Supplementary Data. Reference
|
|
|
- Shawn West
- 5 years ago
- Views:
Transcription
1 Supplementary Data Supplementary Methods Cloning of the mir-155 and mir-221/222 genes Genomic DNA was isolated from hmsc-tert20 cells using the High Pure PCR Template Preparation Kit (Roche Applied Science, Indianapolis, IN), according to the manufacturer s recommendations. The mir-155 and mir-221/222 genes with flanking sequences were amplified using standard PCR and cloning procedures. mir-155 was isolated using primers mir-155_2_f and mir-155_2_r, which generated a 371-bp fragment. The resulting expression vector was designated pmahygro/to/mir-155. mir-221 and mir-222 have the same seed sequence and are separated by less than bp in the genome, suggesting that they are processed from a single transcript. As the expression patterns of mir-221 and mir-222 during differentiation were similar, we cloned both mirna genes and expressed them simultaneously from the same vector. A genomic fragment of 1,299 bp harboring the mir-221 and mir-222 genes were cloned using primers mir-221/222_f_1 and R_1 (Supplementary Table S1). The primers were designed with overhangs for either BamH I or Hind III, but T/A cloning was used. The expression vector with these mirnas was designated pmahygro/to/mir- 221/222. The CMV-driven expression vector pmahygro/to is a modified version of the pcdna5/frt/to vector (Life Technologies, Carlsbad, CA), in which the FRT site and ATG-lacking Hygromycin gene have been removed and replaced with a Hygromycin expression cassette obtained from the psilencer 3.1-H1 Hygro vector (Life Technologies). Briefly, adenosine-tailed PCR products (using standard procedures) were T/A cloned into pmahygro/to. To allow T/A cloning, the vector was first digested with EcoR V (New England Biolabs, Ipswich, MA) and subsequently thymidinetailed using standard procedures. Expression of mature mirna sequences was confirmed by transient transfection of HEK-293 cells with Lipofectamine 2000 (Life Technologies) followed by qpcr using TaqMan MiRNA Assays (Life Technologies) (Supplementary Fig. S1). pmaegfp was used to optimize the transfection conditions. This construct was generated by cloning an enhanced green fluorescent protein expression cassette from padtrack-cmv [1] (Addgene plasmid 16405) into the pgem-t Easy vector (Promega, Madison, WI). Reference 1. He T-C, S Zhou, LT da Costa, J Yu, KW Kinzler and B Vogelstein. (1998). A simplified system for generating recombinant adenoviruses. Proc Natl Acad Sci USA 95: SUPPLEMENTARY FIG. S1. Ectopic expression of mir-155 and mir-222 in HEK-293 cells. The mirna expression contructs pmahygro/to/mir-155 and pmahygro/to/mir were transiently transfected into HEK-293 cells using Lipofectamine 2000 as described by the manufacturer. Expression of mature mirnas at the indicated time points was confirmed by qpcr. pmahygro/to/mir generates both mir-221 and mir-222, illustrated by expression of mir The average of RNU6B and RNU24 was used for normalization. h: hours after transfection.
2 SUPPLEMENTARY FIG. S2. Changes in gene expression patterns of adipocyte markers during differentiation of hmsc- Tert20 cells. hmsc-tert20 cells were differentiated for 21 days, and total RNA was isolated at the indicated time points. The expression of CCAAT/enhancer-binding protein beta, (CEBPB), CCAAT/enhancer-binding protein delta (CEBPD) 1, peroxisome proliferator-activated receptor gamma (PPARG), CCAAT/enhancer-binding protein alpha (CEBPA), complement factor D (CFD) 1, and ADIPOQ 1 was analyzed using qpcr. The average of TATA-binding protein (TBP) 1 and human acidic ribosomal phosphoprotein (RPLPO) 1,orofTBP and glyceraldehyde 3-phosphate dehydrogenase (GAPDH), was used to normalize the data for the SYBR Green or TaqMan Gene Expression Assays, respectively. Representative data from 2 independent experiments are shown. 1 SYBR Green assays. d and h, days and hours after induction of adipogenesis, respectively.
3 SUPPLEMENTARY FIG. S3. Adipocyte differentiation of primary hmscs. Primary hmscs were differentiated for 21 days. (A) Representative images of differentiated and undifferentiated cells at day 21 after induction of adipogenesis are shown at 10 magnification. In addition, the cells were fixed and stained with Oil Red O to demonstrate adipocyte differentiation. (B) The expression levels of CEBPB, PPARG, CEBPA, and ADIPOQ 1 at the indicated time points after induction of adipogenesis were determined using qpcr. The average of TBP 1 and RPLPO 1,orofTBP and GAPDH, was used to normalize the data for the SYBR Green or TaqMan Gene Expression Assays, respectively. 1 SYBR Green assays. d and h, days and hours after induction of adipogenesis, respectively.
4 SUPPLEMENTARY FIG. S4. mirna expression analysis during adipocyte differentiation of hmsc-tert20. An initial microarray-based global profiling revealed that 66 mirnas were expressed in hmsc-tert20 during adipogenesis. Twenty mirnas that were differentially expressed were further investigated (bold). Two more biological replicates of differentiation were performed and the expression levels of the 20 mirnas in all 3 biological experiments were quantified using qpcr. Twelve mirnas showed consistent expression patterns in all 3 biological experiments (italic). Let-7a, mir-21, mir-26a, mir- 30d, mir-34a, and mir-92 were upregulated and mir-18a, mir-146a, mir-155, mir-214, mir-221, and mir-222 showed decreased expression levels during adipocyte differentiation. d and h: days and hours after induction of adipogenesis, respectively.
5 SUPPLEMENTARY FIG. S5. GeneGO network based on the 32 common targets. A network was generated based on the 32 common targets of mir-155 and mir-221/mir-222. Additional proteins were included to establish interactions between the predicted targets, creating a network of up to 50 proteins using GeneGO and the algorithm auto expand. Supplementary Table S1. Overview of Primers Used to Clone mir-155 and mir-221/222 from hmsc-tert20 Genomic DNA Primer Sequence (5 3 ) mir-155_2_f mir-155_2_r mir-221/222_f_1 mir-221/222_r_1 CTGTCACTCCAGCTTTATAACCG GTTTAAGGTTGAACATCCCAGTG TACGATGGATCCTCACAAGGAATCATGTATGCTG TACGATAAGCTTCTCCACTGGTTTATACCTCCTG The primers used to clone mir-221/222 (mir_221/222_f-1 and mir_221/222_r-1) have restriction sites for BamH I and Hind III (bold letters), although T/A cloning was used in the present study.
6 Supplementary Table S2. Overview of Primers Used for SYBR Green Quantitative Real-Time Polymerase Chain Reaction Gene symbol Primer names Sequence (5 3 ) References ADIPOQ ADIPOQ_F ATGGTCCTGTGATGCTTTGA [1] ADIPOQ_R GTTGAGTGCGTATGTTATTTTT CFD CFD_F ATCACCGAGCGCTTGATGTG CFD_R CTTCTTGCGGTTGCCGCAAA CEBPD CEBPD_F ACAGCCTCGCTTGGACGCAGAG [2] CEBPD_R AGTCGATGTAGGCGCTGAAGTC RPLPO RPLPO_F CGCTGCTGAACATGCTCAAC RPLPO_R TCGAACACCTGCTGGATGAC TBP TBP_F GCCCGAAACGCCGAATAT TBP_R CGTGGCTCTCTTATCCTCATGA ADIPOQ, adiponectin; CFD, complement factor D; CEBPD, CCAAT/enhancer-binding protein delta; RPLPO, human acidic ribosomal phosphoprotein PO; TBP, TATA-binding protein. [1]. Heidi SB, PL Staale, S Axel, M Marta, T Liv, AD Christian, S Unni and ER Janne. (2004). Adiponectin and its receptors are expressed in bone-forming cells. Bone 35: [2]. Tang D, GS Sivko and JW DeWille. (2006). Promoter methylation reduces C/EBPo (CEBPD) gene expression in the SUM-52PE human breast cancer cell line and in primary breast tumors. Breast Cancer Res Treat 95: Supplementary Table S3. Overview of TaqMan Gene and mirna Expression Assays Gene symbol Assay ID/part number Target genes CEBPB Hs _s1 PPARG Hs _m1 CEBPA Hs _s1 CDKN1B Hs _m1 SCD Hs _m1 Endogenous controls Human TBP E Human GAPD (GAPDH) E Target genes hsa-mir hsa-mir hsa-mir-30d hsa-mir hsa-mir-146a hsa-mir hsa-mir-20a hsa-mir hsa-mir hsa-mir hsa-mir hsa-let-7a hsa-let-7c hsa-mir-26a hsa-mir-18a hsa-mir-199a hsa-mir hsa-mir-19b hsa-mir-125a hsa-mir Endogenous controls RNU RNU6B CEBPB, CCAAT/enhancer-binding protein beta; PPARG, peroxisome proliferator-activated receptor gamma; CEBPA, CCAAT/ enhancer-binding protein alpha; CDKN1B, cyclin-dependent kinase inhibitor 1B; SCD, stearoyl coenzyme A desaturase 1; GAPDH, glyceraldehyde 3-phosphate dehydrogenase.
7 Supplementary Table S4. Common Predicted Targets for mir-155 and mir-221/222 Gene symbol Gene name Ref. Seq. ID ACVR2B Activin A receptor, type IIB NM_ BRWD1 Bromodomain and WD repeat domain containing 1 NM_ C14orf147 Chromosome 14 open reading frame 147 NM_ CBL Cas-Br-M (murine) ecotropic retroviral transforming sequence NM_ CD47 Homo sapiens CD47 molecule NM_ E2F2 E2F transcription factor 2 NM_ ETS1 v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) NM_ FOS FBJ murine osteosarcoma viral oncogene homolog NM_ GABRA1 Gamma-aminobutyric acid (GABA) A receptor, alpha 1 NM_ H3F3A H3 histone, family 3A NM_ HNRNPA3 heterogeneous nuclear ribonucleoprotein A3 NM_ KCNA1 Potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic NM_ ataxia with myokymia) KIAA1267 KIAA1267 NM_ KPNA1 Karyopherin alpha 1 (importin alpha 5) NM_ KSR1 Kinase suppressor of ras 1 NM_ MIDN Midnolin NM_ MYBL1 v-myb myeloblastosis viral oncogene homolog (avian)-like 1 NM_ MYO10 Myosin X NM_ NFAT5 Nuclear factor of activated T-cells 5, tonicity-responsive NM_ NOVA1 Neuro-oncological ventral antigen 1 NM_ NUFIP2 Nuclear fragile X mental retardation protein interacting protein 2 NM_ PIK3R1 Phosphoinositide 3-kinase regulatory subunit 1 (alpha) NM_ SHANK2 SH3 and multiple ankyrin repeat domains 2 NM_ SOCS1 Suppressor of cytokine signaling 1 NM_ SOX10 SRY (sex determining region Y)-box 10 NM_ SOX11 SRY (sex determining region Y)-box 11 NM_ TCF4 Transcription factor 4 NM_ TRPS1 Trichorhinophalangeal syndrome I NM_ WEE1 WEE1 homolog (S. pombe) NM_ ZMYM2 Zinc finger, MYM-type 2 NM_ ZNF618 Zinc finger protein 618 NM_ AAK1 AP2 associated kinase 1 NM_ The database TargetScan5.1 was used to identify candidate targets for either mir-155 or mir-221/222. This resulted in 281 and 306 conserved putative targets for mir-155 and mir-221/222, respectively (data not shown), and 32 common targets for all 3 mirnas.
8 Supplementary Table S5. GeneGO Analysis of Common Candidate Targets for mir-155 and mir-221/222 Name Network objects P value Top GeneGO Maps Cell differentiation 11/ E-06 Cell cycle and its regulation 7/ E-05 Apoptosis 9/ E-05 Hematopoiesis 5/ E-04 Vascular development (angiogenesis) 6/ E-03 Mitogenic signaling 6/ E-03 Inflammatory response 6/ E-03 Hypoxia response regulation 2/ E-03 Immune system response 7/ E-03 Protein synthesis 4/ E-03 Top GeneGo Pathway Maps Development_EGFR signaling via small GTPases 3/ E-05 Apoptosis and survival_ceramides signaling pathway 3/ E-05 Development_ERBB-family signaling 3/ E-05 Development_Thrombopoietin-regulated cell processes 3/ E-04 Development_GM-CSF signaling 3/ E-04 G-protein signaling_proinsulin C-peptide signaling 3/ E-04 Immune response_ifn gamma signaling pathway 3/ E-04 Immune response_trem1 signaling pathway 3/ E-04 Development_Prolactin receptor signaling 3/ E-04 Development_Gastrin in cell growth and proliferation 3/ E-04 Top GeneGo Process Network Development_Hemopoiesis, Erythropoietin pathway 6/ E-06 Cell cycle_g1-s Growth factor regulation 6/ E-05 Inflammation_IL-6 signaling 5/ E-05 Reproduction_Feeding and Neurohormones signaling 6/ E-05 Reproduction_FSH-beta signaling pathway 5/ E-04 Apoptosis_Anti-Apoptosis mediated by external signals via MAPK and JAK/STAT 5/ E-04 Immune response_phagocytosis 5/ E-04 Translation_Regulation of initiation 4/ E-04 Cell cycle_g1-s Interleukin regulation 4/ E-04 Development_EMT_Regulation of epithelial-to-mesenchymal transition 5/ E-04 Functional analysis of the common targets was performed in GeneGO. An enrichment analysis was performed and the 10 most significant findings for each of the functional categories are presented.
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
SUPPLEMENTARY INFORMATION
 -. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
-. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
U118MG. Supplementary Figure 1 U373MG U118MG 3.5 A CCF-SSTG
 A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
A172 CCF-SSTG1 15 - - - 1 1 1 2 2 3 4 6 7 8 10101112131617192022-1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22 T98G U373MG - - - 1 1 1 2 2 3 4 6 7 8 10 10 11 12 13 16 17 19 20 22-1 1 1 2 2 3 4 6 7
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Figure S1: Heat map based on the relative expression of genes and mirnas in human
 Figure S1: Heat map based on the relative expression of genes and mirnas in human prostate tissue. Columns represent genes and mirnas; rows represent prostate tissue samples (red squares: Tu samples, green
Figure S1: Heat map based on the relative expression of genes and mirnas in human prostate tissue. Columns represent genes and mirnas; rows represent prostate tissue samples (red squares: Tu samples, green
Table S1 Differentially expressed genes showing > 2 fold changes and p <0.01 for 0.01 mm NS-398, 0.1mM ibuprofen and COX-2 RNAi.
 Table S1 Differentially expressed genes showing > 2 fold changes and p
Table S1 Differentially expressed genes showing > 2 fold changes and p
SUPPLEMENTAY FIGURES AND TABLES
 SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Mouse Meda-4 : chromosome 5G bp. EST (547bp) _at. 5 -Meda4 inner race (~1.8Kb)
 Supplemental.Figure1 A: Mouse Meda-4 : chromosome 5G3 19898bp I II III III IV V a b c d 5 RACE outer primer 5 RACE inner primer 5 RACE Adaptor ORF:912bp Meda4 cdna 2846bp Meda4 specific 5 outer primer
Supplemental.Figure1 A: Mouse Meda-4 : chromosome 5G3 19898bp I II III III IV V a b c d 5 RACE outer primer 5 RACE inner primer 5 RACE Adaptor ORF:912bp Meda4 cdna 2846bp Meda4 specific 5 outer primer
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at
 Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
SUPPLEMENTARY DATA. Assessment of cell survival (MTT) Assessment of early apoptosis and cell death
 Overexpression of a functional calcium-sensing receptor dramatically increases osteolytic potential of MDA-MB-231 cells in a mouse model of bone metastasis through epiregulinmediated osteoprotegerin downregulation
Overexpression of a functional calcium-sensing receptor dramatically increases osteolytic potential of MDA-MB-231 cells in a mouse model of bone metastasis through epiregulinmediated osteoprotegerin downregulation
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Validated Mouse Quantitative RT-PCR Genes Gene Gene Bank Accession Number Name
 5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
ONCOGENES. Michael Lea
 ONCOGENES 2011 Michael Lea ONCOGENES - Lecture Outline I. Introduction 2. Identification of oncogenic genes in retroviruses 3. Homologous sequences in transformed and untransformed cells 4. Methods of
ONCOGENES 2011 Michael Lea ONCOGENES - Lecture Outline I. Introduction 2. Identification of oncogenic genes in retroviruses 3. Homologous sequences in transformed and untransformed cells 4. Methods of
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion
 Probe Set ID RefSeq Transcript ID Gene Title Gene Symbol 205980_s_at NM_001017526 /// NM_181334 /// NM_181335 Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion ARHGAP8 /// LOC553158 FC ALK shrna
Probe Set ID RefSeq Transcript ID Gene Title Gene Symbol 205980_s_at NM_001017526 /// NM_181334 /// NM_181335 Rho GTPase activating protein 8 /// PRR5- ARHGAP8 fusion ARHGAP8 /// LOC553158 FC ALK shrna
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array.
 Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Marta Puerto Plasencia. microrna sponges
 Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
Prof. R. V. Skibbens. Cell Cycle, Cell Division and Cancer (Part 2)
 Prof. R. V. Skibbens November 22, 2010 BIOS 10: BioScience in the 21 st Century Cell Cycle, Cell Division and Cancer (Part 2) Directionality - clocks go in only one direction G1 doesn t have replication-inducing
Prof. R. V. Skibbens November 22, 2010 BIOS 10: BioScience in the 21 st Century Cell Cycle, Cell Division and Cancer (Part 2) Directionality - clocks go in only one direction G1 doesn t have replication-inducing
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
SUPPLEMENTARY FIGURES AND TABLES
 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR
 Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) cadl CTTGGGGGCGCGTCT CTGTTCTTTTGTGCCGTTTCG cyl-coenzyme Dehydrogenase, very
Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) cadl CTTGGGGGCGCGTCT CTGTTCTTTTGTGCCGTTTCG cyl-coenzyme Dehydrogenase, very
Supplementary Figure 1: Fn14 is upregulated in the epidermis and dermis of mice
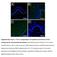 Supplementary Figure 1: Fn14 is upregulated in the epidermis and dermis of mice undergoing AD- and psoriasis-like disease. Immunofluorescence staining for Fn14 (green) and DAPI (blue) in skin of naïve
Supplementary Figure 1: Fn14 is upregulated in the epidermis and dermis of mice undergoing AD- and psoriasis-like disease. Immunofluorescence staining for Fn14 (green) and DAPI (blue) in skin of naïve
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
BL 424 Test pts name Multiple choice have one choice each and are worth 3 points.
 BL 424 Test 3 2010 150 pts name Multiple choice have one choice each and are worth 3 points. 1. The plasma membrane functions as a a. selective barrier to the passage of molecules. b. sensor through which
BL 424 Test 3 2010 150 pts name Multiple choice have one choice each and are worth 3 points. 1. The plasma membrane functions as a a. selective barrier to the passage of molecules. b. sensor through which
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy
 Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Viruses Tomasz Kordula, Ph.D.
 Viruses Tomasz Kordula, Ph.D. Resources: Alberts et al., Molecular Biology of the Cell, pp. 295, 1330, 1431 1433; Lehninger CD Movie A0002201. Learning Objectives: 1. Understand parasitic life cycle of
Viruses Tomasz Kordula, Ph.D. Resources: Alberts et al., Molecular Biology of the Cell, pp. 295, 1330, 1431 1433; Lehninger CD Movie A0002201. Learning Objectives: 1. Understand parasitic life cycle of
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis
 High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis Supplementary Material Supplementary Figure S1. Representative CRBP-1 immunostaining of non-neoplastic
High expression of cellular retinol binding protein-1 in lung adenocarcinoma is associated with poor prognosis Supplementary Material Supplementary Figure S1. Representative CRBP-1 immunostaining of non-neoplastic
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
REGULATION OF MDM2 MEDIATED NFκB2 PATHWAY IN HUMAN LUNG CANCER
 Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2008 REGULATION OF MDM2 MEDIATED NFκB2 PATHWAY IN HUMAN LUNG CANCER Lathika Mohanraj Virginia Commonwealth
Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2008 REGULATION OF MDM2 MEDIATED NFκB2 PATHWAY IN HUMAN LUNG CANCER Lathika Mohanraj Virginia Commonwealth
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Supplementary information for: Community detection for networks with. unipartite and bipartite structure. Chang Chang 1, 2, Chao Tang 2
 Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS
 CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
ulcer healing role 118 Bicarbonate, prostaglandins in duodenal cytoprotection 235, 236
 Subject Index Actin cellular forms 48, 49 epidermal growth factor, cytoskeletal change induction in mucosal repair 22, 23 wound repair 64, 65 polyamine effects on cytoskeleton 49 51 S-Adenosylmethionine
Subject Index Actin cellular forms 48, 49 epidermal growth factor, cytoskeletal change induction in mucosal repair 22, 23 wound repair 64, 65 polyamine effects on cytoskeleton 49 51 S-Adenosylmethionine
microrna Presented for: Presented by: Date:
 microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Herpesviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003)
 MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
VIRUSES. Biology Applications Control. David R. Harper. Garland Science Taylor & Francis Group NEW YORK AND LONDON
 VIRUSES Biology Applications Control David R. Harper GS Garland Science Taylor & Francis Group NEW YORK AND LONDON vii Chapter 1 Virus Structure and 2.2 VIRUS MORPHOLOGY 26 Infection 1 2.3 VIRAL CLASSIFICATION
VIRUSES Biology Applications Control David R. Harper GS Garland Science Taylor & Francis Group NEW YORK AND LONDON vii Chapter 1 Virus Structure and 2.2 VIRUS MORPHOLOGY 26 Infection 1 2.3 VIRAL CLASSIFICATION
Fig. S1. Validation of ChIP-seq binding sites by single gene ChIP-PCR Fig. S2. Transactivation potential of PPAR
 Fig. S1. Validation of ChIP-seq binding sites by single gene ChIP-PCR ChIP-PCR was performed on PPARγ and RXR-enriched chromatin harvested during adipocyte differentiation at day and day 6 as described
Fig. S1. Validation of ChIP-seq binding sites by single gene ChIP-PCR ChIP-PCR was performed on PPARγ and RXR-enriched chromatin harvested during adipocyte differentiation at day and day 6 as described
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α
 Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α Mahnaz Janghorban, PhD Dr. Rosalie Sears lab 2/8/2015 Zanjan University Content 1. Background (keywords: c-myc, PP2A,
Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α Mahnaz Janghorban, PhD Dr. Rosalie Sears lab 2/8/2015 Zanjan University Content 1. Background (keywords: c-myc, PP2A,
RALYL Hypermethylation: A Potential Diagnostic Marker of Esophageal Squamous Cell Carcinoma (ESCC) Junwei Liu, MD
 RALYL Hypermethylation: A Potential Diagnostic Marker of Esophageal Squamous Cell Carcinoma (ESCC) Junwei Liu, MD Aurora Healthcare, Milwaukee, WI INTRODUCTION v Epigenetic aberration and genetic alteration
RALYL Hypermethylation: A Potential Diagnostic Marker of Esophageal Squamous Cell Carcinoma (ESCC) Junwei Liu, MD Aurora Healthcare, Milwaukee, WI INTRODUCTION v Epigenetic aberration and genetic alteration
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature7221 Brown fat selective genes 12 1 Control Q-RT-PCR (% of Control) 8 6 4 2 Ntrk3 Cox7a1 Cox8b Cox5b ATPase b2 ATPase f1a1 Sirt3 ERRα Elovl3/Cig3 PPARα Zic1 Supplementary Figure S1. stimulates
doi: 1.138/nature7221 Brown fat selective genes 12 1 Control Q-RT-PCR (% of Control) 8 6 4 2 Ntrk3 Cox7a1 Cox8b Cox5b ATPase b2 ATPase f1a1 Sirt3 ERRα Elovl3/Cig3 PPARα Zic1 Supplementary Figure S1. stimulates
A bioinformatics driven cancer study
 The 11th China-Japan-Korea Bioinformatics Training Course & Symposium, June 16-18, Suzhou, China A bioinformatics driven cancer study Chao Li and Yixue Li yxli@sibs.ac.cn Shanghai Institutes for Bio-Medicine
The 11th China-Japan-Korea Bioinformatics Training Course & Symposium, June 16-18, Suzhou, China A bioinformatics driven cancer study Chao Li and Yixue Li yxli@sibs.ac.cn Shanghai Institutes for Bio-Medicine
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Non-Size Indications for Aortic Replacement
 Blood Test for Thoracic Aortic Aneurysm 2 nd International Meeting on Aortic Diseases September 30-October 2, 2010 Liege, Belgium John A. Elefteriades, MD William W.L. Glenn Professor of Cardiothoracic
Blood Test for Thoracic Aortic Aneurysm 2 nd International Meeting on Aortic Diseases September 30-October 2, 2010 Liege, Belgium John A. Elefteriades, MD William W.L. Glenn Professor of Cardiothoracic
Supplemental Information
 Supplemental Information Screening of strong constitutive promoters in the S. albus transcriptome via RNA-seq The total RNA of S. albus J1074 was isolated after 24 hrs and 72 hrs of cultivation at 30 C
Supplemental Information Screening of strong constitutive promoters in the S. albus transcriptome via RNA-seq The total RNA of S. albus J1074 was isolated after 24 hrs and 72 hrs of cultivation at 30 C
DNA context and promoter activity affect gene expression in lentiviral vectors
 ACTA BIOMED 2008; 79: 192-196 Mattioli 1885 O R I G I N A L A R T I C L E DNA context and promoter activity affect gene expression in lentiviral vectors Gensheng Mao 1, Francesco Marotta 2, Jia Yu 3, Liang
ACTA BIOMED 2008; 79: 192-196 Mattioli 1885 O R I G I N A L A R T I C L E DNA context and promoter activity affect gene expression in lentiviral vectors Gensheng Mao 1, Francesco Marotta 2, Jia Yu 3, Liang
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Supplemental Table 1 Age and gender-specific cut-points used for MHO.
 Supplemental Table 1 Age and gender-specific cut-points used for MHO. Age SBP (mmhg) DBP (mmhg) HDL-C (mmol/l) TG (mmol/l) FG (mmol/l) Boys 6-11 90th * 90th * 1.03 1.24 5.6 12 121 76 1.13 1.44 5.6 13 123
Supplemental Table 1 Age and gender-specific cut-points used for MHO. Age SBP (mmhg) DBP (mmhg) HDL-C (mmol/l) TG (mmol/l) FG (mmol/l) Boys 6-11 90th * 90th * 1.03 1.24 5.6 12 121 76 1.13 1.44 5.6 13 123
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
NOVEL FUNCTION OF mirnas IN REGULATING GENE EXPRESSION. Ana M. Martinez
 NOVEL FUNCTION OF mirnas IN REGULATING GENE EXPRESSION Ana M. Martinez Switching from Repression to Activation: MicroRNAs can Up-Regulate Translation. Shoba Vasudevan, Yingchun Tong, Joan A. Steitz AU-rich
NOVEL FUNCTION OF mirnas IN REGULATING GENE EXPRESSION Ana M. Martinez Switching from Repression to Activation: MicroRNAs can Up-Regulate Translation. Shoba Vasudevan, Yingchun Tong, Joan A. Steitz AU-rich
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Genome of Hepatitis B Virus. VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department
 Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
VIRUSES AND CANCER Michael Lea
 VIRUSES AND CANCER 2010 Michael Lea VIRAL ONCOLOGY - LECTURE OUTLINE 1. Historical Review 2. Viruses Associated with Cancer 3. RNA Tumor Viruses 4. DNA Tumor Viruses HISTORICAL REVIEW Historical Review
VIRUSES AND CANCER 2010 Michael Lea VIRAL ONCOLOGY - LECTURE OUTLINE 1. Historical Review 2. Viruses Associated with Cancer 3. RNA Tumor Viruses 4. DNA Tumor Viruses HISTORICAL REVIEW Historical Review
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/4/199/ra75/dc1 Supplementary Materials for Signaling by the Matrix Proteoglycan Decorin Controls Inflammation and Cancer Through PDCD4 and MicroRNA-21 Rosetta
www.sciencesignaling.org/cgi/content/full/4/199/ra75/dc1 Supplementary Materials for Signaling by the Matrix Proteoglycan Decorin Controls Inflammation and Cancer Through PDCD4 and MicroRNA-21 Rosetta
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts
 Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Oncolytic virus strategy
 Oncolytic viruses Oncolytic virus strategy normal tumor NO replication replication survival lysis Oncolytic virus strategy Mechanisms of tumor selectivity of several, some of them naturally, oncolytic
Oncolytic viruses Oncolytic virus strategy normal tumor NO replication replication survival lysis Oncolytic virus strategy Mechanisms of tumor selectivity of several, some of them naturally, oncolytic
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Aberrant Promoter CpG Methylation is a Mechanism for Lack of Hypoxic Induction of
 Aberrant Promoter CpG Methylation is a Mechanism for Lack of Hypoxic Induction of PHD3 in a Diverse Set of Malignant Cells Abstract The prolyl-hydroxylase domain family of enzymes (PHD1-3) plays an important
Aberrant Promoter CpG Methylation is a Mechanism for Lack of Hypoxic Induction of PHD3 in a Diverse Set of Malignant Cells Abstract The prolyl-hydroxylase domain family of enzymes (PHD1-3) plays an important
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Hepatitis B virus X gene in the development of hepatocellular carcinoma. Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p.
 Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
RNA interference (RNAi)
 RN interference (RNi) Natasha aplen ene Silencing Section Office of Science and Technology Partnerships Office of the Director enter for ancer Research National ancer Institute ncaplen@mail.nih.gov Plants
RN interference (RNi) Natasha aplen ene Silencing Section Office of Science and Technology Partnerships Office of the Director enter for ancer Research National ancer Institute ncaplen@mail.nih.gov Plants
Certificate of Analysis
 Certificate of Analysis Catalog No. Amount Lot Number 631987 10 μg Specified on product label. Product Information plvx-ef1α-ires-mcherry is a bicistronic lentiviral expression vector that can be used
Certificate of Analysis Catalog No. Amount Lot Number 631987 10 μg Specified on product label. Product Information plvx-ef1α-ires-mcherry is a bicistronic lentiviral expression vector that can be used
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Bhatnagar et al, 2010 Cell Death and Disease Manuscript # CDDIS T
 Bhatnagar et al, Cell Death and Disease Manuscript # CDDIS--98-T Supplemental Materials. Supplemental Figure Legends Supplemental Figure (A) WPE-NA and WPE-NB6 cells were treated with 4 nm of Docetaxel
Bhatnagar et al, Cell Death and Disease Manuscript # CDDIS--98-T Supplemental Materials. Supplemental Figure Legends Supplemental Figure (A) WPE-NA and WPE-NB6 cells were treated with 4 nm of Docetaxel
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney
 BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
Probe. Hind III Q,!?R'!! /0!!!!D1"?R'! vector. Homologous recombination
 Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha
 Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha system. The dcas9-vpr effector is fused to the C-terminus
Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha system. The dcas9-vpr effector is fused to the C-terminus
