 Supplementary Figure 4. Histone genes are enriched with histone acetylation marks. Expected Vs. observed percentage of histones genes with modification.
5 Supplementary Figure 5. Percentage of histone genes bound by Tet1, within each subtype.
6 Supplementary Figure 6. Histone genes are enriched with Med1, Med12, Smc1, Smc3, Nipbl and Tet1. Read density of various TFs within locus chr13:
7 Supplementary Table 1 References to and datasets TF/Histone modification Nanog, Pou5f1, Stat3, Smad1, Sox2, Zfx, c- Myc, n-myc, Klf4, Esrrb, Tcfcp2l1, E2f1 and CTCF Med1, Med12, Nipbl, Smc1, Smc3 Atrx Brg1 p300 Tet1 Sall1 Setdb1 Tcf3, H3K79me2 Jarid2 Smad3 Cnot3, Trim28 Reference Chen X, Xu H, Yuan P, Fang F, Huss M, et al. (2008) Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133: Kagey MH, Newman JJ, Bilodeau S, Zhan Y, Orlando DA, et al. (2010) Mediator and cohesin connect gene expression and chromatin architecture. Nature 467: Law MJ, Lower KM, Voon HP, Hughes JR, Garrick D, et al. (2010) ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size-dependent manner. Cell 143: Ho L, Jothi R, Ronan JL, Cui K, Zhao K, et al. (2009) An embryonic stem cell chromatin remodeling complex, esbaf, is an essential component of the core pluripotency transcriptional network. Proc Natl Acad Sci U S A 106: Schnetz MP, Handoko L, Akhtar-Zaidi B, Bartels CF, Pereira CF, et al. (2010) CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS Genet 6: e Xu Y, Wu F, Tan L, Kong L, Xiong L, et al. (2011) Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell 42: Karantzali E, Lekakis V, Ioannou M, Hadjimichael C, Papamatheakis J, et al. (2011) Sall1 regulates embryonic stem cell differentiation in association with nanog. J Biol Chem 286: Bilodeau S, Kagey MH, Frampton GM, Rahl PB, Young RA (2009) SetDB1 contributes to repression of genes encoding developmental regulators and maintenance of ES cell state. Genes Dev 23: Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, et al. (2008) Connecting microrna genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134: Shen X, Kim W, Fujiwara Y, Simon MD, Liu Y, et al. (2009) Jumonji modulates polycomb activity and selfrenewal versus differentiation of stem cells. Cell 139: Mullen AC, Orlando DA, Newman JJ, Loven J, Kumar RM, et al. (2011) Master transcription factors determine cell-type-specific responses to TGF-beta signaling. Cell 147: Hu G, Kim J, Xu Q, Leng Y, Orkin SH, et al. (2009) A Type of Experiment
8 Ezh1 Ezh2, YY1 (ESC,NPC) CTtr9 Nr0b(Dax1), Rex1, Zfp281, Nax1, c-myc Nr5a2 Gcn5, Tip60, E2f4 Sall4a, Sall4b Smad2 p300 (limb bud, forebrain, midbrain) Smad4 E2f4 (B-cell lymphoma, myoblast, erythroleukemia) E2f4 (liver, adipocyte) genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev 23: Shen X, Liu Y, Hsu YJ, Fujiwara Y, Kim J, et al. (2008) EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol Cell 32: Mendenhall EM, Koche RP, Truong T, Zhou VW, Issac B, et al. (2010) GC-rich sequence elements recruit PRC2 in mammalian ES cells. PLoS Genet 6: e Ding L, Paszkowski-Rogacz M, Nitzsche A, Slabicki MM, Heninger AK, et al. (2009) A genome-scale RNAi screen for Oct4 modulators defines a role of the Paf1 complex for embryonic stem cell identity. Cell Stem Cell 4: Kim J, Chu J, Shen X, Wang J, Orkin SH (2008) An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132: Heng JC, Feng B, Han J, Jiang J, Kraus P, et al. (2010) The nuclear receptor Nr5a2 can replace Oct4 in the reprogramming of murine somatic cells to pluripotent cells. Cell Stem Cell 6: Kim J, Woo AJ, Chu J, Snow JW, Fujiwara Y, et al. (2010) A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 143: Rao S, Zhen S, Roumiantsev S, McDonald LT, Yuan GC, et al. (2010) Differential roles of Sall4 isoforms in embryonic stem cell pluripotency. Mol Cell Biol 30: L ee KL, Lim SK, Orlov YL, Yit le Y, Yang H, et al. (2011) Graded Nodal/Activin signaling titrates conversion of quantitative phospho-smad2 levels into qualitative embryonic stem cell fate decisions. PLoS Genet 7: e Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, et al. (2009) accurately predicts tissue-specific activity of enhancers. Nature 457: Fei T, Xia K, Li Z, Zhou B, Zhu S, et al. (2010) Genome-wide mapping of Smad target genes reveals the role of BMP signaling in embryonic stem cell fate determination. Genome Res 20: ENCODE Project Consortium, Myers RM, Stamatoyannopoulos J, Snyder M, Dunham I, Hardison RC, Bernstein BE, Gingeras TR, Kent WJ, Birney E et al. (2011) A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9: e MacIsaac KD, Lo KA, Gordon W, Motola S, Mazor T, et al. (2010) A quantitative model of transcriptional regulation reveals the influence of binding location on expression. PLoS Comput Biol. 6: e
9 H3K9ac H3K27ac, H3K4me1, H3K4me3 H3K9me3, H3K27me3, H3K36me3, H4K20me3 H3K4me2 RNA Pol II Hezroni H, Sailaja BS, Meshorer E (2011) Pluripotency-related, valproic acid (VPA)-induced genome-wide histone H3 lysine 9 (H3K9) acetylation patterns in embryonic stem cells. J Biol Chem 286: Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, et al. (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107: Mikkelsen TS, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature. 2007;448: Meissner A, et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells.nature 2008;454: Seila AC, et al. Divergent transcription from active promoters. Science :
10 Supplementary Table 2 List of primer sequences used for ChIP Name Gene Sequence Smad1_H1 Hist1h1d F: TTTCCAATCCGGCTACACAAGGTG R: CTGATTGGTGCAACACCCATGACT Smad1_H4 Hist1h4i F: TGCCTACTGGAACTAAATGCGTGC R: GCATCCTCGTTCAGTCCATAATTGGG Zfx_H1 Hist1h1a F: CTTCGTCTTGGGTATAGCTGTTGT R: ACGCTTCGCCATGTCCGAGA E2f1_H1 Hist1h1b F: AACAGCCTTAGTGATGAGCTCGGA R: TCCCGCCAAGAAGAAGACAACGAA E2f1_H2A Hist1h2ae F: TGTTTGCCACGTCCAGACATTGAC R: CGAGTATAAGTAGCTTCTCGTCTGGC E2f1_H2B Hist1h2bk F: TACCCGTGGTGTTCCTATCTGCTT R: AGGCTCAGGCATGGCGTAGAAAT E2f1_H3 Hist1h3a F: CTAGAAAGCAAGCTGGTGAGCACT R: TCATCCAATCACTATCCTTTGCAT E2f1_H4 Hist1h4n F: TGCCACGACCAGACATGATGAAGA R: TCCGGTCCGCAAGTTCCCTATAA Genic Serpin F: GCTGGAAGCACTTACTTGTGTTTCTGAAGG R: AGATCTCAGGATCAGCCACACACA Intergenic F: TGCATGTATTACTTTGTGGAATTT R: CCTTTCTCACAAACTAGCTCATTAC
11 Supplementary Table 3 Smad2 knockout under TGF-β stimulated and non-stimulated conditions Fold change TGF-β (+)/TGF-β (-) Affymetrix ID Gene SMAD2 WT SMAD2 KO No TGF-β stimulation Fold change (KO/WT) TGF-β stimulated _a_at Hist1h1c _at Hist1h2bc _at Hist2h2aa _x_at Hist2h2aa1, Hist2h2aa _at Hist1h1e _s_at H _at Hist1h4i _x_at Hist1h2bp _at Hist1h3d _at Hist1h4h _at Hist1h3f, Hist1h3e _at Hist3h2a _s_at Hist3h2a _at Hist1h2ao _at Hist1h1t _a_at H2b _at Hist1h3d _s_at H
12 Supplementary Table 4 Smad2 activation/inhibition Affymetrix ID Gene Activation fold change Inhibition fold change Controlaverage Activationaverage Inhibitionaverage _a_at Hist1h1c _at Hist1h2bc _at Hist2h2aa _x_at Hist2h2aa1, Hist2h2aa _s_at H _at Hist1h3f, Hist1h3e _at Hist3h2a _a_at Hist1h1c _at Hist1h2ao _at Hist3h2ba _a_at H2b _at Hist1h3d _s_at H
Genome Control in Cell Identity and Disease! Development and cell identity Loss of cell identity and disease New diagnostics and therapeutics
 Genome Control in Cell Identity and Disease! Development and cell identity Loss of cell identity and disease New diagnostics and therapeutics Development and cell identity! 30,000,000,000 cells!! Control
Genome Control in Cell Identity and Disease! Development and cell identity Loss of cell identity and disease New diagnostics and therapeutics Development and cell identity! 30,000,000,000 cells!! Control
ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to
 Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Supplemental Methods RNA-seq ESCs were lysed with Trizol reagent (Life technologies) and RNA was extracted according to the manufacturer's instructions. RNAse free DNaseI (Sigma) was used to eliminate
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
ubiquitinylation succinylation butyrylation hydroxylation crotonylation
 Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells
 Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq Data
 BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
BioMed Volume 2016, Article ID 2395341, 6 pages http://dx.doi.org/10.1155/2016/2395341 Research Article Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
H3K27me3 may be associated with Oct4 and Sox2 in mouse preimplantation embryos
 H3K27me3 may be associated with Oct4 and Sox2 in mouse preimplantation embryos F.-R. Wu, Y. Zhang, B. Ding, X.-H. Lei, J.-C. Huang, C.-H. Wang, Y. Liu, R. Wang and W.-Y. Li School of Biological Science
H3K27me3 may be associated with Oct4 and Sox2 in mouse preimplantation embryos F.-R. Wu, Y. Zhang, B. Ding, X.-H. Lei, J.-C. Huang, C.-H. Wang, Y. Liu, R. Wang and W.-Y. Li School of Biological Science
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Long noncoding RNAs in DNA methylation: new players stepping into the old game
 DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
DOI 10.1186/s13578-016-0109-3 Cell & Bioscience REVIEW Long noncoding RNAs in DNA methylation: new players stepping into the old game Yu Zhao 1, Hao Sun 1,2* and Huating Wang 1,3* Open Access Abstract
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Cluster Dendrogram. dist(cor(na.omit(tss.exprs.chip[, c(1:10, 24, 27, 30, 48:50, dist(cor(na.omit(tss.exprs.chip[, c(1:99, 103, 104, 109, 110,
 A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
1,2,3 1,2 1 1,2 1,2 1 1,2 1,2 (1.,, ) (2.,, ) (3.,, )
 34 0 Vol.34 No.0 204 0 Oct. 204 Reproduction & ontraception doi: 0.7669/j.issn.0253-357X.204.0.0789 E-mail: randc_journal@63.com 5-23 2 2 2 2 2. 236037) 2. 236037) 3. 23060) : 5-5-azacytidine ) : 0. μmol/l
34 0 Vol.34 No.0 204 0 Oct. 204 Reproduction & ontraception doi: 0.7669/j.issn.0253-357X.204.0.0789 E-mail: randc_journal@63.com 5-23 2 2 2 2 2. 236037) 2. 236037) 3. 23060) : 5-5-azacytidine ) : 0. μmol/l
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Figure S1, Beyer et al.
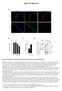 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells
 SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
Replication-dependent histone genes are actively transcribed in differentiating and aging retinal neurons
 REPORT Cell Cycle 13:16, 2526--2541; August 15, 2014; 2014 Taylor & Francis Group, LLC Replication-dependent histone genes are actively transcribed in differentiating and aging retinal neurons Abdul Rouf
REPORT Cell Cycle 13:16, 2526--2541; August 15, 2014; 2014 Taylor & Francis Group, LLC Replication-dependent histone genes are actively transcribed in differentiating and aging retinal neurons Abdul Rouf
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
STEM CELL GENETICS AND GENOMICS
 STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
STEM CELL GENETICS AND GENOMICS Concise Review: Roles of Polycomb Group Proteins in Development and Disease: A Stem Cell Perspective VINAGOLU K. RAJASEKHAR, a MARTIN BEGEMANN b a Memorial Sloan-Kettering
Imprinting. Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821
 Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Imprinting Joyce Ohm Cancer Genetics and Genomics CGP-L2-319 x8821 Learning Objectives 1. To understand the basic concepts of genomic imprinting Genomic imprinting is an epigenetic phenomenon that causes
Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen
 Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen Receptor Signaling to PI3K/AKT Tiffany G. Bredfeldt, Kristen L. Greathouse, Stephen H. Safe, Mien-Chie Hung, Mark
Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen Receptor Signaling to PI3K/AKT Tiffany G. Bredfeldt, Kristen L. Greathouse, Stephen H. Safe, Mien-Chie Hung, Mark
Super-Enhancers in the Control of Cell Identity and Disease
 Resource Super-Enhancers in the Control of Cell Identity and Disease Denes Hnisz,, Brian J. Abraham,, Tong Ihn Lee,, Ashley Lau,, Violaine Saint-André, Alla A. Sigova, Heather A. Hoke,, and Richard A.
Resource Super-Enhancers in the Control of Cell Identity and Disease Denes Hnisz,, Brian J. Abraham,, Tong Ihn Lee,, Ashley Lau,, Violaine Saint-André, Alla A. Sigova, Heather A. Hoke,, and Richard A.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
review Enhancers: emerging roles in cell fate specification review Chin-Tong Ong & Victor G. Corces +
 review Enhancers: emerging roles in cell fate specification Chin-Tong Ong & Victor G. Corces + Department of Biology, Emory University, Atlanta, Georgia, USA Enhancers are regulatory DNA elements that
review Enhancers: emerging roles in cell fate specification Chin-Tong Ong & Victor G. Corces + Department of Biology, Emory University, Atlanta, Georgia, USA Enhancers are regulatory DNA elements that
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Mesenchymal progenitor cells in intramuscular connective tissue development
 Mesenchymal progenitor cells in intramuscular connective tissue development Min Du, Ph.D. Professor and Endowed Chair Department of Animal Sciences Washington State University Pullman, WA Beef Quality:
Mesenchymal progenitor cells in intramuscular connective tissue development Min Du, Ph.D. Professor and Endowed Chair Department of Animal Sciences Washington State University Pullman, WA Beef Quality:
arxiv: v1 [q-bio.gn] 15 Oct 2010
![arxiv: v1 [q-bio.gn] 15 Oct 2010 arxiv: v1 [q-bio.gn] 15 Oct 2010](/thumbs/77/76121917.jpg) Mapping Dynamic Histone Acetylation Patterns to Gene Expression 1 arxiv:1010.3268v1 [q-bio.gn] 15 Oct 2010 Mapping Dynamic Histone Acetylation Patterns to Gene Expression in Nanog-depleted Murine Embryonic
Mapping Dynamic Histone Acetylation Patterns to Gene Expression 1 arxiv:1010.3268v1 [q-bio.gn] 15 Oct 2010 Mapping Dynamic Histone Acetylation Patterns to Gene Expression in Nanog-depleted Murine Embryonic
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Connecting microrna Genes to the Core Transcriptional Regulatory Circuitry of Embryonic Stem Cells
 Resource Connecting microrna Genes to the Core Transcriptional Regulatory Circuitry of Embryonic Stem Cells Alexander Marson, 1,2,5 Stuart S. Levine, 1,5 Megan F. Cole, 1,2 Garrett M. Frampton, 1,2 Tobias
Resource Connecting microrna Genes to the Core Transcriptional Regulatory Circuitry of Embryonic Stem Cells Alexander Marson, 1,2,5 Stuart S. Levine, 1,5 Megan F. Cole, 1,2 Garrett M. Frampton, 1,2 Tobias
Genomic Methods in Cancer Epigenetic Dysregulation
 Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Genomic Methods in Cancer Epigenetic Dysregulation Clara, Lyon 2018 Jacek Majewski, Associate Professor Department of Human Genetics, McGill University Montreal, Canada A few words about my lab Genomics
Transcriptional repression of Xi
 Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
Transcriptional repression of Xi Xist Transcription of Xist Xist RNA Spreading of Xist Recruitment of repression factors. Stable repression Translocated Xic cannot efficiently silence autosome regions.
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies
 CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Promiscuous RNA binding by Polycomb Repressive Complex 2
 Promiscuous RNA binding by Polycomb Repressive Complex 2 Chen Davidovich 1,2, Leon Zheng 3, Karen J. Goodrich 1,2, and Thomas R. Cech 1,2,4 1 Department of Chemistry & Biochemistry, Howard Hughes Medical
Promiscuous RNA binding by Polycomb Repressive Complex 2 Chen Davidovich 1,2, Leon Zheng 3, Karen J. Goodrich 1,2, and Thomas R. Cech 1,2,4 1 Department of Chemistry & Biochemistry, Howard Hughes Medical
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
IN THE NAME OF GOD CANCER CELL REPROGRAMING
 IN THE NAME OF GOD CANCER CELL REPROGRAMING Qazvin university of medical science Presented by: fatane abedy GUIDANCE: Dr. gheibi 1 CONTENT: CANCER REPROGRAMMING OSKM microrna PROBLEM AND ADVANTAGE REFRANCE
IN THE NAME OF GOD CANCER CELL REPROGRAMING Qazvin university of medical science Presented by: fatane abedy GUIDANCE: Dr. gheibi 1 CONTENT: CANCER REPROGRAMMING OSKM microrna PROBLEM AND ADVANTAGE REFRANCE
Supplementary Data. (continued)
 Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
Supplementary Data SUPPLEMENTARY FIG. S1. Gating strategy for Figure 1. Splenocytes were gated for dimension and for positivity to the different markers representing different cell populations. (A) B cells,
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
MicroRNA-mediated incoherent feedforward motifs are robust
 The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
This is a published version of a paper published in PLoS genetics. Access to the published version may require subscription.
 Umeå University This is a published version of a paper published in PLoS genetics. Citation for the published paper: Holmqvist, P., Boija, A., Philip, P., Crona, F., Stenberg, P. et al. (2012) "Preferential
Umeå University This is a published version of a paper published in PLoS genetics. Citation for the published paper: Holmqvist, P., Boija, A., Philip, P., Crona, F., Stenberg, P. et al. (2012) "Preferential
Wnt/β-catenin s regulatory role in embryonic stem cell pluripotency and differentiation
 Wnt/β-catenin s regulatory role in embryonic stem cell pluripotency and differentiation Wim de Jonge 3361039 Supervisor: Prof. Dr. S.J.L. van den Heuvel Group: Developmental Biology, Faculty of Science,
Wnt/β-catenin s regulatory role in embryonic stem cell pluripotency and differentiation Wim de Jonge 3361039 Supervisor: Prof. Dr. S.J.L. van den Heuvel Group: Developmental Biology, Faculty of Science,
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
The genetics of heterochromatin. in metazoa. mutations by means of X-ray irradiation" "for the discovery of the production of
 The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
The genetics of heterochromatin in metazoa 1 Hermann Joseph Muller 1946 Nobel Prize in Medicine: "for the discovery of the production of mutations by means of X-ray irradiation" 3 4 The true meaning of
Sudin Bhattacharya Institute for Integrative Toxicology
 Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Histone modifications in kidney disease
 Histone modifications in kidney disease Masaomi Nangaku Division of Nephrology and Endocrinology The University of Tokyo Graduate School of Medicine Japan Mimura, Tanaka, & Nangaku. Semin Nephrol 2013
Histone modifications in kidney disease Masaomi Nangaku Division of Nephrology and Endocrinology The University of Tokyo Graduate School of Medicine Japan Mimura, Tanaka, & Nangaku. Semin Nephrol 2013
ChipSeq. Technique and science. The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation
 Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins?
 REVIEW ARTICLE The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins? Andrea Scelfo, Andrea Piunti and Diego Pasini Department
REVIEW ARTICLE The controversial role of the Polycomb group proteins in transcription and cancer: how much do we not understand Polycomb proteins? Andrea Scelfo, Andrea Piunti and Diego Pasini Department
Yingying Wei George Wu Hongkai Ji
 Stat Biosci (2013) 5:156 178 DOI 10.1007/s12561-012-9066-5 Global Mapping of Transcription Factor Binding Sites by Sequencing Chromatin Surrogates: a Perspective on Experimental Design, Data Analysis,
Stat Biosci (2013) 5:156 178 DOI 10.1007/s12561-012-9066-5 Global Mapping of Transcription Factor Binding Sites by Sequencing Chromatin Surrogates: a Perspective on Experimental Design, Data Analysis,
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition
 AD Award Number: W81XWH-13-1-0336 TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition PRINCIPAL INVESTIGATOR: Mary-Claire King,
AD Award Number: W81XWH-13-1-0336 TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition PRINCIPAL INVESTIGATOR: Mary-Claire King,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature19357 Figure 1a Chd8 +/+ Chd8 +/ΔSL Chd8 +/+ Chd8 +/ΔL E10.5_Whole brain E10.5_Whole brain E10.5_Whole brain E14.5_Whole brain E14.5_Whole brain E14.5_Whole
SUPPLEMENTARY INFORMATION doi:10.1038/nature19357 Figure 1a Chd8 +/+ Chd8 +/ΔSL Chd8 +/+ Chd8 +/ΔL E10.5_Whole brain E10.5_Whole brain E10.5_Whole brain E14.5_Whole brain E14.5_Whole brain E14.5_Whole
Reprogramming through micrornas Stefanie Dimmeler
 Klinikum der Johann Wolfgang Goethe Universität Frankfurt am Main Reprogramming through micrornas Stefanie Dimmeler Conflict of interest: T2cure GmbH, Miragen Non-coding DNA & RNA and micrornas Human Genome
Klinikum der Johann Wolfgang Goethe Universität Frankfurt am Main Reprogramming through micrornas Stefanie Dimmeler Conflict of interest: T2cure GmbH, Miragen Non-coding DNA & RNA and micrornas Human Genome
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
Name: Xueming Zhao. Professional Title: Professor. Animal embryo biotechnology, mainly including in vitro maturation (IVM), in vitro fertilization
 Name: Xueming Zhao Professional Title: Professor Telephone:86-010-62815892 Fax:86-010-62895971 E-mail: zhaoxueming@caas.cn Website: http://www.iascaas.net.cn/yjspy/dsjj/sssds/dwyzyzypz1/62040.htm Research
Name: Xueming Zhao Professional Title: Professor Telephone:86-010-62815892 Fax:86-010-62895971 E-mail: zhaoxueming@caas.cn Website: http://www.iascaas.net.cn/yjspy/dsjj/sssds/dwyzyzypz1/62040.htm Research
Large conserved domains of low DNA methylation maintained by Dnmt3a
 Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC)
 Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Lung Met 1 Lung Met 2 Lung Met Lung Met H3K4me1. Lung Met H3K27ac Primary H3K4me1
 a Gained Met-VELs 1.5 1.5 -.5 Lung Met 1 Lung Met Lung Met 3 1. Lung Met H3K4me1 Lung Met H3K4me1 1 Lung Met H3K4me1 Lung Met H3K7ac 1.5 Lung Met H3K7ac Lung Met H3K7ac.8 Primary H3K4me1 Primary H3K7ac
a Gained Met-VELs 1.5 1.5 -.5 Lung Met 1 Lung Met Lung Met 3 1. Lung Met H3K4me1 Lung Met H3K4me1 1 Lung Met H3K4me1 Lung Met H3K7ac 1.5 Lung Met H3K7ac Lung Met H3K7ac.8 Primary H3K4me1 Primary H3K7ac
SetDB1 Contributes to Repression of Genes Encoding Developmental Regulators and Maintenance of ES Cell State
 SUPPLEMENTAL INFORMATION SetDB1 Contributes to Repression of Genes Encoding Developmental Regulators and Maintenance of ES Cell State Steve Bilodeau, Michael Kagey, Garrett M. Frampton, Peter B. Rahl,
SUPPLEMENTAL INFORMATION SetDB1 Contributes to Repression of Genes Encoding Developmental Regulators and Maintenance of ES Cell State Steve Bilodeau, Michael Kagey, Garrett M. Frampton, Peter B. Rahl,
Activin/Nodal signaling and NANOG orchestrate human embryonic stem cell fate decisions by controlling the H3K4me3 chromatin mark
 Activin/Nodal signaling and NANOG orchestrate human embryonic stem cell fate decisions by controlling the H3K4me3 chromatin mark Alessandro Bertero, 1 Pedro Madrigal, 1,2 Antonella Galli, 2 Nina C. Hubner,
Activin/Nodal signaling and NANOG orchestrate human embryonic stem cell fate decisions by controlling the H3K4me3 chromatin mark Alessandro Bertero, 1 Pedro Madrigal, 1,2 Antonella Galli, 2 Nina C. Hubner,
Epigene.cs: What is it and how it effects our health? Overview. Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa
 Epigene.cs: What is it and how it effects our health? Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa Overview Basic Background Epigene.cs in general Epigene.cs in cancer Epigene.cs
Epigene.cs: What is it and how it effects our health? Dr. Bill Stanford, PhD OFawa Hospital Research Ins.tute University of OFawa Overview Basic Background Epigene.cs in general Epigene.cs in cancer Epigene.cs
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors
 Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION. Jude Fitzgibbon
 FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
FOLLICULAR LYMPHOMA- ILLUMINA METHYLATION Jude Fitzgibbon j.fitzgibbon@qmul.ac.uk Molecular Predictors of Clinical outcome Responders Alive Non-responders Dead TUMOUR GENETICS Targeted therapy, Prognostic
Fragile X Syndrome. Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype
 Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Fragile X Syndrome Genetics, Epigenetics & the Role of Unprogrammed Events in the expression of a Phenotype A loss of function of the FMR-1 gene results in severe learning problems, intellectual disability
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Seq-ing Answers to Questions of Transcriptional Regulation and Epigenetics. Theodore J. Perkins OttBUG Seminar Nov.
 Seq-ing Answers to Questions of Transcriptional Regulation and Epigenetics Theodore J. Perkins www.perkinslab.ca OttBUG Seminar Nov. 7, 2016 Outline ChIP-seq What s it for? How does it work? Two examples
Seq-ing Answers to Questions of Transcriptional Regulation and Epigenetics Theodore J. Perkins www.perkinslab.ca OttBUG Seminar Nov. 7, 2016 Outline ChIP-seq What s it for? How does it work? Two examples
The role of G9a gene in various cancers
 Health Biotechnology and Biopharma (2017), 1(3): 5-11 Minireview Article The role of G9a gene in various cancers Hassan Taghizadeh 1, Eskandar Taghizadeh 2 * 1 Cellular and Molecular Research Center, Yasuj
Health Biotechnology and Biopharma (2017), 1(3): 5-11 Minireview Article The role of G9a gene in various cancers Hassan Taghizadeh 1, Eskandar Taghizadeh 2 * 1 Cellular and Molecular Research Center, Yasuj
Transcript-indexed ATAC-seq for immune profiling
 Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Statistical Assessment of the Global Regulatory Role of Histone. Acetylation in Saccharomyces cerevisiae. (Support Information)
 Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation
 Published online: June 1, 15 Article Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation Evelyn N Kouwenhoven 1,, Martin Oti, Hanna Niehues 3, Simon J van
Published online: June 1, 15 Article Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation Evelyn N Kouwenhoven 1,, Martin Oti, Hanna Niehues 3, Simon J van
3) The sheer number of and inconsistency between different animal models used make the paper difficult to follow and may impact data interpretation:
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Pasricha et al. show that iron deficiency (ID) and stimulated erythropoiesis suppress hepcidin via distinct processes. They
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Pasricha et al. show that iron deficiency (ID) and stimulated erythropoiesis suppress hepcidin via distinct processes. They
