MicroRNAs coordinate an alternative splicing network during mouse postnatal heart development
|
|
|
- Ralph Harmon
- 5 years ago
- Views:
Transcription
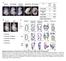
1 RESEARCH COMMUNICATION MicroRNAs coordinate an alternative splicing network during mouse postnatal heart development Auinash Kalsotra, 1 Kun Wang, 2 Pei-Feng Li, 2 and Thomas A. Cooper 1,3,4 1 Department of Pathology and Immunology, Baylor College of Medicine, Houston, Texas 77030, USA; 2 Division of Cardiovascular Research, National Key Laboratory of Biomembrane and Membrane Biotechnology, Institute of Zoology, Chinese Academy of Sciences, Beijing , People s Republic of China; 3 Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas 77030, USA Alternative splicing transitions have been identified recently as a conserved component of vertebrate heart remodeling during postnatal development. Here we report that the targeted deletion of Dicer, specifically in adult mouse myocardium, reveals the role of micrornas (mirnas) in regulating networks of postnatal splicing transitions and in maintaining adult splicing programs. We demonstrate a direct role for mir-23a/b in the dramatic postnatal down-regulation of CUGBP and ETR-3-like factor (CELF) proteins that regulate nearly half of developmentally regulated splicing transitions in the heart. These findings define a hierarchy in which rapid postnatal upregulation of specific mirnas controls expression of alternative splicing regulators and the subsequent splicing transitions of their downstream targets. Supplemental material is available at Received December 7, 2009; revised version accepted February 17, MicroRNAs (mirnas) are key components of posttranscriptional gene regulation with diverse roles in tissue development, differentiation, and homeostasis (van Rooij et al. 2007; Zhao et al. 2007; Bartel 2009). Mature mirnas are formed by two sequential processing reactions: Primary transcripts of mirna genes are first cleaved into hairpin-containing intermediates (pre-mirnas) by the Drosha microprocessor complex, and pre-mirnas are then processed into mature mirnas by Dicer (Kim 2005). While mirna-mediated regulation stimulates quantitative changes in protein and mrna expression (Baek et al. 2008; Selbach et al. 2008), alternative pre-mrna splicing controls qualitative gene output in the expression of diverse mrna isoforms (Blencowe 2006). Recent genome-wide studies revealed that >90% of human intron-containing [Keywords: Alternative splicing networks; CELF proteins; heart development; microrna] 4 Corresponding author. tcooper@bcm.edu; FAX (713) Article published online ahead of print. Article and publication date are online at genes are alternatively spliced, and more than half of alternative splicing events differ between tissues, revealing an extensive level of regulation (Castle et al. 2008; Pan et al. 2008; Wang et al. 2008). The mammalian heart undergoes a period of dramatic remodeling during the first 3 wk after birth that requires transcriptional and post-transcriptional regulatory programs not yet fully understood (Xu et al. 2005; Olson 2006). We recently identified a conserved set of alternative splicing transitions during mouse heart development, and demonstrated that nearly half are responsive to the CUGBP and ETR-3-like factor (CELF) family of splicing regulators (Kalsotra et al. 2008). Two CELF proteins, CUGBP1 and CUGBP2, are highly expressed in the fetal heart, but are down-regulated >10 fold within 3 wk after birth without a change in mrna levels. Here we used targeted deletion of the Dicer gene specifically in adult mouse myocardium to test the role of mirnas in the mechanism of CELF protein downregulation during postnatal development. Both CELF proteins were dramatically up-regulated within 2 d of Dicer knockout, as was a subset of five other splicing regulators, while spliceosomal components and an additional set of splicing regulators were not affected. Using multiple independent assays, including antagomir delivery to adult mice, we demonstrate a direct role for mir- 23a/b in the postnatal down-regulation of the two CELF proteins. Using a data set of postnatal splicing transitions for which we identified cognate regulators, we demonstrate splicing networks that are and are not mirnadependent in both the Dicer knockout and antagomirtreated hearts. The effects of the Dicer knockout are specific, as neither the expression of splicing regulators nor their target splicing events are affected in three different mouse models of cardiomyopathy. Our results define an important hierarchy in which mirna expression controls expression of alternative splicing regulators during a critical period of heart development. Results and Discussion The two CELF paralogs expressed in the heart, CUGBP1 and CUGBP2, exhibit a respective 10-fold and 18-fold decrease in protein during the first 2 wk after birth without a decrease in mrna levels, indicative of posttranscriptional regulation (Kalsotra et al. 2008). CUGBP1 is stabilized by phosphorylation, which decreases during postnatal heart development (Supplemental Fig. 1; Kuyumcu-Martinez et al. 2007). In contrast, CUGBP2 exhibits no change in its phosphorylation state during postnatal development, indicating a different mechanism for its down-regulation (Supplemental Fig. 1). To determine whether mirnas are required for repression of CUGBP1 and CUGBP2 proteins in adult hearts, we generated tamoxifen (Tam)-inducible, heart-specific Dicer knockout mice using a Cre-loxP approach (Supplemental Fig. 2A). Mice contain a single Dicer gene that is required to generate mature mirnas (Birchler and Kavi 2008). Inducible ablation of Dicer in adult hearts overcame previously described early developmental defects (Chen et al. 2008) and allowed timed analysis to distinguish primary effects from secondary compensatory changes. PCR analysis on DNA from heart tissue of Dicer GENES & DEVELOPMENT 24: Ó 2010 by Cold Spring Harbor Laboratory Press ISSN /10; 1
2 Kalsotra et al. homozygous floxed (f/f); MerCreMer (MCM) mice showed recombination only upon Tam treatment, indicating minimal recombination in uninduced mice (Supplemental Fig. 2B). Western blot analysis showed a more than threefold decrease in Dicer protein 8 wk following Tam treatment (Fig. 1A). The loss of mature mirnas and accumulation of unprocessed precursors confirmed the functional loss of Dicer (Fig. 1B). Residual levels of Dicer protein and mature mirnas (Fig. 1A,B) in recombined heart tissue were most likely from noncardiomyocytes as well as incomplete recombination and/or long-lived mirnas. The cardiomyocyte-specific loss of Dicer in adult mice resulted in dilated cardiomyopathy and premature death within 10 wk of Tam treatment (Fig. 1C). Histology and echocardiography studies revealed dilation of the left ventricle, thinning of the left ventricular wall, and reduced fractional shortening accompanied by systolic and diastolic dysfunction in Dicer f/f ; MCM mice treated with Tam (Fig. 1D; Supplemental Fig. 3A D). We also detected profound myofiber disarray (Fig. 1E), increased heartto-body weight ratios, widespread apoptosis (Supplemental Fig. 3E,F), myocyte hypertrophy, inflammatory cell infiltration, and focal areas of mild interstitial fibrosis (data not shown). Tam-treated Dicer f/f mice and MCM controls showed no changes in several parameters examined (Supplemental Fig. 4A,B; data not shown). Similar effects have been described 1 wk after Tam induction (da Costa Martins et al. 2008). Consistent with a role for mirna-mediated repression in the adult heart, CUGBP1 and CUGBP2 proteins were strongly up-regulated in heart tissue from Dicer knockout mice 8 wk following Tam treatment compared with uninduced mice (Fig. 1F). To determine whether CELF protein up-regulation was a general response to cardiac injury, we assayed CUGBP1 and CUGBP2 protein levels in three different mouse models of heart disease (Sano et al. 2002; Braz et al. 2004). While each of these models exhibited altered expression of marker genes, CUGBP1 and CUGBP2 levels were unaffected (Supplemental Fig. 4C). Consistent with elevated CELF protein expression in Dicer knockout hearts, previously identified CUGBP1 target exons that switch their splicing pattern during normal postnatal heart development (Kalsotra et al. 2008) reverted to the embryonic pattern, but were unaffected in the other three models of heart disease (Fig. 1G; Supplemental Fig. 4D). Several constitutive and alternative exons that are not regulated during postnatal heart development showed no changes in Dicer knockout hearts (Supplemental Fig. 5). We conclude that CELF protein induction and the subsequent reversion to embryonic splicing patterns is due to the loss of mirnas rather than a secondary response to cardiac injury. We found that CUGBP1 and CUGBP2 up-regulation in Dicer knockout hearts was relatively rapid, and was observed within 48 h after the last dose of Tam injections (Fig. 2A). Real-time RT PCR results (Fig. 2B) confirmed that CUGBP1 and CUGBP2 mrna levels in Dicer knockout hearts 48 h after Tam treatment remain constant despite striking changes in protein expression. Intriguingly, several other splicing regulators were also up-regulated rapidly after Dicer loss, including Fox-1, Fox-2, PTB, Tia1, and hnrnp H (Fig. 2A). Fox-2 showed a dramatic change in the ratio of its protein isoforms in response to the loss of Dicer, suggestive of a change in splicing pattern. In contrast, the expression of MBNL1, ASF/SF2, hnrnp C, and hnrnp L splicing regulators was unaffected (Fig. 2A). Figure 1. Cardiac-specific deletion of Dicer results in dilated cardiomyopathy and initiates an embryonic splicing program in the adult heart. (A) Western blot showing decrease in Dicer protein in Dicer knockout hearts. Fold decrease in Dicer protein (shown in parentheses) was determined by relative quantification of band intensities, normalized to GAPDH. (B) Northern blots showing depletion of mature mirnas and accumulation of pre-mirnas in Dicer knockout hearts. U6 snrna is the loading control. (C) Kaplan-Meier survival curves for Dicer f/f ; MCM mice with and without Tam treatment. (D,E) Gross morphology and representative images of H&E staining of transverse sections of hearts from Dicer f/f ; MCM mice with and without Tam treatment. (F) Western blots showing increase in steady-state levels of CUGBP1 and CUGBP2 proteins in Dicer knockout hearts. (G) Misregulation of CUGBP1 target exons in Dicer knockout hearts but not in other models of heart disease such as PKC-a transgenic (Tg) mice, CycT1/Gq bitransgenic (BiTg) mice, or wild-type mice after 8 wk of transverse aortic constriction (TAC). Each bar represents mean 6 SD (of four biological replicates) for the percent inclusion of the specified exon. P < 0.05; asterisk (*) indicates significantly different from wild-type mice. 2 GENES & DEVELOPMENT
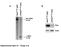
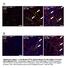
3 mirna control of alternative splicing in the heart Figure 2. Up-regulation of a subset of splicing regulators in Dicer knockout hearts occurs early and specifically in cardiomyocytes. (A) Steadystate protein levels of alternative splicing regulators. Fold changes are shown in parentheses. (B) CUGBP1 and CUGBP2 mrna levels, detected by real-time RT PCR, were unaffected by Dicer knockout. (C) Steady-state protein levels of basal spliceosomal factors. (D) Induced splicing factors in Dicer knockout hearts localize to cardiomyocyte nuclei. The top panel demonstrates immunoreactive nuclei (DAB staining) for CUGBP1, CUGBP2, and Fox-1, as indicated by black arrows. CUGBP1 expression in Dicer knockout hearts was elevated only in the nuclei of cells that costained with desmin (white arrows) and not in the nuclei of other cells (white outline). Cell nuclei were stained with DAPI. All images were taken using the same exposure time. Original magnification: C, top three panels, 340; C, bottom panel, 363. Bar, 50 mm. Embryonic reversion of splicing for CUGBP1 target exons (E) but not for MBNL1 target exons (F) in adult hearts of Dicer knockouts. (P < 0.05). (a) Significantly different from E14; (b) significantly different from Dicer f/f ; MCM Tam. In addition, none of the basal spliceosomal components tested showed a change in steady-state protein levels in Dicer knockout hearts (Fig. 2C). Therefore, these results identified a specific set of auxiliary splicing factors that are under regulatory control of mirnas in the mouse heart. Inflammatory cells were detected by 48 h after Dicer loss (data not shown), and we considered the possibility that these infiltrating cells were the source of the increased splicing factors. Immunohistochemistry for CUGBP1, CUGBP2, and Fox-1 in Dicer knockout hearts 48 h after Tam treatment demonstrated nuclear localization (Fig. 2D, black arrows, top panel). Coimmunofluorescent staining of CUGBP1 with the cardiomyocyte marker desmin confirmed that CUGBP1 protein was elevated only in the nuclei of cardiomyocytes (Fig. 2D, white arrows) but not other cell types (Fig. 2D, white circle, third panel from top). CUGBP1 expression in Dicer f/f ; MCM mice without Tam treatment remained low (Fig. 2D, second panel from top). Thus, we conclude that the rapid up-regulation of specific splicing factors following Dicer gene ablation is specific to cardiomyocytes. To determine whether early up-regulation of CELF proteins resulted in the embryonic splicing program in adult hearts, we examined splicing of five CUGBP1-regulated exons along with three MBNL1-regulated exons as controls for a splicing regulator not affected by Dicer knockout. As shown in Figure 2, E and F, all eight splicing events underwent robust splicing shifts between embryonic day 14 (E14) and the adult heart. However, only splicing events regulated by CUGBP1 and not those regulated by MBNL1 shifted to the embryonic pattern in Dicer knockout hearts, consistent with the respective changes in protein expression (Fig. 2A,E,F). These results provide strong evidence that, by regulating key splicing factors, mirnas coordinate developmental transitions in alternative splicing in the postnatal heart. mirna-based down-regulation of CELF protein expression during postnatal development could result from either a switch to longer 39-untranslated regions (UTRs) introducing new mirna-binding sites (Sandberg et al. 2008; Ji et al. 2009), or up-regulation of specific mirnas that target CUGBP1 and CUGBP2 mrnas. Northern blot analysis of CUGBP1 and CUGBP2 mrnas from E14 and adult mouse heart RNA revealed the absence of large structural changes in these mrnas during development (Supplemental Fig. 6). Three different algorithms were used to identify candidate mirnas that potentially regulate CUGBP1 and CUGBP2 expression. A number of mirnas targeting both CUGBP1 and CUGBP2 were predicted by each program, butonlymir-96andmir-23werepredictedbyallthree algorithms. Consistent with earlier reports (Chen et al. 2008), we found that mir-96 is not expressed in the heart (Fig. 3B), and therefore focused on mir-23. mir-23 includes mir-23a and mir-23b, which are transcribed from different chromosomes, have identical seed sequences, and differ by only 1 nucleotide on their 39 ends. There are three conserved mir-23a/b seed matches in the 39-UTR of CUGBP1 and two in CUGBP2 (Fig. 3A). Strikingly, mir-23a/b expression displays an inverse temporal relationship to CUGBP1 and CUGBP2 protein expression during a narrow period of postnatal heart development. For instance, between postnatal days 6 and 14, mir-23a/b levels increase nearly fourfold, whereas CUGBP1 and CUGBP2 levels decline nearly fivefold and fourfold, respectively (Fig. 3B,C). GENES & DEVELOPMENT 3
4 Kalsotra et al. Figure 3. mir-23a/b coordinates the developmental loss of CELF proteins by targeting multiple conserved sites in their 39-UTRs. (A) Multiple putative mir-23a/b seed elements located in the 39-UTRs of CUGBP1 and CUGBP2 mrnas. (B) mir-23a/b exhibits a reciprocal expression pattern to CUGBP1 and CUGBP2 proteins during a narrow time frame of postnatal heart development. (C) Quantitation of steady-state levels of mir-23a/b, CUGBP1, and CUGBP2 proteins. (D) Relative luciferase activity derived from CUGBP1 and CUGBP2 39-UTR reporter constructs transfected in HEK293T cells after overexpression of NS or mir-23b. All values are normalized to b-gal activity produced from a cotransfected control plasmid. (E) Northern blot analysis demonstrating increased mature mir-23b levels in HL-1 cells infected with mir-23b lentivirus. (F) Significant decrease in CUGBP1 and CUGBP2 proteins in HL-1 cardiomyocytes upon mir- 23b overexpression. MBNL1 protein levels remain unchanged. Relative quantification of band intensities, normalized to GAPDH levels, are shown below blots. (G) Steady-state mrna levels of CUGBP1 and CUGBP2 by real-time RT PCR show no significant change after mir-23b overexpression in HL-1 cells. (H) Splicing of CUGBP1 target exons in HL-1 cells shifts from embryonic to adult patterns upon mir-23b overexpression. P < 0.05; asterisk (*) indicates significantly different from NS. To determine whether mir-23b can suppress endogenous CELF protein expression, we overexpressed mir-23b in HL-1 mouse cardiomyocytes (Claycomb et al. 1998). Both CUGBP1 and CUGBP2 are abundantly expressed in these cells, consistent with low expression of endogenous mir-23a/b and expression of the embryonic splicing program for nine out of the 10 developmentally regulated and CUGBP1-responsive splicing events that were screened (data not shown). We overexpressed mir-23b or NS using a lentivirus expression vector (Fig. 3E), and assayed steadystate protein and mrna levels for CUGBP1 and CUGBP2 by Western blotting and real-time RT PCR, respectively. CELF protein levels (Fig. 3F), but not mrna levels (Fig. 3G), were significantly decreased in the presence of mir- 23b when compared with NS. Consistent with protein down-regulation, CUGBP1 pre-mrna targets shifted from embryonic to adult splicing patterns in HL-1 cells overexpressing mir-23b (Fig. 3H). Furthermore, mir-23b and mir-23a overexpression in HEK293T cells also resulted in strong down-regulation of CELF protein, demonstrating that mir-23 regulation of CELF proteins is conserved between mice and humans, and suggesting redundancy between mir-23a and mir-23b in regulation of CELF proteins (Supplemental Figs. 8C, 9). To test whether mir-23a/b are necessary for suppressing CELF protein expression in adult hearts, we infused a mir- 23a or a negative control (NC) antagomir in wild-type adult mice for systemic delivery to the heart as described previously (Lin et al. 2009). The 25 and 75 mg/kg doses of mir-23a antagomir produced 50% and 65% knockdowns, respectively, of both mir-23a and mir-23b without causing any overt phenotype (Fig. 4A; data not shown). Administration of the mir-23a antagomir resulted in a significant increase in CUGBP1 and CUGBP2 protein levels in To determine whether CUGBP1 and CUGBP2 transcripts are regulated directly by mir-23a/b, we constructed reporter plasmids in which 39-UTR segments containing each of the three (CUGBP1) or two (CUGBP2) predicted binding sites were cloned downstream from a luciferase ORF, with or without mutations that would disrupt the predicted mirna interaction (Supplemental Fig. 7). We overexpressed mir-23b or a nonsilencing control mirna (NS) in human embryonic kidney (HEK) 293T cells using lentiviral constructs that coexpress TurboRed fluorescent protein. TurboRed fluorescence indicated a high efficiency of infection (Supplemental Fig. 8A) and mir-23b overexpression was confirmed using Northern blots (Supplemental Fig. 8B). mir-23b significantly repressed luciferase activity of each of the five constructs containing wild-type CUGBP1 and CUGBP2, but not the mutant reporter constructs, while cotransfection with NS had no effect (Fig. 3D). These results demonstrate a direct functional interaction between mir-23b and each of these regions within the 39-UTR of CUGBP1 and CUGBP2. Figure 4. mir-23a/b knockdown in adult mouse hearts reverses CUGBP1 and CUGBP2 protein repression and induces embryonic splicing patterns of CELF targets. (A) Knockdown efficiency of mir- 23a and mir-23b in adult hearts after mir-23a antagomir treatment (25 or 75 mg/kg per day) was determined by real-time RT PCR. (B) Steady-state mrna levels of CUGBP1 and CUGBP2 assayed by real-time RT PCR show no significant change after mir-23a/b knockdown. (C) Representative Western blots showing CUGBP1 and CUGBP2 up-regulation in adult hearts after mir-23a antagomir treatment. (D) Relative quantitation of CUGBP1 and CUGBP2 levels normalized to GAPDH. (E) Embryonic shift in splicing of CUGBP1 target exons upon mir-23a/b knockdown. Each bar represents mean 6 SD from six wild-type untreated control animals, six wild-type animals treated with NC antagomir, or six wild-type animals treated with mir-23a antagomir. P < 0.05; asterisk (*) indicates significantly different from NC anatgomir. 4 GENES & DEVELOPMENT
5 mirna control of alternative splicing in the heart adult hearts (Fig. 4C,D) without affecting their respective mrna levels (Fig. 4B). Importantly, the increase in CELF proteins correlated directly with an adult to embryonic shift in the splicing of CUGBP1 pre-mrna targets (Fig. 4E). In contrast, mir-23a/b knockdown had no effect on the splicing of three MBNL1-regulated exons (Supplemental Fig. 10), demonstrating specificity to CELF pre-mrna targets. These data indicate that postnatal up-regulation of mir-23a/b suppresses CUGBP1 and CUGBP2 protein expression in adult hearts via direct binding to their respective 39-UTRs, resulting in a physiological shift in the alternative splicing of a subset of splicing events. mirna-based gene silencing and alternative splicing have emerged as two key post-transcriptional mechanisms that function on a genome-wide scale to regulate tissue development and differentiation (Blencowe 2006; Bartel 2009). The interplay between the mirna and splicing regulation has been demonstrated in cell culture models in which large-scale changes in splicing occur due to a switch in the expression of splicing regulators PTB/nPTB that is controlled by cell- and tissue-specific mirnas (Boutz et al. 2007; Makeyev et al. 2007). Here we identified a posttranscriptional regulatory network operative during the critical period of postnatal heart development in which coordination of a predominantly conserved set of splicing transitions are driven to the adult pattern, at least in part by mirna-mediated regulation of splicing factor expression. Most of the auxiliary splicing regulators but none of the basal splicing components examined demonstrated Dicer-dependent regulation, indicating a large potential for linked programs of post-transcriptional regulation. These results also demonstrate that, in addition to modulating splicing transitions during postnatal heart development, mirnas also maintain appropriate expression of splicing regulators in adult hearts. Disruption of this regulation could cause previously unrecognized mechanisms of disease. For example, aberrant up-regulation of CUGBP1 in adult hearts is associated with myotonic dystrophy in which expression of embryonic splicing patterns in adult tissues lead to specific disease features (Cooper et al. 2009). Materials and methods Animal models Mice homozygous for the Dicer f/f alleles, were crossed with Tam-inducible MCM transgenic mice that express Cre recombinase protein under the control of the cardiac-specific a-myosin heavy-chain (MHC) promoter (Sohal et al. 2001; Yi et al. 2006). Cre-mediated recombination was induced in 2- to 4-mo-old mice by intraperitoneal injection of 20 mg/kg Tam (Tam; Sigma-Aldrich) once daily for five consecutive days. Dicer f/f ; MCM mice treated with vehicle alone ( Tam) were used as controls. In vivo inhibition of mir-23 was performed by infusing a mir-23a antagomir (25 or 75 mg/kg per day) for 1 wk using Alzet osmotic pumps as described previously (Lin et al. 2009). NC antagomir (59-AGUACUUUUGUGUAGUACAA-39) served as a NC. All experiments were conducted in accordance with the NIH Guide for the Use and Care of Laboratory Animals and were approved by the Institutional Animal Care and Use Committee of Baylor College of Medicine. Cell culture and transfections HL-1 cells were cultured on gelatin (0.02%, w/v)/fibronectin (10 mg/ml)- coated plates and were maintained in Claycomb medium (JRH Biosciences) supplemented with 10% fetal bovine serum (JRH Biosciences), 2 mm L-glutamine, 0.1 mm norepinephrine, 100 U/mL penicillin, and 100 mg/ml streptomycin (Life Technologies) as described previously (Claycomb et al. 1998). HEK293T cells were cultured in high-glucose (4.5 g/l) DMEM supplemented with 10% fetal bovine serum, 2 mm L-glutamine, 13 sodium pyruvate, 100 U/mL penicillin, and 100 mg/ml streptomycin. Viral stocks for mir-23b and a NS (contains a random nontargeting sequence; Open Biosystems) were produced using the lentiviral packaging plasmids pspax2 and pmd2.g (Addgene, Inc.), concentrated and titered as per the manufacturer s instructions. The HL-1 cells were infected in T25 flasks with mir- 23b or NS viral stocks in the presence of 8 mg/ml polybrene. After 48 h of selection in 2 mg/ml puromycin selection media, cells were harvested to isolate total RNA or proteins. For transient transfections, mirna mimics for NS, mir-22, or mir-23a were purchased from Dharmacon and were transfected at 100 nm final concentration using the manufacturer s instructions. Protein and mrna expression analysis Protein expression by Western blot analysis and RNA expression by semiquantitative and real-time RT PCR was performed by using standard procedures. Antibody and primer information is provided in the Supplemental Material. RT PCR splicing analysis Splicing analysis by RT PCR was performed using unique primers around potential splicing regions as described previously (Kalsotra et al. 2008). Histology, immunohistochemistry, Northern blotting, luciferase assay, and cardiac function analysis Details of histology, immunohistochemistry, Northern blotting, luciferase assay, and cardiac function analysis are provided in the Supplemental Material. Statistics Values are presented as mean 6 SD. Statistical significance was determined using two-tailed Student s t-test or one-way ANOVA followed by post hoc Tukey s multiple range test (P < 0.05). Acknowledgments We thank Dr. A. Tarakhovsky for providing Dicer f/f mice, Dr. J. Molkentin for providing MCM mice and heart tissues from PKCa-Tg mice, and Dr. M. Schneider and Dr. X. Wehrens for providing heart tissues from CycT1/Gqa- BiTg mice and TAC operated mice, respectively. We are also grateful to Dr. W. Claycomb for providing HL-1 cells, and to Dr. S. Anakk for help with luciferase assays. This project was funded by the NIH grant R01GM to T.A.C., Myotonic Dystrophy Foundation Postdoctoral award to A.K., and NNSFC grant G to P.F.L. References Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP The impact of micrornas on protein output. Nature 455: Bartel DP MicroRNAs: Target recognition and regulatory functions. Cell 136: Birchler JA, Kavi HH Molecular biology. Slicing and dicing for small RNAs. Science 320: Blencowe BJ Alternative splicing: New insights from global analyses. Cell 126: Boutz PL, Chawla G, Stoilov P, Black DL MicroRNAs regulate the expression of the alternative splicing factor nptb during muscle development. Genes & Dev 21: Braz JC, Gregory K, Pathak A, Zhao W, Sahin B, Klevitsky R, Kimball TF, Lorenz JN, Nairn AC, Liggett SB, et al PKC-a regulates cardiac contractility and propensity toward heart failure. Nat Med 10: Castle JC, Zhang C, Shah JK, Kulkarni AV, Kalsotra A, Cooper TA, Johnson JM Expression of 24,426 human alternative splicing events and predicted cis regulation in 48 tissues and cell lines. Nat Genet 40: Chen JF, Murchison EP, Tang R, Callis TE, Tatsuguchi M, Deng Z, Rojas M, Hammond SM, Schneider MD, Selzman CH, et al Targeted GENES & DEVELOPMENT 5
6 Kalsotra et al. deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure. Proc Natl Acad Sci 105: Claycomb WC, Lanson NA Jr, Stallworth BS, Egeland DB, Delcarpio JB, Bahinski A, Izzo NJ Jr HL-1 cells: A cardiac muscle cell line that contracts and retains phenotypic characteristics of the adult cardiomyocyte. Proc Natl Acad Sci 95: Cooper TA, Wan L, Dreyfuss G RNA and disease. Cell 136: da Costa Martins PA, Bourajjaj M, Gladka M, Kortland M, van Oort RJ, Pinto YM, Molkentin JD, De Windt LJ Conditional dicer gene deletion in the postnatal myocardium provokes spontaneous cardiac remodeling. Circulation 118: Ji Z, Lee JY, Pan Z, Jiang B, Tian B Progressive lengthening of 39 untranslated regions of mrnas by alternative polyadenylation during mouse embryonic development. Proc Natl Acad Sci 106: Kalsotra A, Xiao X, Ward AJ, Castle JC, Johnson JM, Burge CB, Cooper TA A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc Natl Acad Sci 105: Kim VN MicroRNA biogenesis: Coordinated cropping and dicing. Nat Rev Mol Cell Biol 6: Kuyumcu-Martinez NM, Wang GS, Cooper TA Increased steadystate levels of CUGBP1 in myotonic dystrophy 1 are due to PKCmediated hyperphosphorylation. Mol Cell 28: Lin Z, Murtaza I, Wang K, Jiao J, Gao J, Li PF mir-23a functions downstream of NFATc3 to regulate cardiac hypertrophy. Proc Natl Acad Sci 106: Makeyev EV, Zhang J, Carrasco MA, Maniatis T The microrna mir-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mrna splicing. Mol Cell 27: Olson EN Gene regulatory networks in the evolution and development of the heart. Science 313: Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ Deep surveying of alternative splicing complexity in the human transcriptome by highthroughput sequencing. Nat Genet 40: Sandberg R, Neilson JR, Sarma A, Sharp PA, Burge CB Proliferating cells express mrnas with shortened 39 untranslated regions and fewer microrna target sites. Science 320: Sano M, Abdellatif M, Oh H, Xie M, Bagella L, Giordano A, Michael LH, DeMayo FJ, Schneider MD Activation and function of cyclin T Cdk9 (positive transcription elongation factor-b) in cardiac muscle-cell hypertrophy. Nat Med 8: Selbach M, Schwanhausser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N Widespread changes in protein synthesis induced by micro- RNAs. Nature 455: Sohal DS, Nghiem M, Crackower MA, Witt SA, Kimball TR, Tymitz KM, Penninger JM, Molkentin JD Temporally regulated and tissuespecific gene manipulations in the adult and embryonic heart using a tamoxifen-inducible Cre protein. Circ Res 89: van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J, Olson EN Control of stress-dependent cardiac growth and gene expression by a microrna. Science 316: Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, Kingsmore SF, Schroth GP, Burge CB Alternative isoform regulation in human tissue transcriptomes. Nature 456: Xu X, Yang D, Ding JH, Wang W, Chu PH, Dalton ND, Wang HY, Bermingham JR Jr, Ye Z, Liu F, et al ASF/SF2-regulated CaMKIId alternative splicing temporally reprograms excitation-contraction coupling in cardiac muscle. Cell 120: Yi R, O Carroll D, Pasolli HA, Zhang Z, Dietrich FS, Tarakhovsky A, Fuchs E Morphogenesis in skin is governed by discrete sets of differentially expressed micrornas. Nat Genet 38: Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, Tsuchihashi T, McManus MT, Schwartz RJ, Srivastava D Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking mirna-1-2. Cell 129: GENES & DEVELOPMENT
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained
 Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Probe. Hind III Q,!?R'!! /0!!!!D1"?R'! vector. Homologous recombination
 Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Postn MCM Smad2 fl/fl Postn MCM Smad3 fl/fl Postn MCM Smad2/3 fl/fl. Postn MCM. Tgfbr1/2 fl/fl TAC
 A Smad2 fl/fl Smad3 fl/fl Smad2/3 fl/fl Tgfbr1/2 fl/fl 1. mm B Tcf21 MCM Tcf21 MCM Smad3 fl/fl Tcf21 MCM Smad2/3 fl/fl Tcf21 MCM Tgfbr1/2 fl/fl αmhc MCM C 1. mm 1. mm D Smad2 fl/fl Smad3 fl/fl Smad2/3
A Smad2 fl/fl Smad3 fl/fl Smad2/3 fl/fl Tgfbr1/2 fl/fl 1. mm B Tcf21 MCM Tcf21 MCM Smad3 fl/fl Tcf21 MCM Smad2/3 fl/fl Tcf21 MCM Tgfbr1/2 fl/fl αmhc MCM C 1. mm 1. mm D Smad2 fl/fl Smad3 fl/fl Smad2/3
c Ischemia (30 min) Reperfusion (8 w) Supplementary Figure bp 300 bp Ischemia (30 min) Reperfusion (4 h) Dox 20 mg/kg i.p.
 a Marker Ripk3 +/ 5 bp 3 bp b Ischemia (3 min) Reperfusion (4 h) d 2 mg/kg i.p. 1 w 5 w Sacrifice for IF size A subset for echocardiography and morphological analysis c Ischemia (3 min) Reperfusion (8
a Marker Ripk3 +/ 5 bp 3 bp b Ischemia (3 min) Reperfusion (4 h) d 2 mg/kg i.p. 1 w 5 w Sacrifice for IF size A subset for echocardiography and morphological analysis c Ischemia (3 min) Reperfusion (8
MicroRNA dysregulation in cancer. Systems Plant Microbiology Hyun-Hee Lee
 MicroRNA dysregulation in cancer Systems Plant Microbiology Hyun-Hee Lee Contents 1 What is MicroRNA? 2 mirna dysregulation in cancer 3 Summary What is MicroRNA? What is MicroRNA? MicroRNAs (mirnas) -
MicroRNA dysregulation in cancer Systems Plant Microbiology Hyun-Hee Lee Contents 1 What is MicroRNA? 2 mirna dysregulation in cancer 3 Summary What is MicroRNA? What is MicroRNA? MicroRNAs (mirnas) -
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
The role of mirnas in cardiac hypertrophy
 The role of mirnas in cardiac hypertrophy Fabian Pruissen Biology of Disease 2011 Supervised by Sjoukje Lok, MD and Roel de Weger, PhD Abstract Cardiac hypertrophy and increased cardiomyocytes size are
The role of mirnas in cardiac hypertrophy Fabian Pruissen Biology of Disease 2011 Supervised by Sjoukje Lok, MD and Roel de Weger, PhD Abstract Cardiac hypertrophy and increased cardiomyocytes size are
Supplemental Figure 1. Western blot analysis indicated that MIF was detected in the fractions of
 Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Supplemental Information. Menin Deficiency Leads to Depressive-like. Behaviors in Mice by Modulating. Astrocyte-Mediated Neuroinflammation
 Neuron, Volume 100 Supplemental Information Menin Deficiency Leads to Depressive-like Behaviors in Mice by Modulating Astrocyte-Mediated Neuroinflammation Lige Leng, Kai Zhuang, Zeyue Liu, Changquan Huang,
Neuron, Volume 100 Supplemental Information Menin Deficiency Leads to Depressive-like Behaviors in Mice by Modulating Astrocyte-Mediated Neuroinflammation Lige Leng, Kai Zhuang, Zeyue Liu, Changquan Huang,
Supplemental Information. Myocardial Polyploidization Creates a Barrier. to Heart Regeneration in Zebrafish
 Developmental Cell, Volume 44 Supplemental Information Myocardial Polyploidization Creates a Barrier to Heart Regeneration in Zebrafish Juan Manuel González-Rosa, Michka Sharpe, Dorothy Field, Mark H.
Developmental Cell, Volume 44 Supplemental Information Myocardial Polyploidization Creates a Barrier to Heart Regeneration in Zebrafish Juan Manuel González-Rosa, Michka Sharpe, Dorothy Field, Mark H.
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons
 Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003)
 MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
MicroRNAs: Genomics, Biogenesis, Mechanism, and Function (D. Bartel Cell 2004) he micrornas of Caenorhabditis elegans (Lim et al. Genes & Development 2003) Vertebrate MicroRNA Genes (Lim et al. Science
Chapter 2. Investigation into mir-346 Regulation of the nachr α5 Subunit
 15 Chapter 2 Investigation into mir-346 Regulation of the nachr α5 Subunit MicroRNA s (mirnas) are small (< 25 base pairs), single stranded, non-coding RNAs that regulate gene expression at the post transcriptional
15 Chapter 2 Investigation into mir-346 Regulation of the nachr α5 Subunit MicroRNA s (mirnas) are small (< 25 base pairs), single stranded, non-coding RNAs that regulate gene expression at the post transcriptional
SUPPLEMENTARY INFORMATION
 a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice
 Research article MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice Thomas E. Callis, 1,2 Kumar Pandya, 3 Hee Young Seok, 1,2 Ru-Hang Tang, 1,2,4 Mariko Tatsuguchi, 1,2 Zhan-Peng
Research article MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice Thomas E. Callis, 1,2 Kumar Pandya, 3 Hee Young Seok, 1,2 Ru-Hang Tang, 1,2,4 Mariko Tatsuguchi, 1,2 Zhan-Peng
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1. Baf60c and baf180 are induced during cardiac regeneration in zebrafish. RNA in situ hybridization was performed on paraffin
 Supplementary Figure 1. Baf60c and baf180 are induced during cardiac regeneration in zebrafish. RNA in situ hybridization was performed on paraffin sections from sham-operated adult hearts (a and i) and
Supplementary Figure 1. Baf60c and baf180 are induced during cardiac regeneration in zebrafish. RNA in situ hybridization was performed on paraffin sections from sham-operated adult hearts (a and i) and
MicroRNA therapeutics for cardiovascular disease. Eva van Rooij, PhD Hubrecht Institute, KNAW and University Medical Center Utrecht, The Netherlands
 Original Article Heart Metab. (2014) 65:4-8 for cardiovascular disease Eva van Rooij, PhD Hubrecht Institute, KNAW and University Medical Center Utrecht, The Netherlands Correspondence: Eva van Rooij,
Original Article Heart Metab. (2014) 65:4-8 for cardiovascular disease Eva van Rooij, PhD Hubrecht Institute, KNAW and University Medical Center Utrecht, The Netherlands Correspondence: Eva van Rooij,
Supporting Information
 Supporting Information Franco et al. 10.1073/pnas.1015557108 SI Materials and Methods Drug Administration. PD352901 was dissolved in 0.5% (wt/vol) hydroxyl-propyl-methylcellulose, 0.2% (vol/vol) Tween
Supporting Information Franco et al. 10.1073/pnas.1015557108 SI Materials and Methods Drug Administration. PD352901 was dissolved in 0.5% (wt/vol) hydroxyl-propyl-methylcellulose, 0.2% (vol/vol) Tween
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
Stress-dependent cardiac remodeling occurs in the absence of microrna-21 in mice
 Brief report Related Commentary, page 3817 Stress-dependent cardiac remodeling occurs in the absence of microrna-21 in mice David M. Patrick, 1 Rusty L. Montgomery, 2 Xiaoxia Qi, 1 Susanna Obad, 3 Sakari
Brief report Related Commentary, page 3817 Stress-dependent cardiac remodeling occurs in the absence of microrna-21 in mice David M. Patrick, 1 Rusty L. Montgomery, 2 Xiaoxia Qi, 1 Susanna Obad, 3 Sakari
International Graduate Research Programme in Cardiovascular Science
 1 International Graduate Research Programme in Cardiovascular Science This work has been supported by the European Community s Sixth Framework Programme under grant agreement n LSHM-CT-2005-01883 EUGeneHeart.
1 International Graduate Research Programme in Cardiovascular Science This work has been supported by the European Community s Sixth Framework Programme under grant agreement n LSHM-CT-2005-01883 EUGeneHeart.
Novel RNAs along the Pathway of Gene Expression. (or, The Expanding Universe of Small RNAs)
 Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Cardiac development begins when mesodermal progenitor
 Molecular Medicine MicroRNA-22 Regulates Cardiac Hypertrophy and Remodeling in Response to Stress Zhan-Peng Huang,* Jinghai Chen,* Hee Young Seok, Zheng Zhang, Masaharu Kataoka, Xiaoyun Hu, Da-Zhi Wang
Molecular Medicine MicroRNA-22 Regulates Cardiac Hypertrophy and Remodeling in Response to Stress Zhan-Peng Huang,* Jinghai Chen,* Hee Young Seok, Zheng Zhang, Masaharu Kataoka, Xiaoyun Hu, Da-Zhi Wang
Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated
 Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated with the H3K27me3 mark in chondrocytes (see Table S1,
Supplementary Figure 1 (Related with Figure 4). Molecular consequences of Eed deletion. (a) ChIP analysis identifies 3925 genes that are associated with the H3K27me3 mark in chondrocytes (see Table S1,
Supplemental Experimental Procedures
 Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
Table S1. Quantitative RT-PCR primers
 Table S1. Quantitative RT-PCR primers Gene Forward Primer Reverse Primer Human ApoB gcaagcagaagccagaagta ccatttggagaagcagtttgg Human ApoA1 gaaagctgcggtgctgac agtggccaggtccttcact Human MTP acggccattcccattgtg
Table S1. Quantitative RT-PCR primers Gene Forward Primer Reverse Primer Human ApoB gcaagcagaagccagaagta ccatttggagaagcagtttgg Human ApoA1 gaaagctgcggtgctgac agtggccaggtccttcact Human MTP acggccattcccattgtg
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Inhibition of mir-29 protects against cardiac hypertrophy and fibrosis: new insight for the role of mir-29 in the heart
 Editorial Page 1 of 6 Inhibition of mir-29 protects against cardiac hypertrophy and fibrosis: new insight for the role of mir-29 in the heart Julie R. McMullen 1,2,3,4,5, Bianca C. Bernardo 1,5,6 1 Baker
Editorial Page 1 of 6 Inhibition of mir-29 protects against cardiac hypertrophy and fibrosis: new insight for the role of mir-29 in the heart Julie R. McMullen 1,2,3,4,5, Bianca C. Bernardo 1,5,6 1 Baker
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway
 Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway S1 Oleksi Petrenko and Ute M. Moll Figure S1. MIF-Deficient Cells Have Reduced Transforming Ability (A) Soft
Supplemental Data Macrophage Migration Inhibitory Factor MIF Interferes with the Rb-E2F Pathway S1 Oleksi Petrenko and Ute M. Moll Figure S1. MIF-Deficient Cells Have Reduced Transforming Ability (A) Soft
Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility. in mice
 Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility in mice Xiaopeng Hu 1 *, Zhenzhou Tang 1,2 *, Yang Li 2 *, Wensheng Liu 2, Shuang Zhang 3, Bingyan Wang 3, Yingpu Tian 2, Yinan
Deletion of tyrosine phosphatase Shp2 in Sertoli cells causes infertility in mice Xiaopeng Hu 1 *, Zhenzhou Tang 1,2 *, Yang Li 2 *, Wensheng Liu 2, Shuang Zhang 3, Bingyan Wang 3, Yingpu Tian 2, Yinan
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.
 Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Kidney. Heart. Lung. Sirt1. Gapdh. Mouse IgG DAPI. Rabbit IgG DAPI
 a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
E10.5 E18.5 P2 10w 83w NF1 HF1. Sham ISO. Bmi1. H3K9me3. Lung weight (g)
 Myociyte cross-sectional Relative mrna levels Relative levels Relative mrna levels Supplementary Figures and Legends a 8 6 4 2 Ezh2 E1.5 E18.5 P2 1w 83w b Ezh2 p16 amhc b-actin P2 43w kd 37 86 16 wt mouse
Myociyte cross-sectional Relative mrna levels Relative levels Relative mrna levels Supplementary Figures and Legends a 8 6 4 2 Ezh2 E1.5 E18.5 P2 1w 83w b Ezh2 p16 amhc b-actin P2 43w kd 37 86 16 wt mouse
Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse
 Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
TITLE: MiR-146-SIAH2-AR Signaling in Castration-Resistant Prostate Cancer
 AWARD NUMBER: W81XWH-14-1-0387 TITLE: MiR-146-SIAH2-AR Signaling in Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr. Goberdhan Dimri, PhD CONTRACTING ORGANIZATION: George Washington University,
AWARD NUMBER: W81XWH-14-1-0387 TITLE: MiR-146-SIAH2-AR Signaling in Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr. Goberdhan Dimri, PhD CONTRACTING ORGANIZATION: George Washington University,
Neurotrophic factor GDNF and camp suppress glucocorticoid-inducible PNMT expression in a mouse pheochromocytoma model.
 161 Neurotrophic factor GDNF and camp suppress glucocorticoid-inducible PNMT expression in a mouse pheochromocytoma model. Marian J. Evinger a, James F. Powers b and Arthur S. Tischler b a. Department
161 Neurotrophic factor GDNF and camp suppress glucocorticoid-inducible PNMT expression in a mouse pheochromocytoma model. Marian J. Evinger a, James F. Powers b and Arthur S. Tischler b a. Department
Marta Puerto Plasencia. microrna sponges
 Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
Marta Puerto Plasencia microrna sponges Introduction microrna CircularRNA Publications Conclusions The most well-studied regions in the human genome belong to proteincoding genes. Coding exons are 1.5%
J Jpn Coll Angiol, 2009, 49:
 Online publication October 6, 2009 48 2 20 J Jpn Coll Angiol, 2009, 49: 293 297 atrial natriuretic peptide, brain natriuretic peptide, guanylyl cyclase-a receptor, cardiac remodeling, cardiac hypertrophy
Online publication October 6, 2009 48 2 20 J Jpn Coll Angiol, 2009, 49: 293 297 atrial natriuretic peptide, brain natriuretic peptide, guanylyl cyclase-a receptor, cardiac remodeling, cardiac hypertrophy
Post-transcriptional regulation of an intronic microrna
 Post-transcriptional regulation of an intronic microrna Carl Novina Dana-Farber Cancer Institute Harvard Medical School Broad Institute of Harvard and MIT Qiagen Webinar 05-17-11 Outline 1. The biology
Post-transcriptional regulation of an intronic microrna Carl Novina Dana-Farber Cancer Institute Harvard Medical School Broad Institute of Harvard and MIT Qiagen Webinar 05-17-11 Outline 1. The biology
Title: Smooth muscle cell-specific Tgfbr1 deficiency promotes aortic aneurysm formation by stimulating multiple signaling events
 Title: Smooth muscle cell-specific Tgfbr1 deficiency promotes aortic aneurysm formation by stimulating multiple signaling events Pu Yang 1, 3, radley M. Schmit 1, Chunhua Fu 1, Kenneth DeSart 1, S. Paul
Title: Smooth muscle cell-specific Tgfbr1 deficiency promotes aortic aneurysm formation by stimulating multiple signaling events Pu Yang 1, 3, radley M. Schmit 1, Chunhua Fu 1, Kenneth DeSart 1, S. Paul
AP VP DLP H&E. p-akt DLP
 A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
A B AP VP DLP H&E AP AP VP DLP p-akt wild-type prostate PTEN-null prostate Supplementary Fig. 1. Targeted deletion of PTEN in prostate epithelium resulted in HG-PIN in all three lobes. (A) The anatomy
Alterations in sarcomere function modify the hyperplastic to hypertrophic transition phase of mammalian cardiomyocyte development
 Alterations in sarcomere function modify the hyperplastic to hypertrophic transition phase of mammalian cardiomyocyte development Benjamin R. Nixon, 1 Alexandra F. Williams, 1 Michael S. Glennon, 1 Alejandro
Alterations in sarcomere function modify the hyperplastic to hypertrophic transition phase of mammalian cardiomyocyte development Benjamin R. Nixon, 1 Alexandra F. Williams, 1 Michael S. Glennon, 1 Alejandro
Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma CH Li, SC Chow, DL Yin,
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Functional characterisation of hepatitis B viral X protein/microrna-21 interaction in HBVassociated hepatocellular carcinoma CH Li, SC Chow, DL Yin,
Utility of Circulating micrornas in Cardiovascular Disease
 Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
Utility of Circulating micrornas in Cardiovascular Disease Pil-Ki Min, MD, PhD Cardiology Division, Gangnam Severance Hospital, Yonsei University College of Medicine Introduction Biology of micrornas Circulating
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
MicroRNAs, RNA Modifications, RNA Editing. Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM
 MicroRNAs, RNA Modifications, RNA Editing Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM Expanding world of RNAs mrna, messenger RNA (~20,000) trna, transfer
MicroRNAs, RNA Modifications, RNA Editing Bora E. Baysal MD, PhD Oncology for Scientists Lecture Tue, Oct 17, 2017, 3:30 PM - 5:00 PM Expanding world of RNAs mrna, messenger RNA (~20,000) trna, transfer
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Bi 8 Lecture 17. interference. Ellen Rothenberg 1 March 2016
 Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
hemodynamic stress. A. Echocardiographic quantification of cardiac dimensions and function in
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Fbn1 C1039G/+ hearts display normal cardiac function in the absence of hemodynamic stress. A. Echocardiographic quantification of cardiac dimensions and
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure 1. Fbn1 C1039G/+ hearts display normal cardiac function in the absence of hemodynamic stress. A. Echocardiographic quantification of cardiac dimensions and
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections
 Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections of normal cartilage were stained with safranin O-fast
Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections of normal cartilage were stained with safranin O-fast
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
MicroRNA in Cancer Karen Dybkær 2013
 MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
MicroRNA in Cancer Karen Dybkær RNA Ribonucleic acid Types -Coding: messenger RNA (mrna) coding for proteins -Non-coding regulating protein formation Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Targeted mass spectrometry (LC/MS/MS) for Olaparib pharmacokinetics. For LC/MS/MS of Olaparib pharmacokinetics metabolites were extracted from
 Supplementary Methods: Targeted mass spectrometry (LC/MS/MS) for Olaparib pharmacokinetics For LC/MS/MS of Olaparib pharmacokinetics metabolites were extracted from mouse tumor samples and analyzed as
Supplementary Methods: Targeted mass spectrometry (LC/MS/MS) for Olaparib pharmacokinetics For LC/MS/MS of Olaparib pharmacokinetics metabolites were extracted from mouse tumor samples and analyzed as
Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at
 Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at E10.5 were double-stained for TUNEL (red) and PECAM-1 (green).
Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at E10.5 were double-stained for TUNEL (red) and PECAM-1 (green).
Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions. 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19
 Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19 1 MicroRNA biogenesis 1. Pri mirna: primary mirna 2. Drosha: RNaseIII 3. DCR 1: Dicer 1 RNaseIII
Cellular MicroRNA and P Bodies Modulate Host-HIV-1 Interactions 指導教授 : 張麗冠博士 演講者 : 黃柄翰 Date: 2009/10/19 1 MicroRNA biogenesis 1. Pri mirna: primary mirna 2. Drosha: RNaseIII 3. DCR 1: Dicer 1 RNaseIII
SOPten flox/flox (KO) Pten flox/flox (WT) flox allele 6.0 kb. Pten. Actin. ! allele 2.3 kb. Supplementary Figure S1. Yanagi, et al.
 s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
s1 A Pten flox/flox () SOPten flox/flox () flox allele 6. kb B Pten flox/flox () SOPten flox/flox () Pten Actin! allele 2.3 kb Supplementary Figure S1. Yanagi, et al. A B BrdU BrdU positive cells ( ) 3
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Tcf21 MCM ; R26 mtmg Sham GFP Col 1/3 TAC 8W TAC 2W. Postn MCM ; R26 mtmg Sham GFP Col 1/3 TAC 8W TAC 2W
 A Tcf21 MCM ; R26 mtmg Sham GFP Col 1/3 Tcf21 MCM ; R26 mtmg TAC 2W Tcf21 MCM ; R26 mtmg TAC 8W B Postn MCM ; R26 mtmg Sham GFP Col 1/3 Postn MCM ; R26 mtmg TAC 2W Postn MCM ; R26 mtmg TAC 8W Supplementary
A Tcf21 MCM ; R26 mtmg Sham GFP Col 1/3 Tcf21 MCM ; R26 mtmg TAC 2W Tcf21 MCM ; R26 mtmg TAC 8W B Postn MCM ; R26 mtmg Sham GFP Col 1/3 Postn MCM ; R26 mtmg TAC 2W Postn MCM ; R26 mtmg TAC 8W Supplementary
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Table 1. List of primers used in this study
 Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
Supplementary Information
 Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer
 AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis
 Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Research Communication
 IUBMB Life, 64(7): 628 635, July 2012 Research Communication MicroRNA-181b Targets camp Responsive Element Binding Protein 1 in Gastric Adenocarcinomas Lin Chen*, Qian Yang*, Wei-Qing Kong*, Tao Liu, Min
IUBMB Life, 64(7): 628 635, July 2012 Research Communication MicroRNA-181b Targets camp Responsive Element Binding Protein 1 in Gastric Adenocarcinomas Lin Chen*, Qian Yang*, Wei-Qing Kong*, Tao Liu, Min
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10188 Supplementary Figure 1. Embryonic epicardial genes are down-regulated from midgestation stages and barely detectable post-natally. Real time qrt-pcr revealed a significant down-regulation
doi:10.1038/nature10188 Supplementary Figure 1. Embryonic epicardial genes are down-regulated from midgestation stages and barely detectable post-natally. Real time qrt-pcr revealed a significant down-regulation
Berberine Sensitizes Human Ovarian Cancer Cells to Cisplatin Through mir-93/ PTEN/Akt Signaling Pathway
 Chen Accepted: et al.: February Berberine 24, Sensitizes 2015 Ovarian Cancer Cells to Cisplatin www.karger.com/cpb 956 1421-9778/15/0363-0956$39.50/0 Original Paper This is an Open Access article licensed
Chen Accepted: et al.: February Berberine 24, Sensitizes 2015 Ovarian Cancer Cells to Cisplatin www.karger.com/cpb 956 1421-9778/15/0363-0956$39.50/0 Original Paper This is an Open Access article licensed
Supplementary Figure 1
 Supplementary Figure 1 Kif1a RNAi effect on basal progenitor differentiation Related to Figure 2. Representative confocal images of the VZ and SVZ of rat cortices transfected at E16 with scrambled or Kif1a
Supplementary Figure 1 Kif1a RNAi effect on basal progenitor differentiation Related to Figure 2. Representative confocal images of the VZ and SVZ of rat cortices transfected at E16 with scrambled or Kif1a
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Supplemental Information. Metabolic Maturation during Muscle Stem Cell. Differentiation Is Achieved by mir-1/133a-mediated
 Cell Metabolism, Volume 27 Supplemental Information Metabolic Maturation during Muscle Stem Cell Differentiation Is Achieved by mir-1/133a-mediated Inhibition of the Dlk1-Dio3 Mega Gene Cluster Stas Wüst,
Cell Metabolism, Volume 27 Supplemental Information Metabolic Maturation during Muscle Stem Cell Differentiation Is Achieved by mir-1/133a-mediated Inhibition of the Dlk1-Dio3 Mega Gene Cluster Stas Wüst,
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
MicroRNAs: Regulatory Function and Potential for Gene Therapy
 Lidia Park Genomics and Medicine November 23, 2008 MicroRNAs: Regulatory Function and Potential for Gene Therapy When Victor Ambros, Rosalind Lee, and Rhonda Feinbaum published the first finding of a microrna,
Lidia Park Genomics and Medicine November 23, 2008 MicroRNAs: Regulatory Function and Potential for Gene Therapy When Victor Ambros, Rosalind Lee, and Rhonda Feinbaum published the first finding of a microrna,
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
In vitro functional study of Cardiomyocyte
 In vitro functional study of Cardiomyocyte Gwang Hyeon Eom Department of Pharmacology and Medical Research Center for Gene Regulation Chonnam National University Medical School, Gwangju, South Korea Heart
In vitro functional study of Cardiomyocyte Gwang Hyeon Eom Department of Pharmacology and Medical Research Center for Gene Regulation Chonnam National University Medical School, Gwangju, South Korea Heart
