Supporting Information
|
|
|
- Alexia Barton
- 5 years ago
- Views:
Transcription
1 Supporting Information Gack et al /pnas SI Text Cell Culture, Transfection, and Reagents. HEK293T, MEF, Vero, EcoPack2-293 (D iosciences), HeLa, HCT116, Huh7, NHLF, 2fTGH wild-type, U3A, and U6A cells (provided by George Stark, Cleveland Clinic Foundation, Cleveland, OH) were cultured in Dulbecco s Modified Eagle s Medium (DMEM) supplemented with 1% FS, 2 mm l-glutamine, and 1% penicillinstreptomycin (GICO RL). LnCap human prostate cancer cells were cultured in RPMI medium164 supplemented with 1% FS, 2 mm l-glutamine, and 1% penicillin-streptomycin. Lymphatic endothelial cells (LECs) were cultured in endothelial growth medium (EGM-2) supplemented with the microvascular supplement pack (Clonetics). Transient transfections were performed with Lipofectamine (Invitrogen), FuGENE 6 (Roche) or calcium phosphate (Clontech) according to the manufacturer s instructions. RIG-I and knockout MEFs were immortalized as described in ref. 1. To obtain stable RIG-I MEF cells, pabe-puro vector, pabe-puro-rig-i, pabe- Puro-RIG-I T 55 I, or pabe-puro-rig-i retrovirus was produced in Ecopack After retroviral infection of RIG-I knockout MEFs, cells were selected by using 1 g/ml puromycin. Plasmid Construction. All constructs for transient and stable expression in mammalian cells were derived from peg GST fusion vector, pef-ires-puro, or pabe-puro expression vector. DNA fragments corresponding to the coding sequence of the RIG-I and TRIM genes were amplified from template DNA by PCR and subcloned into plasmid peg between KpnI and NotI, into pef-ires-puro between AflII and NotI or into pabe-puro between amhi and SalI restriction sites. V5-, Flag-, or Myc-tagged TRIM and RIG-I constructs were expressed from a modified pires-puro encoding a C-terminal V5, Flag, or Myc tag, respectively. RIG-I mutants were generated by PCR using site-directed mutagenesis or overlapping PCR. All constructs were sequenced using an AI PRISM 377 automatic DNA sequencer to verify % conformance with the original sequence. RT-PCR. HEK293T cells and LECs were mock-treated, infected with HA units/ml SeV (Cantrell strain; Charles River), or stimulated with 1, units/ml IFN- (Sigma) or 1 M all-trans retinoic acid (Sigma). Furthermore, HCT116, Huh7, HeLa, LnCap, NHLF, 2fTGH wild-type, U3A, and U6A cells were treated with 1, units/ml IFN- (Sigma) or 1, units/ml IFN- (GeneTex). Total cellular RNA was extracted by using the RNeasy Plus mini kit (Qiagen). RNA (1 5 g) was used for cdna synthesis using the SuperScript III First Strand kit (Invitrogen), followed by PCR using RIG-I gene-specific primers. To screen for splice variants of the RIG-I CARDs, the forward primer 5 -(ATAGGATCCATCCTGAGCTACATG- GCCCCCTGG)-3 and the reverse primer 5 -(TTCCACAAC- CTGTAGGAGCACATA)-3 were used. To amplify specifically RIG-I or RIG-I splice variant transcripts, we used the following primer sets: forward 5 -(TGGTTCCGTGGCTTTT- TGGATGCC)-3 and reverse 5 -(TTCCACAACCTGTAG- GAGCACATA)-3 for RIG-I ; forward 5 -(CCCTGGTT- TAGGGAGGGTTATTCT)-3 and reverse 5 -(TTCCACAAC- CTGTAGGAGCACATA)-3 for the RIG-I splice variant. -Actin was amplified by using TGGACATCCGCAAAGACCTG (forward) and CCGATCCACACGGAGTACTT (reverse). Molecular Cloning of RIG-I Splice Variant. HEK293T cells were treated with IFN- (1, units/ml) for 24 h. Total RNA was extracted by using the RNeasy Plus mini kit (Qiagen) and reverse transcribed by using Long Range 2Step RT-PCR kit (Qiagen). From the resulting cdnas, RIG-I and splice variant were amplified by using the following primers: forward 5 - (ATGACCACCGAGCAGCGACGCAGC)-3 and reverse 5 - (TCATTTGGACATTTCTGCTGGATC)-3. The resulting amplicon of 2.7 kb was digested with SacI to specifically digest RIG-I but not RIG-I splice variant transcripts. After cloning into the pgem-teasy vector (Promega), clones were subjected to sequence analysis using an AI PRISM 377 automatic DNA sequencer. GST Pulldown Assay, Immunoprecipitation, and Immunoblot Analysis. HEK293T cells were lysed in Nonidet P-4 buffer [5 mm Hepes (ph 7.4), 15 mm NaCl, 1% (vol/vol) Nonidet P-4, and protease inhibitor mixture (Roche)]. GST pulldown and immunoprecipitations were performed as described in ref. For immunoblotting, proteins were resolved by SDS/PAGE and transferred onto a PVDF membrane. The following primary antibodies were used: anti-v5 (1:5,) (Invitrogen), anti-flag (1:5,) (Sigma), anti-ha (1:5,) (Sigma), anti-myc (1:5,) (Convance), anti- GST (1:1,) (Sigma), anti-actin (1:1,) (Abcam), antiubiquitin (P4D1; Santa Cruz iotechnology), anti-trim (D iosciences), monoclonal anti-rig-i (1:1,) (Alexis), polyclonal anti-rig-i (1:) (IL), anti-irf3 (1:5) (Santa Cruz iotechnology), and anti-phospho-ser 396 -IRF3 (1:) (Upstate iotechnology). The proteins were visualized by an enhanced chemiluminescence reagent (Pierce) and detected by a phospho imager (Fuji LAS-4). Protein Expression, Fluorescence and Anisotropy Measurement. Protein expressions of RIG-I and RIG-I splice variant in insect cells were performed as described in ref. 2. Proteins were purified to homogeneity by using metal affinity (Qiagen), anion exchange and gel filtration chromatography with standard protocols. Fluorescence anisotropy experiments were performed with a FluoroMax-P fluorimeter (Horiba Jobin Yvon), equipped with a Glan Thompson prism polarizer. Typically, 1.2 ml of buffer [ mm Tris HCl (ph 7.5), 15 mm NaCl, 2 mm DTT, and 1 M ZnCl 2 ] and 5 nm RNA (in vitro-transcribed ppprvl with incorporated Alexa Fluor UTP) were preequilibrated in a quartz cuvette at 12 C. Protein samples were added in a stepwise manner and briefly mixed by magnetic stirring. After 3 min of reequilibration, the anisotropy data were measured in triplets for each titration step by using an excitation wavelength of 492 nm and monitoring the emission 516 nm. The band pass was 5 nm for excitation and 5 nm for emission. Luciferase Reporter Assay. HEK293T, HCT116, Huh7, and HeLa cells were seeded into six-well plates. At 24 h, the cells were transfected with an IFN- or NF- luciferase construct together with constitutive -gal-expressing pgk- -gal. At 24 h after transfection, the cells were mock-treated or infected with SeV (5 HA units/ml) for 16 h. s were prepared and subjected to a luciferase assay (Promega). Luciferase values were normalized to -galactosidase to measure transfection efficiency. IFN- ELISA and V replication. HEK293T cells were seeded into a 12-well plate and mock-infected or infected with SeV (5 HA Gack et al. 1of11
2 units/ml) for h. The supernatants were collected and analyzed for IFN- production by using enzyme-linked immunosorbent assays (PL iomedical Laboratories). For viral replication assays, stable HEK293T or RIG-I knockout MEFs were seeded into six-well plates and infected with V-eGFP at MOI.5. At h after infection, the culture medium was harvested and the virus yield determined by plaque assay on Vero cells. 1. Gack MU, et al. (7) TRIM RING-finger E3 ubiquitin ligase is essential for RIG-Imediated antiviral activity. Nature 446: Cui S, et al. (8) The C-terminal regulatory domain is the RNA 5 -triphosphate sensor of RIG-I. Mol Cell 29: Gack et al. 2of11
3 2CARD 1 st CARD 2 nd CARD tor TRIM-V NF-κ luciferase activity 5 37 I: αgst I: αub TRIM GST 2CARD 1 st CARD 2 nd CARD Fig. S1. The intact tandem CARD of RIG-I is required for TRIM-mediated RIG-I ubiquitination and downstream signaling. (A) oth CARDs are required for TRIM-mediated RIG-I ubiquitination. HEK293T cells were transfected with GST-RIG-I 2CARD, GST-RIG-I first CARD, or GST-RIG-I second CARD with or without V5-tagged TRIM. s were used for GST pull down (GST-PD), followed by I with -GST (Top)or -Ub (Middle) to show the ubiquitination of RIG-I constructs. s were used for immunoblotting (I) with -V5 (ottom). Arrows indicate the ubiquitinated bands. () oth CARDs are necessary for TRIM-induced RIG-I NF- activation. HEK293T were transfected with GST, GST-RIG-I 2CARD, GST-RIG-I first CARD, or GST-RIG-I second CARD with or without TRIM together with NF- luciferase and constitutive -gal-expressing pgk- -gal reporter. Luciferase values were determined and normalized to -galactosidase. Data represent the mean SD (n 3). Gack et al. 3of11
4 GST T 55I I: αtrim RIG-I 2CARD-Flag GST + - GST-MAVS-CARD-PRD I: αgst I: αtrim I: αgst C TRIM shrna GST GST-RIG-I 2CARD MAVS CARD-PRD-Flag I: αgst I: αub I: αtrim I: αactin (%) % 85.1% total RIG-I () ubiquitinated ubiquitinated non-ubiquitinated nonubiquitinated 49.8% 5.92% RIG-I bound to MAVS (GST-PD) Fig. S2. TRIM-mediated RIG-I ubiquitination is critical for RIG-I-MAVS interaction. (A) T 55 I mutation of RIG-I abolishes TRIM binding. At 48 h after transfection with GST, GST-RIG-I 2CARD, or GST-RIG-I 2CARD T 55 I mutant, s were used for GST-PD, followed by I with -TRIM (Top)or -GST (Middle). s were immunoblotted with -TRIM (ottom). Arrows indicate the ubiquitinated bands. () MAVS preferentially interacts with ubiquitinated RIG-I. (Upper) At 48 h after transfection with GST or GST-MAVS-CARD-PRD together with Flag-tagged RIG-I 2CARD, HEK293T s were used for GST-PD, followed by I with -Flag (Top) or -GST (Middle). s were used for I with -Flag to determine the expression of RIG-I 2CARD (ottom). Arrows indicate the ubiquitinated bands. (Lower) The quantitation of ubiquitinated and nonubiquitinated RIG-I bands in the s and in the GST-MAVS-CARD-PRD complex by using a Fuji phospho imager. (C) TRIM-mediated ubiquitination is necessary for efficient RIG-I-MAVS interaction. At 48 h after transfection with GST or GST-RIG-I 2CARD together with MAVS-CARD-PRD-Flag and increasing amount of TRIM shrna-specific retroviral psuper vector, HEK293T s were used for GST-PD, followed by I with -Flag, -GST, or -Ub. Arrows indicate the ubiquitinated bands. s were used for I with -TRIM or -Flag to show the expression of TRIM and MAVS-CARD-PRD, respectively. Loading control was determined by using an -actin antibody. Gack et al. 4of11
5 IFN-β luciferase activity 5 37 GST T 55S T 55W T 55V I: αgst GST T 55S T 55W T 55V I: αub NF-κ luciferase activity I: αtrim I: αtrim GST T 55S T 55W T 55V Fig. S3. Mutation of T 55 to hydrophobic residues abolishes RIG-I CARD ubiquitination, TRIM binding, and RIG-I signaling. (A) Mutation of T 55 to hydrophobic residues abolishes RIG-I CARD ubiquitination and TRIM binding. At 48 h after transfection with GST, GST-RIG-I 2CARD, GST-RIG-I 2CARD T 55 S, GST-RIG-I 2CARD T 55 W, or GST-RIG-I 2CARD T 55 V, HEK293T s were subjected to GST-PD, followed by I with -GST, -Ub or -TRIM. To determine endogenous TRIM expression, s were subjected to I with -TRIM. Arrows indicate the ubiquitinated bands. () RIG-I T 55 W and T 55 V mutants show a near complete loss of downstream signaling activity. GST, GST-RIG-I 2CARD, GST-RIG-I 2CARD T 55 S, GST-RIG-I 2CARD T 55 W, or GST-RIG-I 2CARD T 55 V was expressed in HEK293T together with IFN- or NF- luciferase and pgk- -gal. At 36 h after transfection, the cells were harvested and the luciferase and -galactosidase values determined. Luciferase values, normalized to -galactosidase activity, are presented as. Data represent the mean SD (n 3). Gack et al. 5of11
6 735 bp 63 bp - SeV 12 h 24 h 48 h - IFN-β RA 12 h 24 h 48 h 48 h 72 h 588 bp 588 bp 65 bp 65 bp actin - Flag - Flag actin IP: αrig-i I: αrig-i C I αrig-i (helicase) αrig-i (1 st CARD) IP: αrig-i Fig. S4. RIG-I splice variant transcript and protein. (A) RIG-I and splice variant transcripts and proteins in HEK293T upon SeV infection. HEK293T were mock-infected or infected with HA units/ml SeV for the indicated number of hours. To identify potential splice variants of the RIG-I CARDs, total cellular RNA was isolated and subjected to RT-PCR to amplify the RIG-I CARDs (exon 1 3) (Top). Transcript levels of RIG-I and the splice variant () were determined using primers specific for each isoform (Second and Third images). Actin transcripts were determined as control. Endogenous protein levels of RIG-I and upon SeV infection were evaluated by IP with -RIG-I followed by I with -RIG-I (ottom). Flag-tagged RIG-I and were loaded as controls. () RIG-I and splice variant transcript levels in LECs. LECs were mock-treated, treated with 1, units/ml IFN-, or1 all-trans retinoic acid (RA) for the indicated hours. Total cellular RNA was isolated and subjected to RT-PCR with primers specific for RIG-I or, respectively. Actin transcripts were determined as control.(c) Control IP for the RIG-I splice variant. s of HEK293T cells, treated with IFN- (1, U/ml) for 24 h, were used for immunoprecipitation with an -RIG-I antibody recognizing the central helicase domain (residues 1 713). Precipitated endogenous RIG-I and splice variant were determined by I with RIG-I antibodies, recognizing the helicase domain (residues 1 713) or first CARD (residues 37 55), respectively. Arrows indicate RIG-I or the splice variant. Gack et al. 6of11
7 Fig. S5. IFN-dependent expression of RIG-I splice variant in various cell lines. (A) HeLa, HCT116, Huh7, LnCap, and NHLF cells were mock-treated or stimulated with 1, units/ml IFN- for 24 h. Total cellular RNA was isolated and subjected to RT-PCR with primers specific for RIG-I or, respectively. Actin transcripts were determined as control. () The 2fTGH wild-type, STAT1-deficient (U3A), and STAT2-deficient (U6A) human fibroblasts were mock-treated or treated with 1, units/ml IFN- or IFN- for 24 h. Total cellular RNA was isolated and subjected to RT-PCR with primers specific for RIG-I or, respectively. Actin transcripts were determined as control. Gack et al. 7of11
8 TRIM SPRY-V GST 2CARD 2CARD I: αgst TRIM-V5 T 55 I K 172 R C T 55 I K 172 R : HA-K 63only -Ub D I: αha I: αha I: αactin GST NF-κ luciferase activity TRIM-V CARD 2CARD Fig. S6. Abolished TRIM binding, ubiquitination, and signaling activity of RIG-I splice variant. (A) RIG-I but not RIG-I splice variant interacts with TRIM-SPRY. V5-tagged TRIM-SPRY together with GST, GST-RIG-I 2CARD, or GST-RIG 2CARD were expressed in HEK293T cells. At 48 h after transfection, cells were lysed, and s were subjected to GST-PD, followed by I with -V5 (Top) or -GST (Middle). s were used for I with -V5 to determine the expression of TRIM-SPRY (ottom). () RIG-I splice variant and RIG-I T 55 I do not interact with TRIM. After transfection of V5-tagged TRIM together with vector, Flag-tagged RIG-I, RIG-I T 55 I, RIG-I, or RIG-I K 172 R, HEK293T s were subjected to IP with -Flag, followed by I with -V5 (Top)or -Flag (Middle). s were used for I with -V5 to test the TRIM expression (ottom). (C) RIG-I splice variant exhibits a near-complete loss of K63-linked ubiquitination. Flag-tagged RIG-I, RIG-I T 55 I, RIG-I or RIG-I K 172 R together with HA-tagged K 63only -ubiquitin were expressed in HEK293T cells. At 24 h after transfection, the cells were infected with SeV ( HA units/ml) for 1 h. s were used for IP with -Flag, followed by I with -HA (Top) or -Flag (second). s were subjected to I with -HA to determine HA-ubiquitin expression (third). Loading control was determined by using an -actin antibody (fourth). (D) Lack of signaling activity of RIG-I splice variant. HEK293T were transfected with GST, GST-RIG-I 2CARD, or GST-RIG-I 2CARD with or without V5-tagged TRIM together with NF- luciferase and pgk- -gal. At 36 h after transfection, the cells were harvested, and luciferase and -galactosidase values were determined. Luciferase values were normalized to -galactosidase activity and are presented as. Data represent the mean SD (n 3). Gack et al. 8of11
9 NF-κ luciferase activity T 55 I Mock SeV Fig. S7. RIG-I splice variant inhibits SeV-induced NF- promoter activation. HEK293T were transfected with vector or increasing amount of RIG-I T 55 I or RIG-I splice variant () together with NF- luciferase and pgk- -gal. At 24 h after transfection, the cells were mock-infected or infected with SeV for 16 h. Luciferase and -galactosidase values were determined and NF- luciferase values normalized to -galactosidase activity for transfection efficiency control. Data represent the mean SD (n 3). Gack et al. 9of11
10 HCT116 IFN-β luciferase activity Mock SeV Huh7 IFN-β luciferase activity Mock SeV HeLa IFN-β luciferase activity Mock SeV Fig. S8. RIG-I splice variant inhibits SeV-induced IFN- promoter activation in various cell lines. HCT116, Huh7, and HeLa cells were transfected with vector or increasing amount of RIG-I splice variant () together with IFN- luciferase and pgk- -gal. At 24 h after transfection, the cells were mock-infected or infected with SeV for 16 h. Luciferase and -galactosidase values were determined and IFN- luciferase values normalized to -galactosidase activity for transfection efficiency control. Data represent the mean SD (n 3). Gack et al. 1 of 11
11 .1 Anisotropy Protein [nm] -V : RIG-I -Flag : RIG-I -Myc I: αmyc C T 55I :SeV : MAVS-Flag : RIG-I -Myc I: αmyc I: αmyc I: αmyc Fig. S9. (A) Fluorescence anisotropy changes of fluorescently labeled ppprvl in response to titration with RIG-I (filled square kd nm) and RIG-I (open circles, kd nm). The plot was non-linearly fitted according to a one-site binding model A max * [Protein] / (kd [Protein]). () RIG-I splice variant interferes with virus-induced RIG-I multimerization. HEK293T cells were transfected with vector, Myc-tagged RIG-I, Flag-tagged RIG-I and increasing amount of V5-tagged RIG-I. At 24 h posttransfection, the cells were infected with SeV for 16 h. Co-immunoprecipitation of Myc-RIG-I with Flag-RIG-I was determined by IP using -Flag followed by I with -Myc or -Flag. s were subjected to I with -Myc or -V5. (C) RIG-I splice variant inhibits the virus-induced RIG-I-MAVS-complex formation. HEK293T were cotransfected with Flag-tagged full-length MAVS and Myc-tagged RIG-I together with vector, V5-tagged RIG-I T 55 I, or splice variant () and subsequently either mock-treated or infected with SeV (5 HA units/ml) for 16 h. s were subjected to IP with -Flag, followed by I with -Myc or -Flag. s were subjected to I with -Myc or -V5. Gack et al of 11
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
Conventional PKC-α/β Negatively Regulate RIG-I Antiviral Signal Transduction
 JVI Accepts, published online ahead of print on 23 November 2011 J. Virol. doi:10.1128/jvi.06543-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JVI Accepts, published online ahead of print on 23 November 2011 J. Virol. doi:10.1128/jvi.06543-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
Species-Specific Inhibition of RIG-I Ubiquitination and IFN Induction by the Influenza A Virus NS1 Protein
 Species-Specific Inhibition of RIG-I Ubiquitination and IFN Induction by the Influenza A Virus NS1 Protein Ricardo Rajsbaum 1,2, Randy A. Albrecht 1,2, May K. Wang 3, Natalya P. Maharaj 3, Gijs A. Versteeg
Species-Specific Inhibition of RIG-I Ubiquitination and IFN Induction by the Influenza A Virus NS1 Protein Ricardo Rajsbaum 1,2, Randy A. Albrecht 1,2, May K. Wang 3, Natalya P. Maharaj 3, Gijs A. Versteeg
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary Figure 1
 Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
(Stratagene, La Jolla, CA) (Supplemental Fig. 1A). A 5.4-kb EcoRI fragment
 SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Materials and Methods
 Supplementary Materials and Methods Reagents and antibodies was purchased from iaffin GmbH & Co KG. Cisplatin (ristol-myers Squibb Co.) and etoposide (Sandoz Pharma Ltd.) were used. Antibodies recognizing
Supplementary Materials and Methods Reagents and antibodies was purchased from iaffin GmbH & Co KG. Cisplatin (ristol-myers Squibb Co.) and etoposide (Sandoz Pharma Ltd.) were used. Antibodies recognizing
Supporting Information
 Supporting Information Zhu et al. 1.173/pnas.11167618 SI Materials and Methods DNA Construction. The plasmid pcdna4to/myc-rzap, which expresses myc-tagged full-length rat ZAP, has been described previously
Supporting Information Zhu et al. 1.173/pnas.11167618 SI Materials and Methods DNA Construction. The plasmid pcdna4to/myc-rzap, which expresses myc-tagged full-length rat ZAP, has been described previously
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3
 Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
TRIM25 Is Required for the Antiviral Activity of Zinc-finger Antiviral Protein
 JVI Accepted Manuscript Posted Online 15 February 2017 J. Virol. doi:10.1128/jvi.00088-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 TRIM25 Is Required for the Antiviral
JVI Accepted Manuscript Posted Online 15 February 2017 J. Virol. doi:10.1128/jvi.00088-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 TRIM25 Is Required for the Antiviral
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supporting Information
 Supporting Information Kim et al. 10.1073/pnas.0912180106 SI Materials and Methods DNA Constructs. The N-terminally-tagged mouse Gli2 expression construct, pcefl/3 HA-Gli2, was constructed by ligating
Supporting Information Kim et al. 10.1073/pnas.0912180106 SI Materials and Methods DNA Constructs. The N-terminally-tagged mouse Gli2 expression construct, pcefl/3 HA-Gli2, was constructed by ligating
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth.
 Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
The SAMHD1-mediated block of LINE-1 retroelements is regulated by phosphorylation
 Herrmann et al. Mobile DNA (2018) 9:11 https://doi.org/10.1186/s13100-018-0116-5 RESEARCH Open Access The SAMHD1-mediated block of LINE-1 retroelements is regulated by phosphorylation Alexandra Herrmann
Herrmann et al. Mobile DNA (2018) 9:11 https://doi.org/10.1186/s13100-018-0116-5 RESEARCH Open Access The SAMHD1-mediated block of LINE-1 retroelements is regulated by phosphorylation Alexandra Herrmann
William C. Comb, Jessica E. Hutti, Patricia Cogswell, Lewis C. Cantley, and Albert S. Baldwin
 Molecular Cell, Volume 45 Supplemental Information p85 SH2 Domain Phosphorylation by IKK Promotes Feedback Inhibition of PI3K and Akt in Response to Cellular Starvation William C. Comb, Jessica E. Hutti,
Molecular Cell, Volume 45 Supplemental Information p85 SH2 Domain Phosphorylation by IKK Promotes Feedback Inhibition of PI3K and Akt in Response to Cellular Starvation William C. Comb, Jessica E. Hutti,
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection
 Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplemental Figure 1
 Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
A. List of selected proteins with high SILAC (H/L) ratios identified in mass
 Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element
 Research Article ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element Evelyn Chukwurah, Indhira Handy and Rekha C. Patel Department
Research Article ADAR1 and PACT contribute to efficient translation of transcripts containing HIV-1 trans-activating response (TAR) element Evelyn Chukwurah, Indhira Handy and Rekha C. Patel Department
316 Cell 141, , April 16, 2010 ª2010 Elsevier Inc.
 Figure 1. In Vitro Reconstitution of the RIG-I Pathway and Regulation of RIG-I by RNA and Ubiquitination (A) Purification of RIG-I protein from Sendai virus-infected (+SeV) or untreated ( SeV) HEK293T
Figure 1. In Vitro Reconstitution of the RIG-I Pathway and Regulation of RIG-I by RNA and Ubiquitination (A) Purification of RIG-I protein from Sendai virus-infected (+SeV) or untreated ( SeV) HEK293T
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Supporting Information
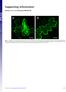 Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
~Lentivirus production~
 ~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator. of the Interaction with Macrophages
 Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator of the Interaction with Macrophages Yohei Sanada, Takafumi Yamamoto, Rika Satake, Akiko Yamashita, Sumire Kanai, Norihisa Kato, Fons AJ van
Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator of the Interaction with Macrophages Yohei Sanada, Takafumi Yamamoto, Rika Satake, Akiko Yamashita, Sumire Kanai, Norihisa Kato, Fons AJ van
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Figure S1! CrFK! DNA Synthesis! Copies viral DNA/ 100 ng DNA (Log 10 ) % Infected (GFP+) cells! 100! 80! 60! 40! 20! 0! 1! 10! 100! 1000!
 Figure S1! A! B! % Infected (GFP+) cells! 100! 80! 60! 40! 20! 0! 1! 10! 100! 1000! Dose μl! CrFK! B-MLV! N-MLV! Template:! Water! Ube2V2! Ube2V1! Primers:! V2! V1! V2! V1! V2! V1! bp! 500! 300! 200! 150!
Figure S1! A! B! % Infected (GFP+) cells! 100! 80! 60! 40! 20! 0! 1! 10! 100! 1000! Dose μl! CrFK! B-MLV! N-MLV! Template:! Water! Ube2V2! Ube2V1! Primers:! V2! V1! V2! V1! V2! V1! bp! 500! 300! 200! 150!
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Nature Immunology doi: /ni Supplementary Figure 1. Raf-1 inhibition does not affect TLR4-induced type I IFN responses.
 Supplementary Figure 1 Raf-1 inhibition does not affect TLR4-induced type I IFN responses. Real-time PCR analyses of IFNB, ISG15, TRIM5, TRIM22 and APOBEC3G mrna in modcs 6 h after stimulation with TLR4
Supplementary Figure 1 Raf-1 inhibition does not affect TLR4-induced type I IFN responses. Real-time PCR analyses of IFNB, ISG15, TRIM5, TRIM22 and APOBEC3G mrna in modcs 6 h after stimulation with TLR4
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Mapping the Ligand-binding Site on a GPCR Using Genetically-encoded Photocrosslinkers
 Mapping the Ligand-binding Site on a GPCR Using Genetically-encoded Photocrosslinkers Amy Grunbeck, Thomas Huber, Pallavi Sachdev, Thomas P. Sakmar Laboratory of Molecular Biology and Biochemistry, The
Mapping the Ligand-binding Site on a GPCR Using Genetically-encoded Photocrosslinkers Amy Grunbeck, Thomas Huber, Pallavi Sachdev, Thomas P. Sakmar Laboratory of Molecular Biology and Biochemistry, The
The Cellular RNA Helicase DDX1 Interacts with Coronavirus Nonstructural Protein 14 and Enhances Viral Replication
 JOURNAL OF VIROLOGY, Sept. 2010, p. 8571 8583 Vol. 84, No. 17 0022-538X/10/$12.00 doi:10.1128/jvi.00392-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The Cellular RNA Helicase
JOURNAL OF VIROLOGY, Sept. 2010, p. 8571 8583 Vol. 84, No. 17 0022-538X/10/$12.00 doi:10.1128/jvi.00392-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The Cellular RNA Helicase
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Senior Thesis. Presented to. The Faculty of the School of Arts and Sciences Brandeis University
 Greenwald 1 Mouse intercellular adhesion molecule 1 (ICAM-1) isoforms demonstrate different binding affinities to mouse macrophage-1 antigen (Mac-1) and preliminary evidence for alternatively-spliced variants
Greenwald 1 Mouse intercellular adhesion molecule 1 (ICAM-1) isoforms demonstrate different binding affinities to mouse macrophage-1 antigen (Mac-1) and preliminary evidence for alternatively-spliced variants
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
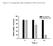 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Innate Immunity & Inflammation
 Innate Immunity & Inflammation The innate immune system is an evolutionally conserved mechanism that provides an early and effective response against invading microbial pathogens. It relies on a limited
Innate Immunity & Inflammation The innate immune system is an evolutionally conserved mechanism that provides an early and effective response against invading microbial pathogens. It relies on a limited
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document
 Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell
 Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Appendix. Table of Contents
 Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
SUPPLEMENTARY INFORMATION
 Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Supplementary Figures
 Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Peli1 negatively regulates T-cell activation and prevents autoimmunity
 Peli1 negatively regulates T-cell activation and prevents autoimmunity Mikyoung Chang 1,*, Wei Jin 1,5,*, Jae-Hoon Chang 1, Yi-chuan Xiao 1, George Brittain 1, Jiayi Yu 1, Xiaofei Zhou 1, Yi-Hong Wang
Peli1 negatively regulates T-cell activation and prevents autoimmunity Mikyoung Chang 1,*, Wei Jin 1,5,*, Jae-Hoon Chang 1, Yi-chuan Xiao 1, George Brittain 1, Jiayi Yu 1, Xiaofei Zhou 1, Yi-Hong Wang
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Ubiquitin-Regulated Recruitment of I B Kinase ε to the MAVS Interferon Signaling Adapter
 MOLECULAR AND CELLULAR BIOLOGY, June 2009, p. 3401 3412 Vol. 29, No. 12 0270-7306/09/$08.00 0 doi:10.1128/mcb.00880-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Ubiquitin-Regulated
MOLECULAR AND CELLULAR BIOLOGY, June 2009, p. 3401 3412 Vol. 29, No. 12 0270-7306/09/$08.00 0 doi:10.1128/mcb.00880-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Ubiquitin-Regulated
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplemental information contains 7 movies and 4 supplemental Figures
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplemental information contains 7 movies and 4 supplemental Figures Movies: Movie 1. Single virus tracking of A4-mCherry-WR MV
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplemental information contains 7 movies and 4 supplemental Figures Movies: Movie 1. Single virus tracking of A4-mCherry-WR MV
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ).
 Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
FIG S1 Examination of eif4b expression after virus infection. (A) A549 cells
 Supplementary Figure Legends FIG S1 Examination of expression after virus infection. () 549 cells were infected with herpes simplex virus (HSV) (MOI = 1), and harvested at the indicated times, followed
Supplementary Figure Legends FIG S1 Examination of expression after virus infection. () 549 cells were infected with herpes simplex virus (HSV) (MOI = 1), and harvested at the indicated times, followed
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway
 Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
HDAC6 regulates cellular viral RNA sensing by deacetylation of RIG-I
 Article HDAC6 regulates cellular viral RNA sensing by deacetylation of RIG-I Su Jin Choi 1,, Hyun-Cheol Lee 2,, Jae-Hoon Kim 2, Song Yi Park 1, Tae-Hwan Kim 2, Woon-Kyu Lee 3, Duk-Jae Jang 2, Ji-Eun Yoon
Article HDAC6 regulates cellular viral RNA sensing by deacetylation of RIG-I Su Jin Choi 1,, Hyun-Cheol Lee 2,, Jae-Hoon Kim 2, Song Yi Park 1, Tae-Hwan Kim 2, Woon-Kyu Lee 3, Duk-Jae Jang 2, Ji-Eun Yoon
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
