Supplementary Figure 1 b
|
|
|
- Marianna Robertson
- 6 years ago
- Views:
Transcription
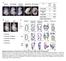
1 Supplementary Figure 1 b 7 c BT-474 MDA-MB-231 migrated cells/field m G oc F k BR P-s i BRL-s L- i1 si BR N 2 M C S1 L 7 BR M S1 N C FP -s i BR Lsi 1 BR Lsi 2 G m oc k L a 3 ****** 2 ### 1 BT-474 MDA-MB-231 mrna expression e d 1.5 vimentin fibronectin 1. ** **.5 Supplementary Figure 1. inhibits migration and EMT of breast cancer cells. (ab) Western blotting of in BT-474 cells that were transfected () or transfected with RNA () or either of the two -sirnas (BRL-si) (a) and in MDA-MB-231 cells that were transfected with pcdna3 vector overexpressing () or an empty vector as negative control () (b). was used as a loading control. (c) Boyden chamber assays for migration of BT-474 cells that were transfected as in (a) and MDA-MB-231 cells transfected as in (b). Bars correspond to mean ± SD. ***, P <.1 as compared with RNA. ###, P <.1, as compared with. Scale bars correspond to 5 µm.(d) Phase-contrast microscopy for MDA-MB-231 cells that were treated as in (b). Scale bar corresponds to 5 µm.(e) qrt-pcr for vimentin and fibronectin mrna expression in MDA-MB-231 cells that were treated as in (b). Bars correspond to mean ± SD. **, P <.1 as compared with.these data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups. BRL,.
2 Supplementary Figure 2 a c e mrna expression/gapdh FZD1 + cells(%) normalized mrna UT FZD1 FZD2 FZD3 FZD4 FZD1 FZD5 FZD6 FZD7 FZD8 FZD9 FZD1 *** *** r=-.762 P< *** b d f mrna expression/gapdh normalized mrna FZD1 FZD1 FZD2 FZD3 FZD1 *** FZD4 FZD5 FZD6 FZD7 FZD8 ### *** FZD9 FZD1 MCF-1A T47D MCF-7 BT-474 MDA-MB-436 MDA-MB cells(%) Supplementary Figure 2. FZD1 expression is negatively associated with. (ab) qrt-pcr for 1 FZD family members in MDA-MB-231 cells transfected with pcdna3 vector overexpressing () or negative control () (a) and in T47D cells that were transfected (), or transfected with RNA or either of the two sirnas (b). Bars correspond to mean ± SD. ***, P <.1 as compared with or GFPsiRNA. (c-d) qrt-pcr for and FZD1 expression in T47D cells that were untreated (UT) or transfected as in b (c) and in MDA-MB-231 cells that were transfected or transfected as in a (d). Bars correspond to mean ± SD. ***, P <.1 as compared with GFPsiRNA or. ###, P <.1 as compared with. (e) Correlation between the percentages of + and FZD1 + breast cancer cells. Linear regression was used to analyze the correlation.(f) FZD1 protein expression measured by western blotting in various breast cell lines. All the data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups. BRL,.
3 Supplementary Figure 3. enhances HDAC1/2 binding to FZD1 promoter and inhibits H3K9 acetylation. (a) MDA-MB-231 cells with stable transfection of pgl3-fzd1- promoter were transfected with pcdna3 vector carrying () or negative control () and assayed for luciferase activity. ***, P <.1 as compared with. (b) Schematic diagram shows primer pairs of the FZD1 proximal and distal promoter regions for ChIP assay.(c) ChIP analysis for binding to FZD1 promoter in MBA-MD-231 cells transfected with Myc- or. (d-e) ChIP analysis for HDAC1 binding to FZD1 promoter in T47D cells transfected with -sirnas or RNA (d) and in MDA-MB-231 cells transfected with pcdna3 vector overexpressing () or negative control () (e) using primer pairs of FZD1 distal promoter as a control. (f-g) ChIP analysis for H3K9Ac binding to FZD1 promoter in T47D cells transfected with -sirnas or GFPsiRNA (f) and in MDA-MB-231 cells transfected with pcdna3 vector overexpressing () or negative control () (g) using primer pairs of FZD1 distal promoter as a control. These data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups.brl,. Supplementary Figure 3 a luciferase activity *** b pgl3-basic pgl3-fzd1 c Distal d e Distal Distal ChIP-IgG ChIP-IgG ChIP-IgG 1. ChIP-anti-Myc 4 ChIP-anti-HDAC1 3 ChIP-anti-HDAC f Bound / input Bound / input Distal Myc- Bound / input 1. ChIP-IgG Distal g Distal Bound / input Bound / input Proximal ChIP-IgG ChIP-anti-H3K9Ac FZD1
4 Supplementary Figure 4 a SP1 b d GAGGAGAGGCGGGCGGGACTGCCGGGGAGGG normalized SP ** **.2 SP1-si1 SP1-si2 Distal SP c SP1 FZD1 SP1 1 7 e Bound / input Distal ChIP-IgG.7.6 ChIP-anti-SP f Bound / input Distal.25 ChIP-IgG.2 ChIP-anti-SP Supplementary Figure 4. suppresses SP1-mediated FZD1 transcription. (a) Sequence of predicted SP1-binding site located between -4 to -25 bp of FZD1 promoter. Point mutation sites for FZD1-2-SP1-Mut-Luc recombinant plasmid were indicated as bold.(b) SP1 was efficiently silenced by sirnas. Bars correspond to mean ± SD.**, P <.1 as compared with GFPsiRNA. (c) Ectopic SP1 expression in T47D cells evaluated by western blotting. (d) Schematic diagram shows primer pairs spanning the SP1-binding site and a distal promoter region for ChIP assay. (e-f) ChIP analysis for SP1 binding to FZD1 promoter in MDA-MB-231 cells transfected with or (e) and in T47D cells transfected with -sirnas or RNA (f) using distal primer pairs as control. These data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups. BRL,.
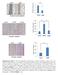
5 Supplementary Figure 5. inhibits activation of canonical Wnt pathway via FZD1. (a-b) qrt-pcr for AXIN2 and TCF-1 expression in MDA-MB-231 cells transfected with pcdna3 vector overexpressing () or negative control () (a) or in T47D cells that were transfected (), or transfected with RNA or two -sirnas (b). Bars correspond to mean ± SD. **, P <.1, ***, P <.1 as compared with -vector or RNA. ##, P <.1, as compared with or RNA. (c-d) Western blotting for total JNK (t-jnk), p-jnk (Thr183/Tyr185) and expression in MDA-MB-231 cells that were transfected as in a (c) and in T47D cells that were treated as in b (d). (e-f) qrt-pcr for AXIN2 and TCF-1 expression in MDA-MB-231 cells (e) and in T47D cells (f) treated as indicated. Bars correspond to mean ± SD. **, P <.1 as compared with p- or -sirna. ##, P <.1 as compared with or -sirna. (g) Immunofluorescent saining for β-catenin in MDA-MB-231 cells treated as in e. Scale bar corresponds to 15 µm. These data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups. BRL,. Supplementary Figure 5 a normalized mrna 1.5 Axin2 TCF-1 p-jnk (Thr183/Tyr185) g t-jnk ** ## b normalized mrna e Axin2 TCF normalized mrna2.5 p-jnk (Thr183/Tyr185) ## ## t-jnk *** *** Axin2 TCF-1 FZD1 c ** ## FZD1 +FZD1 f normalized mrna +FZD Axin2 TCF-1 ** ## **## **##** ## FZD-si1 FZD-si2 FZD-si1 FZD-si2 Merge DAPI β-catenin
6 Supplementary Figure 6 a b mrna expression/gapdh mrna expression/gapdh WNT1 WNT2 WNT1 ** WNT2B WNT3 ** * WNT2 WNT2B WNT3 WNT3A WNT4 WNT5A ** ** WNT3A WNT4 WNT5A WNT5B WNT6 WNT7A WNT7B WNT8A WNT8B WNT9A WNT9B ** WNT5B WNT6 WNT7A WNT7B WNT8A WNT8B WNT9A WNT9B WNT1A * * WNT1A WNT1B WNT11 WNT16 ** WNT1B WNT11 WNT16 Supplementary Figure 6. Screening Wnt ligands responsible for the activation of Wnt pathway via -FZD1. (a-b) qrt-pcr for 19 Wnt family members in MDA-MB-231 cells transfected with pcdna3 vector overexpressing () or negative control () (a) and in T47D cells that were transfected (), or transfected with GFPsiRNA or either of the two -sirnas (b). Bars correspond to mean ± SD. *, P <.5 and **, P <.1 as compared with or RNA. These data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups. BRL,.
7 Supplementary Figure 7 a normalized mrna AXIN2 TCF-1 **## **## WNT3-si1 WNT3-si2 b normalized mrna WNT3-si1 WNT3-si2 WNT3-si1 WNT3-si2 d e f AXIN2 AXIN TCF normalized mrna g normalized mrna WNT3-si1 AXIN2 WNT3-si2 WNT3-si1 WNT3-si2 WNT1B-si1 WNT1B-si normalized mrna h normalized mrna AXIN2 TCF AXIN2 TCF-1 ** ** ** ## ## ## **## WNT3-si1 WNT3-si2 normalized mrna WNT1B-si1 WNT1B-si2 c normalized mrna WNT1B-si1 WNT1B-si AXIN2 AXIN2 TCF-1 WNT3-si1 WNT3-si2 WNT3-si1 WNT3-si2 WNT3-si1 WNT3-si2 Supplementary Figure 7. WNT3 affects the activation of canonical Wnt pathway following silencing. (a-b) qrt-pcr for AXIN2 and TCF-1 expression in MDA-MB-231 cells that were transfected, or transfected with RNA or two WNT3-siRNAs (a) and in T47D cells that were transfected (), transfected with RNA or either of the two -sirnas, or co-transfected with WNT3-siRNAs and -sirnas (b). Bars correspond to mean ± SD. **, P <.1 and ##, P <.1 as compared with RNA or -sirnas. (c-h) qrt-pcr for AXIN2 and TCF-1 expression in MDA-MB-231cells (c, e and g) or in T47D cells (d, f and h) treated as indicated.these data are representative of three independent experiments. Student s t-test was used for the comparison of two independent groups.brl,.
8 Supplementary Figure 8 a b c Relative Lucerfase Activity MCF-1A T47D BT-474 MCF-7 MDA-MB-436 MDA-MB-231 normalized mrna lin4 T47D 16b lin4 16b ASO MDA-MB-231 T47D lin4 16b MDA-MB-231 lin4 16b ASO Supplementary Figure 8. mir-16b post-transcriptionally inhibits expression. (a) Luciferase activity was measured in different breast cell lines that were transfected with PGL3 plasmid containing a 25bp DNA fragment upstream of TSS. (b-c) qrt- PCR (b) and western blotting (c) for in T47D cells that were transfected (), or transfected with lin4 mimics (lin4) or mir-16b mimics (16b) and in MDA-MB-231 cells that were transfected (), or transfected with lin4 mimics (lin4) or mir-16b ASO (16b ASO). Bars correspond to mean ± SD. ***, P <.1, ###, P <.1 as compared with lin4.these data are representative of three independent experiments. Student s t- test was used for the comparison of two independent groups.
9 Supplementary Figure 9 a tumor volum (mm 3 ) c positive cells (%) d FZD1(%) BT-474 GFP-sh BRL-sh1 BRL-sh days after implanation BT-474 FZD1 GFP-sh *** *** *** BRL-sh1 BRL-sh2 BT-474 *** R= -.96 P <.1 b tumor volum (mm 3 ) FZD1(%) positive cells (%) MDA-MB days after implanation FZD1 MDA-MB-231 *** MDA-MB-231 *** R= -.97 P < (%) (%) Supplementary Figure 9. did not affect the primary tumor growth rate in tumor xenografts. (a-b)tumor volume of breast cancer xenografts in the nude mice inoculated with BT-474 that were enforced to stably express two shrnas targeting (-shrnas) or GFP (GFP-shRNA) as negative control (a) or MDA-MB-231 cells stably overxpressing or negative control () (b). BRL,.(c) The positive cells of and FZD1 in BT-474 cells treated as in (a) and MDA-MB-231 cells treated as in (b). ***,P <.1 as compared with GFP-sh or (Student t-test).(d) Correlation between the percentages of + and FZD1 + breast cancer cells.linear regression was used to analyze the correlation.
10 Supplementary Figure 1a Figure 1a MCF-1A T47D BT-474 ZR75-1 MCF-7 MDA-MB-453 BT-549 MDA-MB-436 MDA-MB-231 Figure 2e E-cadherin Vimentin Figure 3c UT FZD1 7 Supplementary Figure 1a. Original blot images of western blotting analysis. Red boxes highlight lanes used in figures.brl,.
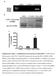
11 Supplementary Figure 1b Figure 3d FZD1 7 Figure 3e E-cadherin UT FZD1-si1 FZD1-si Vimentin 7 Figure 5d β-cat(c) β-cat(n) laminb FZD1 +FZD Figure 5e β-cat(c) β-cat(n) laminb FZD1-si1 FZD1-si2 FZD1-si1 FZD1-si2 FZD1-si1 FZD1-si2 FZD1-si1 FZD1-si Supplementary Figure 1b. Original blot images of western blotting analysis. Red boxes highlight lanes used in figures. β-cat, β-catenin. BRL,.
12 Supplementary Figure 1c Figure 5g β-catenin(c) WNT3-si1 WNT3-si2 1 7 β-catenin(n) 1 7 laminb Figure 5h β-catenin(c) WNT3-si1 WNT3-si2 WNT3-si1 WNT3-si β-catenin(n) 1 7 laminb 7 Supplementary Figure 1c. Original blot images of western blotting analysis. Red boxes highlight lanes used in figures.brl,.
13 Supplementary Figure 1d Figure 6d β-catenin(c) β-catenin(n) laminb lin4 16b 16b +BRL-MUT Figure 6f 1 7 E-cadherin Vimentin lin4 16b 16b +BRL-MUT Figure 6g E-cadherin PBS _ TGF-β Vimentin 7 Supplementary Figure 1d. Original blot images of western blotting analysis. Red boxes highlight lanes used in figures.
14 Supplementary Figure 1e Figure s1a Figure s1b Figure s2f FZD1 Figure s5c MCF-1A p-jnk (Thr183/Tyr185) t-jnk T47D MCF-7 BT-474 MDA-MB-436 MDA-MB Figure s4c SP1 Figure s5d p-jnk (Thr183/Tyr185) t-jnk SP1 1 7 Figure s8c T47D lin4 16b 16b ASO MDA-MB-231 lin4 Supplementary Figure 1e. Original blot images of western blotting analysis. Red boxes highlight lanes used in figures.brl,.
15 Supplementary Table 1. Correlation of Expression with Clinicopathological Status in 428 Cases of Patients with Invasive Breast Cancer IRS>4 IRS 4 P value Age(years) > Size(cm) > Histological grade.19 I II III Stage.21 I II III-IV Positive Lymph node < > ER.287 positive negative 61 HER2.238 negative positive Metastasis <.1 no yes IRS, immunoreactive score.
16 Supplementary Table 2. Multivariate Cox proportional hazard analysis of prognostic variables in 428 patients with breast cancer. Variable Wald(x 2 ) P value Hazard ratio 95.% CI LNM 62.3 < Size < stage < HER HR Age Grade LNM, lymph node metastasis; HER2, human epidermalgrowth factor receptor-2.
17 Supplementary Table 3. Expression of, FZD1, downstream targets of Wnt signal pathway and EMT-markers in normal breast tissue and breast cancer in Oncomine online database. Downregulation of in invasive breast cancer Fold Change P-value Data base (case Invasive ductal breast carcinoma vs normal -1.6 <.1 TCGA breast(593) Invasive ductal breast carcinoma vs normal Zhao breast(64) Invasive ductal breast carcinoma vs normal <.1 Curtis breast(2136) Correlation of and good prognosis in breast cancer Fold Change P-value Data base (case Metastasis at 5 years vs no metastasis at Loi breast(87) years recurrence at 5 years vs no recurrence at Loi breast(87) years Metastasis at 3 years vs no metastasis at Loi breast(87) years Recurrence at 3 years vs no recurrence at Loi breast(87) years Recurrence at 5 years vs no recurrence at Loi breast 3(77) years Metastasis at 5 years vs no metastasis at Loi breast 3(77) years Upregulation of FZD1 in triple negative breast cancer Fold Change P-value Data base (case Triple negative vs other Bonnefoi breast(16) Triple negative vs other Tabchy breast(178) Upregulation of CTNNB1 in invasive breast cancer Fold Change P-value Data base (case Invasive breast carcinoma vs normal 2.33 <.1 Gluck breast(158) Ductal breast carcinoma insitu vs normal Curtis breast(2136) Invasive breast carcinoma vs normal Curtis breast(2136) Invasive ductal and lobular carcinoma vs TCGA(593) normal Upregulation of CTNNB1 in triple negative breast cancer Fold Change P-value Data base (case Triple negative vs other <.1 Curtis breast (2136) Triple negative vs other TCGA(593) Triple negative vs other Zhao breast(64) Triple negative vs other Bonnefoi breast(16) Downregulation of CDH1 in invasive breast cancer Fold Change P-value Data base (case Iinvasive ductal breast carcinoma vs normal Zhao breast(64) Ductal breast carcinoma vs normal <.1 Richardson breast 2 (47) Downregulation of CDH1 in triple negative breast cancer Fold Change P-value Data base (case Triple negative vs other Bittner breast(336) Triple negative vs other <.1 TCGA breast(593) Triple negative vs other <.1 Curtis breast(2136) Triple negative vs other Kao breast(327)
18 Upregulation of VIM in triple negative breast cancer Fold Change P-value Data base (case Triple negative vs other Curtis breast(2136) Triple negative vs other <.1 Tabchy breast(178) Triple negative vs other <.1 Bittner breast(336) Triple negative vs other Esserman breast(13) Upregulation of VIM in breast cancer with positive lymph nodes Fold Change P-value Data base (case N1+ vs N Zhao breast(64) N1+ vs N Weigelt breast(16) N1+ vs No Yu breast 3(96) N1+ vs N Loi breast(87) Upregulation of AXIN2 in breast cancer with positive lymph nodes Fold Change P-value Data base (case N1+ vs N Lu breast(129) N1+ vs N Korde breast(61) N1+ vs N TCGA(593) Upregulation of TCF-1 in triple negative breast cancer Fold Change P-value Data base (case Triple negative vs other Bild breast(158) Triple negative vs other 1.8 <.1 TCGA(593) Triple negative vs other Chin breast(118) Triple negative vs other Curtis breast(2136) Upregulation of FZD1 in high stage breast cancer Fold Change P-value Data base (case Stage IV vs stage I, II and III.15 Chin Breast (118) Grade 3 vs grade Bonnefoi breast (16) Stage III vs stage,1 and 2.11 Curtis breast (2136) Correlation of CTNNB1 and poor Fold P-value Data base (case prognosis in breast cancer Change Dead at 5 years vs no Curtis breast (2136) Dead at 3 years vs no Curtis breast (2136) M1+ vs M Sorlie breast 2 (167) Recurrence at 5 years vs no Sorlie breast 2 (167) Dead at 5 years vs no Sorlie breast 2 (167) Recurrence at 5 years vs no Sorlie breast (85) Dead at 3 years vs no Sorlie breast (85) Metastatic event at 1 year vs no Bos breast(24) Correlation of AXIN2 and poor prognosis Fold P-value Data base (case in breast cancer Change Dead at 5 years vs no Esserman breast (13) Metastatic event at 5 years vs no Loi breast (87) Recurrence at 5 years vs no Loi breast (87) Metastatic event at 5 years vs no Loi breast 3 (77) Recurrence at 5 years vs no Loi breast 3 (77) Correlation of TCF-1 and poor prognosis Fold P-value Data base (case in breast cancer Change Dead at 1 year vs no Curtis breast (2136) Dead at 5 years vs no Curtis breast (2136)
19 Metastatic event at 3 years vs no Curtis breast (2136) Metastasis vs primary site Sorlie breast2 (167) Metastatic event at 3 years vs no 1.86 <.1 Hatzis breast (58) Metastatic event at 5 years vs no <.1 Hatzis breast (58) Correlation of CDH1 and good prognosis Fold P-value Data base (case in breast cancer Change Metastatic event at 5 years vs no <.1 Hatzis breast (58) Metastatic event at 3 years vs no <.1 Hatzis breast (58) Metastatic event at 1 year vs no Hatzis breast (58) Metastatic event at 1 year vs no Desmedt breast (198) Dead at 5 years vs no Curtis breast (2136) Correlation of VIM and poor prognosis Fold P-value Data base (case in breast cancer Change Recurrence at 1 year vs no <.1 Wang breast (286) Metastatic event at 5 years vs no <.1 Wang breast (286) Metastatic event at 3 years vs no 1.29 <.1 Wang breast (286) Dead at 1 years vs no Curtis breast (2136) Dead at 5 years vs no 1.11 <.1 Curtis breast (2136) Dead at 3 years vs no 1.9 <.1 Curtis breast (2136) Metastatic event at 3 years vs no Minn breast 2 (121) Metastatic event at 5 years vs no Minn breast 2 (121) Recurrence at 3 years vs no Sorlie breast 2 (167) Metastatic event at 1 year vs no Desmedt breast (198) Recurrence at 5 years vs no Desmedt breast (198) Recurrence at 5years vs no Ma breast 2 () Metastatic event at 1 year vs no vandevijver breast (295) Dead at 3 years vs no vandevijver breast (295) Dead at 5 years vs no vandevijver breast (295) Metastatic event at 5 years vs no Kao breast (327) Metastatic event at 3 years vs no Kao breast (327) Metastatic event at 3 years vs no <.1 Bos breast (24) Correlation of and FZD1 in breast cancer Correlation P value Data base (case Correlation of and FZD1 in breast cancer Ginestier breast() Correlation of and FZD1 in breast cancer Stickeler breast(57) Correlation of and FZD1 in breast cancer Esserman breast(13) Correlation of and FZD1 in breast cancer Ma breast 4(66) Correlation of and CTNNB1 in breast cancer Correlation P value Data base (case Correlation of and CTNNB1 in breast cancer Esserman breast(13) Correlation of and CTNNB1 in breast cancer <.1 Nikolsky breast(191)
20 Correlation of and CTNNB1 in breast cancer TCGA breast(593) Correlation of and AXIN2 in breast cancer Correlation P value Data base (case Correlation of and AXIN2 in breast cancer <.1 Pawitan breast(159) Correlation of and AXIN2 in breast cancer -.52 <.1 Gluck breast(158) Correlation of and AXIN2 in breast cancer <.1 Miyake breast(115) Correlation of and TCF-1 in breast cancer Correlation P value Data base (case Correlation of and TCF-1 in breast cancer <.1 Gluck breast (158) Correlation of and TCF-1 in breast cancer Loi breast 3 (77) Correlation of and CDH1 in breast cancer Correlation P value Data base (case Correlation of and CDH1 in breast cancer.6.7 Curtis breast 2(1992) Correlation of and CDH1 in breast cancer.66.3 Curtis breast (2136) Correlation of and CDH1 in breast cancer Ma breast 4 (66) Correlation of and VIM in breast cancer Correlation P value Data base (case Correlation of and VIM in breast cancer -.26 <.1 Zhang breast (313) Correlation of and VIM in breast cancer Ma breast 4 (66)
21 Supplementary Table 4. Top 5 up- and downregulated genes after ectopic expression of in MDA-MB-231cells Upregulated genes Downregulated genes Fold change Gene Fold change Gene OR51Q ZFP LOC PKD2L CNTNAP FOXE RSPO PDCD IGLC ALS2CR BICC ZNF LOC RP3-473B DPF LOC ZIC ZNF GALR GRIP C1QL PPP1R9A SLC6A LOC NPFFR LILRB LOC RAD51L MAGEA YLPM OR51D LONRF LRRC SYT FZD GRHL LOC KIAA TREML LOC LOC KIF1B FAM57B FAM26A KLK SUHW LOC USP SCUBE SUB LOC LINS ITGAX FALZ SCUBE TOPORS WDR PLVAP KIR2DL MGC N/A CREM LOC CHIC RP11-41N C9orf ELA3A LOC RAMP KIAA WNT7A KIAA NRAP EXOC EPX GABPB LOC TULP MGC NPM N/A SLC25A LOC FLJ FGF SASS LOC NPM FLJ ANAPC FMN SMAD FFAR SETD KCNT SLC7A LOC C5orf LOC645764
22 Supplementary Table 5. Target scan predicts the mirnas targeting - 3 UTR Preferential conservation mirnas Higher probability* mir-93 mir-2a mir-16a mir-16b mir-17 mir-2b mir-519d (position of 3 UTR) (position of 3 UTR) Lower probability* mir-52a/b/c/d/e mir-32a/b/c/d/e mir-372 (position of 3 UTR) mir-182 (position of 3 UTR) mir-183 (position of 3 UTR) mir-223 (position of 3 UTR) *, sites with higher or lower probability of preferential conservation is defined according to TargetScanHuman 6.2.
23 Supplementary Table 6. Incidence of tumors and liver metastasis from BT-474 and MDA-MB-231 in nude mice. BT-474 (5 1 6 ) MDA-MB-231 (2 1 6 ) Groups Tumors Liver metastasis Groups Tumors Liver metastasis GFP-sh 7/8 1/8 8/8 6/8 sh1 sh2 6/8 4/8 7/8 2/8 7/8 5/8
24 Supplementary Table 7. Primers used for genes and cloning vectors as indicated. Primers for genes FART1-F FART1-R VANGL2-F VANGL2-R BMP8A-F BMP8A-R CAMK2B-F CAMK2B-R WNT1-F WNT1-R WNT2-F WNT2-R WNT2b-F WNT2b-R WNT3-F WNT3-R WNT3A-F WNT3A-R WNT4-F WNT4-R WNT5A-F WNT5A-R WNT5B-F Sequence CTCAGGCCGACCTTGATGG CCTGTGCGGGTTTCTGGTT ACCGCTCTAAGAGTCGAGATG GTTACTACTGTCGTCGTTTCCC ACTCTCGGTGCCTACTTCCT GCAGAGTGACAGCCACAAAC GCACACCAGGCTACCTGTC GGACGGGAAGTCATAGGCA CGATGGTGGGGTATTGTGAAC CCGGATTTTGGCGTATCAGAC CCGAGGTCAACTCTTCATGGT CCTGGCACATTATCGCACAT TGGCGTGCACTCTCAGATTT GACAAGATCAGTCCGGGTGG CCATTTGCGGCTGTGACTC TGGACACTAACACGCCGAAG CAAGATTGGCATCCAGGAGT ATGAGCGTGTCACTGCAAAG ATGGAAGTCACACCCTCTGG CCTGGAAGGACCCACAGATA GGGTGGGAACCAAGAAAAAT TGGAACCTACCCATCCCATA GCGAGAAGACTGGAATCAGG
25 WNT5B-R AACATCTCGGGTCTCTGCAC WNT6-F GTCACGCAGGCCTGTTCTAT WNT6-R CGTCCATAAAGAGCCTCGAC WNT7A-F CTGTGGCTGCGACAAAGAGAA WNT7A-R GCCGTGGCACTTACATTCC WNT7B-F GTTACGGCATCGACTTCTCC WNT7B-R CCGGTCCTCTAGAACCTTCC WNT8A-F GAACCTGTTTATGCTCTGGGC WNT8A-R CAGCGTTCCCAAGCAAACTG WNT8B-F CCGACACCTTTCGCTCCATC WNT8B-R CAGCCCTAGCGTTTTGTTCTC WNT9A-F GCAAGCATCTGAAGCACAAG WNT9A-R TGCTCTCGCAGTTCTTCTCA WNT9B-F TGTGCGGTGACAACCTCAAG WNT9B-R ACAGGAGCCTGATACGCCAT WNT1A-F GGTCAGCACCCAATGACATTC WNT1A-R TGGATGGCGATCTGGATGC WNT1B-F TCTGACAAGGGGACAGAACC WNT1B-R TCATTGCTTAGAGCCCGACT WNT11-F CAGGCAGTGCAACAAGACAT WNT11-R TGAGGGTCCTTGAGCAGAGT WNT16-F AAATGCGCAGGAGAGAAAAA WNT16-R ACCCTCTGATGTACGGTTGC AXIN2-F AGTGTGAGGTCCACGGAAAC AXIN2-R CTGGTGCAAAGACATAGCCA
26 TCF-1-F AGGAGATGAGAGCCAAGGTCA TCF-1-R AGCCTGGGTATAGCTGCATGT GAPDH-F GACTCATGACCACAGTCCATGC GAPDH-R AGAGGCAGGGATGATGTTCTG -F AGTGAAAACGGAACCACCTG -R CCATCAGGCCTCTTAAACCA HPRT-F TTCCTTGGTCAGGCAGTATAATCC HPRT-R AGTCTGGCTTATATCCAACACTTCG 18S rrna-f CGGCTACCACATCCAAGGAA 18S rrna-r GCTGGAATTACCGCGGCT FZD1-F TCAACTACCACTTCCTGGGG FZD1-R CAGCACAGCACTGACCAAAT FZD2-F AGTTCGGTTTTCAGTGGCCC FZD2-R CTCCGTCCTCGGAGTGGTT FZD3-F CAATGGAGCCATTCCACCCT FZD3-R AGGCCAAGGAACACCAAACA FZD4-F GCCAATGTGCACAGAGAAGA FZD4-R GGTTTTGTGAGGTAAGGGCA FZD5-F TTCTGGATAGGCCTGTGGTC FZD5-R AGGTAGCAGGCTGACAGGAA FZD6-F ATGGCCTACAACATGACGTTT FZD6-R GTTTACGACAAGGTGGAACCA FZD7-F CCAACGGCCTGATGTACTTT FZD7-R GAGAACGGTAAAGAGCGTCG FZD8-F ATCGGCTACAACTACACCTACA
27 FZD8-R FZD9-F FZD9-R FZD1-F FZD1-R SP1-F SP1-R Primers for cloning vectors FZD1-2-Luc-F FZD1-2-Luc-R FZD Luc-F FZD1-984-Luc -F FZD1-4-Luc -F FZD1-25-Luc -F FZD1-173-Luc -F FZD1-78-Luc -F FZD1-2-SP1-Mut-Luc-F GTACATGCTGCACAGGAAGAA TGCCCCTCTCTGGCTACCTG GGGCACCGTGTAGAGGATGG GGCGGTGAAGACCATCCTG CAGCTTGTCCGTGTTCTCG ACCAAGCTGAGCTCCATGAT CCTCAGTGCATTGGGTACTTC Sequence ATGGTACCGTTTCTACCTTCTTTTTCACTTTCT ATCAAGCTTGACCAGCCGGCAGCTGTT ATGGTACCCAAACGGGCAGGGGGCAA ATGGTACC GGGCGCTGGCCGGGCACAT ATGGTACC GACCACCCCCAAACCCCT ATGGTACC GCGCGCCAGGCGGGCTGT ATGGTACC TGCAGGCCGGGGGGGATC ATGGTACC AGCAGCGGCGGGGCGGG AGGGCGAGGAGAAGTGAGTGAGATTGCCGG GGAGGG FZD1-2-SP1-Mut-Luc-R CCCTCCCCGGCAATCTCACTCACTTCTCCTCG CCCT pgl3--f pgl3--r pmir-report Mut-F pmir-report Mut-R pmir-report Mut-F ATGGTACCCCATTGTCATCCTGGCCCGT ATCAAGCTTCGGAACGCCCCAACCCA AAGCTAAAAGGAACGCCTCTGTAGAATAGCA AC GTTGCTATTCTACAGAGGCGTTCCTTTTAGCTT TGGTGGTCTTGTACGCCTCAATTTCTGTTACA AT
28 pmir-report Mut-R pmir-report-- WT-F pmir-report-- WT-R ATTGTAACAGAAATTGAGGCGTACAAGACCA CCA AATGAGCTCTGCCTCTTAGGTCCGATGAC TCCAAGCTTATTTGGCAACTATGAAGTAGTGA TC
29 Supplementary methods Preparation of single-cell suspension of primary breast cancer tissue Before digestion with collagenase, primary human tumors with or without lymph node metastases were cut up into small pieces and then minced completely by using sterile blades. To obtain single cell suspensions, the resulting tumor pieces were then mixed with ultra-pure collagenase III in medium 199 (2 25 units of collagenase per ml) and allowed to incubate at 37 C for 3 4 h. Pipetting with a 1-ml pipette was done every 15 2 min. At the end of the incubation, cells were filtered through a 45- μl nylon mesh and washed with RPMI/2% FBS, then washed twice with HBSS. Cells were counted and then transferred to a 5-ml tube, washed twice with HBSS with 2% heat-inactivated calf serum (HICS; 5 min at 1, rpm), then resuspended in 1 μl (per 1 6 cells) of HBSS with 2% HICS. Antibodies (appropriate dilution per antibody) were then added and incubated for 2 min on ice, and then washed twice with HBSS/2% HICS. When needed, a secondary antibody addition was conducted by resuspending in 1 μl (per 1 6 cells) of HBSS/2% HICS, and then adding 1 4 μl of secondary antibody (depending on the secondary antibody and its concentration), followed by a 2-min incubation. The antibodies used were anti-cd44 [allophycocyanin (APC), phycoerythrin (PE), or biotin], anti-cd24 (PE or FITC), anti-b38.1 (APC), anti-epithelial-specific antigen (ESA) FITC (Biomeda, Foster City, CA), and anti-h2kd, (PharMingen). Lineage marker antibodies were anti-cd2, -CD3 -CD1, -CD16, -CD18, -CD31, -CD64, and - CD1b. Unless noted, antibodies were purchased from PharMingen. Antibodies were directly conjugated to various fluorochromes, depending on the experiment. Cell
30 sorting was performed on single-cell suspensions using an Epics Altra flow cytometer (Beckman Coulter). Plasmid construction and establishment of stable cell lines FZD1, SP1 and Myc-tagged were cloned into pcdna3 vector. was also cloned into retrovirus vector pbabe-puro to generate pbabe-. A DNA fragment between -2 to +5 relative to FZD1 transcription start site and a series of sequential deletions of the 5 -flanking region of FZD1 promoter were cloned into pgl3-bsic luciferase reporter vector to generate pgl- FZD1 (FZD1-2-Luc), FZD Luc, FZD1-984-Luc, FZD1-4-Luc, FZD1-25-Luc, FZD1-173-Luc and FZD1-78-Luc recombinant constructs. To construct the promoter recombinant plasmid, a 215bp DNA fragment between to +56 relative to TSS was cloned into pgl3-basic plasmid to generate pgl3- plasmid. FZD1-2-SP1-Mut-Luc Mutant was obtained using the QuikChange site-directed mutagenesis kit (Stratagene), The primers used to generate the recombinant plasmids were listed in Supplementary Table 7. The Oligonucleotides encoding shrna targeting or GFP were synthesized and cloned into psuper-retro-puro plasmid (OligoEngine, Inc, USA). Retrovirus vectors were generated as described(1) by co-transfecting psuper-retro-
31 Puro carrying the shrna expression cassette or pbabe- with helper plasmid pik into 293T cells for 48 hrs. The viral supernatants were collected, filtered, mixed with fresh complete medium (1:1) and 4 µg/ml of polybrene (Sigma, St Louis, Missouri, USA), and then added to breast cancer cells. The stably infected cells were selected with 1µg/mL of puromycin (Sigma, St Louis, Missouri, USA) for two weeks. To generate T47D and MDA-MB-231 cell lines stably expressing pgl3-fzd1, T47D and MDA-MB-231 cell were co-transfected with pgl3-fzd1 and ppru (Clontech, USA) for 24 hrs and the pgl3-fzd1 positive cells were selected with 1 µg/ml of puromycin for two weeks. qrt-pcr Quantitative real-time reverse transcription PCR (qrt-pcr) was performed with a LightCycler 48 instrument (Roche Diagnostics, Switzerland), using the SYBR Premix Ex Taq TM (TaKaRa, Japan) according to the manufacturer s instruction. All reactions were done in a 25ul reaction volume in triplicate. Primers for, FRAT1, VANGL2, BMP8A, CAMK2B, SP1, Vimentin, Fibonectin, GAPDH and 19 Wnt ligands as well as 1 FZD family members were obtained from Invitrogen. Following an initial denaturation at 95 o C for 3 seconds, cycles of PCR amplification were performed at 95 o C for 5 seconds and 6 o C for 2 seconds. Standard curves were generated and the relative amount of target gene mrna was
32 normalized to GAPDH. The specificity was verified by melting curve analysis and agarose gel electrophoresis. To quantify cancer metastasis in mouse livers, qrt-pcr for human hypoxanthine-guanine-phosphoribosyltransferase (hhprt) was performed on Trizol (Invitrogen, Carlsbad, California) -isolated total RNAs using described primers for hhprt and 18S rrna. Following reverse transcription for 45 mins at 42 o C and Taq activation for 3 mins at 95 o C, cycles of PCR at 95 o C for 12 seconds and 6 o C for 15 seconds was performed. All the primers are illustrated in Supplementary Table 7. Dual-luciferase reporter assay The activity of Wnt signal transduction pathways was measured by dual-luciferase reporter assay using Cignal TCF/LEF Reporter (luc) Kit (CCS-18L, QIAGEN) according to the manufacturer s instructions. For FZD1 promoter activity analysis, a series of FZD1 luciferase reporters were co-transfected with prl-tk (Promega, USA) coding for Renilla luciferase into MDA-MB-231 cells for 24 hrs and the luciferase activity was measured using the Dual Luciferase Assay kit (Promega, USA) according to the manufacturer s protocol. For -mediated FZD1 transcriptional repression, T47D or MDA-MB-231 cells stably expressing pgl3-fzd1 were transfected with sirnas or -expressing vector for 48 hrs and the luciferase activity was measured. For
33 promoter activity analysis, pgl3- was transfected into MCF-1A, T47D, BT474, MCF-7, MDA-MB-436 and MDA-MB-231 cells for 24 hrs and the luciferase activity was measured. The prl-tk plasmid was used as a transfection control for all luciferase reporter assays. The firefly luciferase values were normalized to Renilla, and the ratio of firefly/renilla values is presented. The experiments were performed independently in triplicate. For pmir-reporter luciferase assays, 3 UTR target sequences containing potential target sites for micrornas were cloned into the pmir- REPORTER luciferase open reading frame following the manufacture s instruction to construct the pmir--3 UTR-WT vectors. The pmir--3 UTR- MUT vectors containing mutated target sequences were used as negative control. After transfection of 293T cells with pmir--3 UTR-WT/MUT vectors as well as microrna mimics, the luciferase activity was measured. sirna sequences sirnas sequences are as follows: -sirna1: 5 -GAAGACUGGACAACAAUUATT-3, 5 -UAAUUGUUGUCCAGUCUUCTT-3 ; -sirna2: 5 -GUGCUGUAAUUACAACAAUTT-3, 5 - AUUGUUGUAAUUACAGCACTT-3.
34 FZD1-siRNA1: 5 -GCUACAACAUGACUCGUAUTT-3 5 -AUACGAGUCAUGUUGUAGCTT-3 FZD1-siRNA2: 5 -CGCCUCAACAUGGAUUACUTT-3 5 -AGUAAUCCAUGUUGAGGCGTT-3 SP1-siRNA1: 5 -CCAACAGAUUAUCACAAAUTT-3 5 -AUUUGUGAUAAUCUGUUGGTT-3 SP1-siRNA2: 5 -CCUGGAGUGAUGCCUAAUATT UAUUAGGCAUCACUCCAGGTT-3 Supplementary reference 1. Lee SK, Dykxhoorn DM, Kumar P, Ranjbar S, Song E, Maliszewski LE, et al. Lentiviral delivery of short hairpin RNAs protects CD4 T cells from multiple clades and primary isolates of HIV. Blood. 25;16:
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the
 Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting
 mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
Supplementary Table S1. Tumor samples used for analysis Tumor size (cm) BNG (grade) ERα PR. pn-
 Supplementary Table S1. Tumor samples used for analysis Sample# Age Tumor size (cm) pn- Stage Stage BNG (grade) ERα PR HER2 (FISH) Triple negative T1 46 3 N1a III 2 Pos Neg N T2 58 1 N(i-) I 3 Pos Neg
Supplementary Table S1. Tumor samples used for analysis Sample# Age Tumor size (cm) pn- Stage Stage BNG (grade) ERα PR HER2 (FISH) Triple negative T1 46 3 N1a III 2 Pos Neg N T2 58 1 N(i-) I 3 Pos Neg
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplemental Figure 1
 1 Supplemental Figure 1 Effects of DATE shortening on HGF promoter activity. The HGF promoter region (-1037 to +56) containing wild-type (30As) or truncated DATE (26As, 27As, 28A, 29As) from breast cancer
1 Supplemental Figure 1 Effects of DATE shortening on HGF promoter activity. The HGF promoter region (-1037 to +56) containing wild-type (30As) or truncated DATE (26As, 27As, 28A, 29As) from breast cancer
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplemental Table S1
 Supplemental Table S. Tumorigenicity and metastatic potential of 44SQ cell subpopulations a Tumorigenicity b Average tumor volume (mm ) c Lung metastasis d CD high /4 8. 8/ CD low /4 6./ a Mice were injected
Supplemental Table S. Tumorigenicity and metastatic potential of 44SQ cell subpopulations a Tumorigenicity b Average tumor volume (mm ) c Lung metastasis d CD high /4 8. 8/ CD low /4 6./ a Mice were injected
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
Supplementary Figure (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were
 Supplementary Figure 1. Gd@C 82 (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were treated with PBS, Gd@C 82 (OH) 22, C 60 (OH) 22 or GdCl
Supplementary Figure 1. Gd@C 82 (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were treated with PBS, Gd@C 82 (OH) 22, C 60 (OH) 22 or GdCl
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2
 Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Information and Figure legends
 Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Figure 1. Deletion of Smad3 prevents B16F10 melanoma invasion and metastasis in a mouse s.c. tumor model.
 A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
HIF-inducible mir-191 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment.
 HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
An epithelial-to-mesenchymal transition-inducing potential of. granulocyte macrophage colony-stimulating factor in colon. cancer
 An epithelial-to-mesenchymal transition-inducing potential of granulocyte macrophage colony-stimulating factor in colon cancer Yaqiong Chen, Zhi Zhao, Yu Chen, Zhonglin Lv, Xin Ding, Renxi Wang, He Xiao,
An epithelial-to-mesenchymal transition-inducing potential of granulocyte macrophage colony-stimulating factor in colon cancer Yaqiong Chen, Zhi Zhao, Yu Chen, Zhonglin Lv, Xin Ding, Renxi Wang, He Xiao,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
(A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and a
 Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Table S2. Expression of PRMT7 in clinical breast carcinoma samples
 Table S2. Expression of PRMT7 in clinical breast carcinoma samples (All data were obtained from cancer microarray database Oncomine.) Analysis type* Analysis Class(number sampels) 1 2 3 4 Correlation (up/down)#
Table S2. Expression of PRMT7 in clinical breast carcinoma samples (All data were obtained from cancer microarray database Oncomine.) Analysis type* Analysis Class(number sampels) 1 2 3 4 Correlation (up/down)#
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
mir-132 inhibits lung cancer cell migration and invasion by targeting SOX4
 Original Article inhibits lung cancer cell migration and invasion by targeting SOX4 Yang Li, Lingling Zu, Yuli Wang, Min Wang, Peirui Chen, Qinghua Zhou Tianjin Key Laboratory of Lung Cancer Metastasis
Original Article inhibits lung cancer cell migration and invasion by targeting SOX4 Yang Li, Lingling Zu, Yuli Wang, Min Wang, Peirui Chen, Qinghua Zhou Tianjin Key Laboratory of Lung Cancer Metastasis
Predictive PP1Ca binding region in BIG3 : 1,228 1,232aa (-KAVSF-) HEK293T cells *** *** *** KPL-3C cells - E E2 treatment time (h)
 Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas
 Wang et al. Journal of Hematology & Oncology (2016) 9:90 DOI 10.1186/s13045-016-0323-9 RESEARCH Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas Open Access Xiaofeng
Wang et al. Journal of Hematology & Oncology (2016) 9:90 DOI 10.1186/s13045-016-0323-9 RESEARCH Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas Open Access Xiaofeng
FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643
 Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Supplementary Table S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort. Chr. # of Gene 2. Chr. # of Gene 1
 Supplementary Tale S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort TCGA Case ID Gene-1 Gene-2 Chr. # of Gene 1 Chr. # of Gene 2 Genomic coordiante of Gene 1 at fusion junction Genomic
Supplementary Tale S1. List of PTPRK-RSPO3 gene fusions in TCGA's colon cancer cohort TCGA Case ID Gene-1 Gene-2 Chr. # of Gene 1 Chr. # of Gene 2 Genomic coordiante of Gene 1 at fusion junction Genomic
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the. mutated sequence.
 Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
Posttranscriptional upregulation of HER3 by HER2 mrna induces trastuzumab resistance in breast cancer
 Li et al. Molecular Cancer (2018) 17:113 https://doi.org/10.1186/s12943-018-0862-5 RESEARCH Open Access Posttranscriptional upregulation of HER3 by HER2 mrna induces trastuzumab resistance in breast cancer
Li et al. Molecular Cancer (2018) 17:113 https://doi.org/10.1186/s12943-018-0862-5 RESEARCH Open Access Posttranscriptional upregulation of HER3 by HER2 mrna induces trastuzumab resistance in breast cancer
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or
 Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Impact of hyper-o-glcnacylation on apoptosis and NF-κB activity SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
SUPPLEMENTARY METHODS 3D culture and cell proliferation- MiaPaCa-2 cell culture in 3D was performed as described previously (1). Briefly, 8-well glass chamber slides were evenly coated with 50 µl/well
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
TMA-VARESE COHORT-1 TMA-BERN COHORT-2
 Supplementary Figure 1 TMA-VARESE COHORT-1 TOTAL SAMPLES #5 GLEASON SCORE Number Percentage 6 16 32% = 7 17 34% >7 17 34% TUMOR STAGE T2C 28 56% T3A- 21 42% T3C-T4 1 2% NODE STATUS N 42 84% N1 8 16% PSA
Supplementary Figure 1 TMA-VARESE COHORT-1 TOTAL SAMPLES #5 GLEASON SCORE Number Percentage 6 16 32% = 7 17 34% >7 17 34% TUMOR STAGE T2C 28 56% T3A- 21 42% T3C-T4 1 2% NODE STATUS N 42 84% N1 8 16% PSA
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplemental Data. TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim.
 Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538
 Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
SUPPLEMENTARY INFORMATION
 -. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
-. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma CH Li, SC Chow, DL Yin,
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma CH Li, SC Chow, DL Yin,
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535
 Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the
 Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
Doctoral Degree Program in Marine Biotechnology, College of Marine Sciences, Doctoral Degree Program in Marine Biotechnology, Academia Sinica, Taipei,
 Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Hippo component YAP promotes focal adhesion and tumour aggressiveness via transcriptionally activating THBS1/FAK signalling in breast cancer
 Shen et al. Journal of Experimental & Clinical Cancer Research (2018) 37:175 https://doi.org/10.1186/s13046-018-0850-z RESEARCH Open Access Hippo component YAP promotes focal adhesion and tumour aggressiveness
Shen et al. Journal of Experimental & Clinical Cancer Research (2018) 37:175 https://doi.org/10.1186/s13046-018-0850-z RESEARCH Open Access Hippo component YAP promotes focal adhesion and tumour aggressiveness
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Supplemental Information
 Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary Data. Supplementary Methods:
 Supplementary Data Supplementary Methods: Cell viability assay. Cells were seeded overnight at a density of 2,000 cells per well in 96-well plates in RPMI with 10% FBS and then treated with the relevant
Supplementary Data Supplementary Methods: Cell viability assay. Cells were seeded overnight at a density of 2,000 cells per well in 96-well plates in RPMI with 10% FBS and then treated with the relevant
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
mir-542-3p targets sphingosine-1-phosphate receptor 1 and regulates cell proliferation and invasion of breast cancer cells
 European Review for Medical and Pharmacological Sciences 2017; 21: 108-114 mir-542-3p targets sphingosine-1-phosphate receptor 1 and regulates cell proliferation and invasion of breast cancer cells H.-X.
European Review for Medical and Pharmacological Sciences 2017; 21: 108-114 mir-542-3p targets sphingosine-1-phosphate receptor 1 and regulates cell proliferation and invasion of breast cancer cells H.-X.
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 Overview of the transplant procedure and supplementary data to Figure 1. a. Under isofluorane anesthesia, the lumen of the colon is washed by a gentle PBS enema. b. Using a p200
Supplementary Figure 1 Overview of the transplant procedure and supplementary data to Figure 1. a. Under isofluorane anesthesia, the lumen of the colon is washed by a gentle PBS enema. b. Using a p200
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
The pro-metastasis effect of circanks1b in breast cancer
 Zeng et al. Molecular Cancer (2018) 17:160 https://doi.org/10.1186/s12943-018-0914-x RESEARCH Open Access The pro-metastasis effect of circanks1b in breast cancer Kaixuan Zeng 1,2, Bangshun He 1, Burton
Zeng et al. Molecular Cancer (2018) 17:160 https://doi.org/10.1186/s12943-018-0914-x RESEARCH Open Access The pro-metastasis effect of circanks1b in breast cancer Kaixuan Zeng 1,2, Bangshun He 1, Burton
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
(A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939 (5 µm)
 Supplementary Figure Legends Figure S1. Tankyrase inhibition suppresses cell proliferation in an axin/β-catenin independent manner. (A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939
Supplementary Figure Legends Figure S1. Tankyrase inhibition suppresses cell proliferation in an axin/β-catenin independent manner. (A) SW480, DLD1, RKO and HCT116 cells were treated with DMSO or XAV939
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535
 Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.
 Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplementary Figures. T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ
 Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Supplementary Figures T Cell Factor-1 initiates T helper 2 fate by inducing GATA-3 and repressing Interferon-γ Qing Yu, Archna Sharma, Sun Young Oh, Hyung-Geun Moon, M. Zulfiquer Hossain, Theresa M. Salay,
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
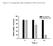 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
The pseudogene TUSC2P promotes TUSC2 function by binding multiple micrornas
 Received 9 Aug 3 Accepted Nov 3 Published 7 Jan DOI:.38/ncomms39 The pseudogene promotes TUSC function by binding multiple micrornas OPEN Zina Jeyapalan Rutnam,, *, William W. Du,, *, Weining Yang, Xiangling
Received 9 Aug 3 Accepted Nov 3 Published 7 Jan DOI:.38/ncomms39 The pseudogene promotes TUSC function by binding multiple micrornas OPEN Zina Jeyapalan Rutnam,, *, William W. Du,, *, Weining Yang, Xiangling
LncRNA TUG1 promoted KIAA1199 expression via mir-600 to accelerate cell metastasis and epithelial-mesenchymal transition in colorectal cancer
 Sun et al. Journal of Experimental & Clinical Cancer Research (2018) 37:106 https://doi.org/10.1186/s13046-018-0771-x RESEARCH Open Access LncRNA TUG1 promoted KIAA1199 expression via mir-600 to accelerate
Sun et al. Journal of Experimental & Clinical Cancer Research (2018) 37:106 https://doi.org/10.1186/s13046-018-0771-x RESEARCH Open Access LncRNA TUG1 promoted KIAA1199 expression via mir-600 to accelerate
Supplemental Experimental Procedures
 Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
ras Multikinase Inhibitor Multikinase Inhibitor 0.1
 a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
a ras ** * ** * ** ** ** ** un in et m lu Se ib SL G 32 W 7 So 50 ra 74 fe ni b LY W 294 or 0 tm 02 R an ap n a in Ev my er cin ol im BE us Z2 3 En P 5 za I13 st 0 au rin D as SP ati 60 nib C 012 is 5
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were
 Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
