Supplementary Data A
|
|
|
- Griffin Joseph
- 5 years ago
- Views:
Transcription
1 Supplementary Data A Proteomic data and MS/MS fragmentation spectra from 2-DE and GeLC-MS/MS analysis of the bovine teat canal lining and the outer teat skin epithelium
2 Table A1: 2-DE proteins spots identified by nanolc-esi-q\tof MS\MS. Spot # Identified protein (species) NCBI accession code Predicted MW (kda) Predicted pi No. peptides matched Sequence coverage (%) 1 heat shock cognate 71 kda protein gi serum albumin precursor gi Chain A, Crystal Structure Of Bovine Serum Albumin gi serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 4-like gi PREDICTED: keratin, type II cytoskeletal 1 gi keratin, type I cytoskeletal 10 gi keratin, type I cytoskeletal 10 gi keratin, type I cytoskeletal 10 gi keratin, type I cytoskeletal 10 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 3 gi PREDICTED: keratin, type II cytoskeletal 3 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 3 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 3 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 3 gi Mowse score
3 18 PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: keratin, type II cytoskeletal 1 gi PREDICTED: SERPIN B3-like gi annexin A2 gi keratin, type I cytoskeletal 17 gi keratin, type II cytoskeletal 4 gi Chain A, The Structure Of Crystalline Profilin-Beta-Actin gi keratin, type II cytoskeletal 4 gi keratin, type I cytoskeletal 14 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 5 gi keratin, type II cytoskeletal 4 gi keratin 6A gi keratin, type II cytoskeletal 5 gi keratin, type II cytoskeletal 79 gi SERPIN B4 protein gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 5 gi PREDICTED: keratin, type II cytoskeletal 1 gi keratin, type II cytoskeletal 6A gi SERPIN B4 protein gi keratin, type II cytoskeletal 5 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 79 gi keratin 6A gi PREDICTED: keratin, type II cytoskeletal 1 gi
4 31 PREDICTED: SERPIN B3-like gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi keratin, type II cytoskeletal 6A gi keratin 6A gi keratin, type II cytoskeletal 79 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 6A gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi keratin 6A gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 6A gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi keratin 6A gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi keratin, type II cytoskeletal 6A gi PREDICTED: keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 79 gi EEF1A1 protein gi PREDICTED: SERPIN B3-like gi PREDICTED: SERPIN B4 gi PREDICTED: SERPIN B4-like gi
5 42 PREDICTED: SERPIN B4-like gi alpha S1 casein, partial gi caspase-14 gi protein sigma (ovis aries) gi PREDICTED: keratin, type II cytoskeletal 1 gi spot not identified 47 PREDICTED: SERPIN B3-like gi PREDICTED: SERPIN B3-like gi annexin I gi annexin A2 gi glyceraldehyde-3-phosphate dehydrogenase gi glyceraldehyde-3-phosphate dehydrogenase gi glyceraldehyde-3-phosphate dehydrogenase gi gasdermin-a gi heat shock protein beta-1 gi phosphoglycerate mutase 1 gi triosephosphate isomerase gi protein S100A9 gi protein S100A9 gi PREDICTED: non-specific cytotoxic cell receptor protein 1 homolog gi Chain X, The Cys121ser Mutant Of Beta-Lactoglobulin gi Chain X, The Cys121ser Mutant Of Beta-Lactoglobulin gi calmodulin gi caspase-14 gi PREDICTED: LOW QUALITY PROTEIN: aspartic peptidase, retroviral-like 1 gi calmodulin-like 5 gi fatty acid-binding protein, adipocyte gi
6 68 PREDICTED: galectin-7 gi fatty acid-binding protein, epidermal gi hemoglobin subunit beta gi Chain A, A Novel Allosteric Mechanism In Haemoglobin. gi protein S100A7 gi PREDICTED: protein S100A7-like gi protein S100A7 gi PREDICTED: protein S100A7-like gi protein S100A7 gi protein S100A7 gi PREDICTED: protein S100A7-like gi PREDICTED: protein S100A7-like gi protein S100A7 gi PREDICTED: protein S100A7-like gi PREDICTED: protein S100A7-like gi protein S100A7 gi S100 calcium binding protein A11 (calgizzarin) gi protein S100A8 gi protein S100A8 gi protein S100A8 gi protein S100A12 gi protein S100A12 gi Chain F, Crystal Structure Of Human Stam1 Vhs Domain In Complex With Ubiquitin gi PREDICTED: galectin-7 gi
7 Table A2: Quantitative changes in 43 differentially expressed teat canal lining and teat skin proteins by 2-DE (p<0.05). Group A - TCL Group B - Teat skin SSP Identified protein Average Spot intensity CV (%) Average Spot intensity CV (%) Fold change p-value (n=3) 105 calmodulin-like protein sigma S100A keratin, type I cytoskeletal S100A S100A caspase S100A7-like keratin, type I cytoskeletal S100A S100A7-like S100A7-like S100A keratin, type II cytoskeletal keratin, type II cytoskeletal keratin, type II cytoskeletal E keratin, type II cytoskeletal keratin, type II cytoskeletal galectin keratin, type II cytoskeletal keratin, type II cytoskeletal
8 5613 keratin, type II cytoskeletal keratin, type II cytoskeletal S100A S100A S100A SERPIN B4-like keratin, type II cytoskeletal 6A keratin, type II cytoskeletal 6A keratin, type II cytoskeletal 6A SERPIN B4-like keratin, type II cytoskeletal 6A keratin, type II cytoskeletal 6A keratin, type II cytoskeletal keratin, type II cytoskeletal keratin, type II cytoskeletal hemoglobin subunit beta hemoglobin subunit alpha keratin, type II cytoskeletal 6A keratin, type II cytoskeletal 6A keratin, type II cytoskeletal keratin, type II cytoskeletal keratin, type II cytoskeletal Spot intensity is an average optical density (OD) of the same spot from three replicate samples belonging to the same experimental group. Coefficient of variation (CV) shows spot variation across these three samples. Fold change is calculated from spot intensity with teat skin as the denominator.
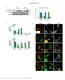
9 Figure A1: PDQuest histograms of 2-DE spot intensities for differentially expressed teat canal lining and teat skin protein spots. The standard spot (SSP) number is displayed beneath each histogram. The number at the upper right hand corner of the histogram is the normalised quantity of the maximum bar in the graph. The other bars are drawn proportionally to the highest bar. Intensity bars from individual teat canal lining and teat skin spots are labelled in green and red, respectively.
10 Table A3: Summary of total MS/MS spectra from each tryptic digest fraction for the teat canal lining group from 6 cows # Fraction # Average SD Total % CV 8.1% SD, standard deviation; CV, coefficient of variation Table A4: Summary of total MS/MS spectra from each tryptic digest fraction for the teat skin group from 6 cows # Fraction # Average SD Total % CV 8.4% SD, standard deviation; CV, coefficient of variation
11 Table A5: Proteins with 2 or more peptides identified by GeLC-MS/MS from TCL and teat skin replicate samples at 5 % FDR Total unique peptide count Teat canal lining Teat skin Identified proteins (species) Accession MW [kda] pi 005 keratin, type II cytoskeletal 1 gi keratin, type I cytoskeletal 10 gi keratin, type II cytoskeletal 6A gi keratin, type II cytoskeletal 6A isoform 3 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 5 gi keratin, type I cytoskeletal 14 gi keratin, type II cytoskeletal 79 gi keratin, type I cytoskeletal 17 gi keratin, type II cytoskeletal 3 gi profilaggrin-like gi SERPIN B3-like gi SERPIN B3-like gi SERPIN B4-like protein gi SERPIN B3-like gi filaggrin gi SERPIN B3-like gi bovine serum albumin gi SERPIN B3 gi protein sigma (ovis aries) gi caspase-14 gi junction plakoglobin gi S100A12 gi
12 S100A9 gi keratin, type II cytoskeletal 2 oral gi annexin A1 gi beta-lactoglobulin gi protein zeta gi hemoglobin subunit beta gi hornerin gi alpha-s1-casein precursor gi desmoplakin gi keratin, type II cytoskeletal 2 epidermal gi ubiquitin gi fatty acid-binding protein, epidermal gi pancreatic adenocarcinoma upregulated factor-like gi glyceraldehyde-3-phosphate dehydrogenase gi nucleoside diphosphate kinase B gi S100A8 gi alpha-s2-casein precursor gi S100A7 gi beta-casein precursor gi S100A7-like gi malate dehydrogenase, cytoplasmic gi triosephosphate isomerase gi actin, beta gi hemoglobin subunit alpha gi gasdermin 1 gi annexin A2 gi heat shock cognate 71 kda protein gi IGL@ protein gi L-lactate dehydrogenase A chain gi
13 tropomyosin 4-like gi casein para kappaa gi repetin gi S100A2 gi transitional endoplasmic reticulum ATPase isoform 3 (canis lupus familiaris) gi SERPIN B12 gi heat shock 27kDa protein 1 gi insulin-degrading enzyme isoform X1 (IDE) gi proteasome subunit beta type-2 isoform 2 gi ganglioside GM2 activator precursor gi retroviral-like aspartic protease 1 gi annexin A8 gi purine nucleoside phosphorylase, chain A gi chloride intracellular channel protein 1 gi collagen, type VI, alpha 3-like isoform 4 gi F-box only protein 50 gi tubulin alpha-1b chain (mus musculus) gi alpha-actinin 1 gi calmodulin-like 5 gi IgM heavy chain constant region gi thioredoxin gi elongation factor 2 gi fatty acid-binding protein, adipocyte gi galectin-7 gi peptidylprolyl isomerase A gi cystatin-a gi histone H2A type 2-C gi histone H2B gi vimentin gi
14 peroxiredoxin-1 gi apolipoprotein A-I precursor gi elongation factor 1 alpha gi profilin gi protein DJ-1 gi S100A11 (calgizzarin) gi S100A14 gi protein beta/alpha (homo sapiens) gi protein epsilon (homo sapiens) gi S ribosomal protein S7 (homo sapiens) gi S ribosomal protein L11 isoform 2 (homo sapiens) gi S ribosomal protein L12 isoform X1 gi ADP-ribosylation factor 3 gi AHNAK nucleoprotein 2 gi aldose reductase gi enolase (alpha) gi angiogenin gi annexin A4 gi annexin V=CaBP33 isoform [cattle, brain, peptide, 320 aa] gi band 6 polypeptide B6P, partial gi basigin precursor gi beta-enolase gi cofilin-1 (sus scrofa) gi collagen alpha-2(vi) chain isoform X1 gi cornulin-like gi cullin 1-like gi cystatin-b gi desmocollin gi
15 desmoglein gi desmosomal cadherin gi epiplakin gi F-actin-binding protein gi fatty acid synthase isoform X1 gi gamma-glutamylcyclotransferase gi gelsolin isoform b gi glutathione S-transferase Mu 1 isoform X1 gi glutathione S-transferase P gi heat shock protein HSP 90-alpha gi heat shock protein HSP 90-beta gi histone H2A type 1 gi histone H3 family 3A gi histone H4 gi keratin 15 gi keratin 77-like gi keratin, type I cytoskeletal 16 gi keratin, type I cytoskeletal 19 gi keratin, type I cytoskeletal 27 gi keratin, type II cytoskeletal 73 gi lactoferrin gi L-lactate dehydrogenase B chain gi mediator of RNA polymerase II transcription subunit 1 gi mitochondrial malate dehydrogenase 2, NAD gi myosin regulatory light chain 12B-like isoform 2 gi macrophage migration-inhibitory factor [cattle, Peptide, 114 aa] gi peroxiredoxin-2 gi phosphatidylethanolamine binding protein gi
16 plakophilin-1 gi plasmalemmal porin gi putative RNA-binding protein 3 gi ras-related protein Rab-10 (mus musculus) gi ras-related protein Rab-11A isoform X1 gi ras-related protein Rab-1A gi ras-related protein Rab-1B gi ribosomal protein L12 gi secernin-3 gi SERPIN B13 isoform 1 gi suprabasin isoform X1 gi titin gi transgelin-2 gi tyrosine 3/tryptophan 5 -monooxygenase activation protein gi ZBTB42 protein-like gi zinc finger protein 638 gi zymogen granule protein 16 homolog B gi Cells highlighted in grey correspond to biological replicates (n = 6) where no peptides were identified in either the TCL or teat skin group.
17 (A) (B) Figure A2: Representative MS/MS spectra identifying the two diagnostic peptides for S100A7. (A) Fragmentation pattern for precursor ion at m/z ; sequence: DNFPNFLGACEK. (B) Fragmentation pattern for precursor ion at m/z ; sequence: GRDYLSNIFEK. Peaks corresponding to b (red) and y (blue) ions are marked. Diagnostic amino acids detected are labelled with an asterisk.
18 (A) K/ ENFPNFLSACEK /R (B) R/ QYLSDIFEK /K Figure A3: Representative MS/MS spectra identifying the two diagnostic peptides for S100A7-like protein. (A) Fragmentation pattern for precursor ion at m/z ; sequence: ENFPNFLSACEK. (B) Fragmentation pattern for precursor ion at m/z ; sequence: QYLSDIFEK. Peaks corresponding to b (red) and y (blue) ions are marked. Diagnostic amino acids detected are labelled with an asterisk.
19 Table A6: Differential expression of keratin proteins in the teat canal lining and teat skin proteomes. Average peptide count (n=6) Identified keratin protein Teat canal lining Teat skin Fold change p-value keratin, type II cytoskeletal x 10-5 keratin, type II cytoskeletal 2 epidermal x keratin, type II cytoskeletal 2 oral x 10-2 keratin, type II cytoskeletal x 10-4 keratin, type II cytoskeletal x 10-7 keratin, type II cytoskeletal x 10-4 keratin, type II cytoskeletal 6A x 10-4 keratin, type II cytoskeletal 6A isoform * 0.01 keratin, type I cytoskeletal x 10-3 keratin, type I cytoskeletal x 10-4 keratin * keratin, type I cytoskeletal * 0.41 keratin, type I cytoskeletal * 0.02 keratin, type I cytoskeletal * keratin, type I cytoskeletal * keratin, type II cytoskeletal * keratin 77-like 0.5* keratin, type II cytoskeletal * 0.02 * Proteins absent in either of the two samples were given a value of 0.5 so that a fold change could be calculated. Fold change was calculated from the average peptide count with TCL as the denominator. p-values were derived from the six teat canal lining and six teat skin samples and were calculated as a two-tailed test with even distribution.
20 Table A7: Proteins identified in all 12 GeLC samples from teat canal lining and teat skin Average peptide count Identified proteins Accession number Teat canal lining Teat skin Fold change Keratins keratin, type II cytoskeletal 1 gi keratin, type I cytoskeletal 10 gi keratin, type II cytoskeletal 2 epidermal gi keratin, type II cytoskeletal 2 oral gi keratin, type II cytoskeletal 3 gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 5 gi keratin, type II cytoskeletal 6A gi keratin, type I cytoskeletal 14 gi Cytoskeleton desmoplakin gi hornerin gi junction plakoglobin gi filaggrin gi profilaggrin-like gi tubulin alpha-1b chain (mus musculus) gi Major milk proteins alpha-s1-casein precursor gi beta-casein precursor gi beta-lactoglobulin gi S100 calcium-binding proteins S100A7 gi S100A7-like gi S100A8 gi S100A9 gi S100A12 gi Cellular proteins actin, beta gi annexin A1 gi annexin A2 gi ubiquitin gi fatty acid-binding protein, epidermal gi heat shock 27kDa protein 1 gi L-lactate dehydrogenase A chain gi malate dehydrogenase, cytoplasmic gi thioredoxin gi transitional endoplasmic reticulum ATPase gi isoform 3 (canis lupus familiaris) Serum proteins bovine serum albumin gi hemoglobin subunit alpha gi hemoglobin subunit beta gi Apoptosis galectin-7 gi caspase-14 gi gasdermin 1 gi Fold change is calculated from average peptide counts with teat skin counts used as the denominator.
21 Supplementary Data B Experiments to determine monoclonal and polyclonal antibody specificity
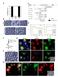
22 B1: Immunohistochemical staining of super mammary lymph node. Figure B2: Immune cell staining of super mammary lymph node tissue. Cryosections of super mammary lymph node tissues were probed with anti-mhc class II monoclonal antibody (a), anti-cd205 monoclonal antibody (b), and anti-cd3 monoclonal antibody (c). The bound anti-mhc class II antibody was detected using Alexa Fluor-594-IgG2a while the anti-cd205 and anti-cd3 antibodies were detected with Alexa Fluor-594-IgG (red signal). Cryosections probed with anti-cd8 monoclonal antibody (d), and anti-cd14 monoclonal antibody (e) were detected with Alexa Fluor-488-IgM labelled secondary antibody (green signal). Nuclei were counterstained with DAPI (blue signal). Section thickness: 5 µm. Scale bar = 100 µm.
23 Supplementary Data C Proteomic data and MS/MS fragmentation spectra from 2-DE and GeLC-MS/MS analysis of late-lactating teat canal lining and 14 d involution teat canal lining
24 Figure C1: PDQuest histograms of 2-DE spot intensities for differentially expressed latelactating and 14 d involution teat canal lining protein spots. The standard spot (SSP) number is displayed beneath each histogram. The number at the upper right hand corner of the histogram is the normalised quantity of the maximum bar in the graph. The other bars are drawn proportionally to the highest bar. Intensity bars from individual latelactating and 14 d involution teat canal lining spots are labelled in red and green, respectively.
25 Table C1: Summary of LC-MS/MS results for the four spiked proteins in E. coli lysate a. Gel slice SpC Peptide count Sequence coverage Conalbumin fmol Ave SD SEM fmol Ave SD SEM fmol Ave SD SEM SpC Peptide count Sequence coverage Ovalbumin fmol Ave SD SEM fmol Ave SD SEM fmol Ave SD SEM SpC Peptide count Sequence coverage Carbonic fmol Ave SD SEM fmol Ave SD SEM fmol Ave SD SEM Anhydrase SpC Peptide count Sequence coverage Lysozyme fmol Ave SD SEM fmol Ave SD SEM fmol Ave SD SEM SpC, spectral count; Ave, average; SD, standard deviation; SEM, standard error of the mean
26 (A) (B) (C) Figure C2: Correlation between SpC, peptide count and sequence coverage with protein abundance level. In each experiment four non- E. coli proteins were added to E. coli lysate at different concentrations representing different protein abundance levels. For all added proteins, a linear correlation was observed between SpC and relative protein abundance (A). Regression coefficients decreased between peptide count and relative protein abundance (B) and sequence coverage and relative protein abundance (C). The results are an average of four technical replicates. Error bars represent the standard error of the mean.
27 Table C2: Summary of LC-MS/MS results for the E. coli proteins in each of the five dilution series lysates. The results are from the four technical replicates. Abbreviations used for E. coli proteins are as follows: PFL, pyruvate formate-lyase [2PFL_A]; DnaK, chaperone protein DnaK [ZP_ ]; TEF Tu, translation elongation factor Tu [EKW71632]; PPH, phosphopyruvate hydratase [ZP_ ]; Oxidoreductase, [ZP_ ]; OMP A, outer membrane protein A [ZP_ ]; Global, global DNA-binding transcriptional dual regulator H-NS [WP_ ]; Stress F, universal stress protein F [WP_ ]. Overall coefficient of variation (CV) was 14.9 %. Gel slice SpC (PFL) SpC (DnaK) Conalbumin fmol Ave. SD SEM %CV fmol Ave. SD SEM %CV % % % % % % % % % % SpC (TEF Tu) SpC (PPH) Ovalbumin fmol Ave. SD SEM %CV fmol Ave. SD SEM %CV % % % % % % % % % % SpC (Oxireductase) SpC (OMP A) Carbonic fmol Ave. SD SEM %CV fmol Ave. SD SEM %CV Anhydrase % % % % % % % % % % SpC (Global) SpC (Stress F) Lysozyme fmol Ave. SD SEM %CV fmol Ave. SD SEM %CV % % % % % % % % % % SpC, spectral count; Ave, average; SD, standard deviation; SEM, standard error of the mean; CV, coefficient of variation
28 (A) (B) (C) (D) Figure C3: Correlation between E. coli protein SpC with added protein abundance levels. In each experiment two of the most abundant E. coli proteins were identified and their SpC plotted relative to the concentration of the added non-e. coli proteins. For all E. coli proteins, a nonlinear correlation was observed between SpC and relative added protein abundance for conalbumin (A), ovalbumin (B), carbonic anhydrase (C), and lysozyme (D). The results are an average of four technical replicates. Error bars represent the standard error of the mean. Abbreviations used for E. coli proteins are as follows: PFL, pyruvate formate-lyase [2PFL_A]; DnaK, chaperone protein DnaK [ZP_ ]; TEF Tu, translation elongation factor Tu [EKW71632]; PPH, phosphopyruvate hydratase [ZP_ ]; Oxidoreductase, [ZP_ ]; OMP A, outer membrane protein A [ZP_ ]; Global, global DNA-binding transcriptional dual regulator H-NS [WP_ ]; Stress F, universal stress protein F [WP_ ].
29 Table C3: Summary of total MS/MS spectra from each tryptic digest fraction for the 14 d involution teat canal lining group from 7 cows Fraction # Average SD Total % CV 8.1% SD, standard deviation; CV, coefficient of variation
30 Table C4: 86 proteins with 2 or more peptides identified by GeLC-MS/MS from 14 d TCL replicate samples at 5 % FDR Identified proteins Accession MW [kda] pi Total unique peptide count 14 d Involution TCL keratin, type II cytoskeletal 1 gi keratin, type I cytoskeletal 10 gi keratin, type II cytoskeletal 6A gi keratin, type II cytoskeletal 4 gi keratin, type II cytoskeletal 79 gi keratin, type II cytoskeletal 3 gi keratin, type II cytoskeletal 6C gi keratin, type I cytoskeletal 14 gi keratin, type II cytoskeletal 5 gi profilaggrin-like gi keratin, type I cytoskeletal 17 gi filaggrin-like gi filaggrin-like, partial gi serpin B3-like gi serpin B3-like gi serpin B4-like protein gi serpin B3-like gi lactoferrin gi keratin, type II cytoskeletal 2, oral gi serum albumin precursor gi
31 keratin, type II cytoskeletal 2, epidermal gi annexin A1 gi protein S100A9 gi protein S100A12 gi annexin A2 gi glyceraldehyde-3-phosphate dehydrogenase gi caspase-14 gi beta-lactoglobulin gi hemoglobin subunit beta gi hornerin gi cytokeratin 8 (370 AA) gi alpha S1 casein, partial gi protein S100A7 gi hemoglobin alpha chain gi protein S100A8 gi protein S100A7-like gi ubiquitin gi imunoglobulin light chain, lambda gi fatty acid-binding protein, epidermal gi desmoplakin gi hornerin-like gi beta-casein gi gasdermin 1 gi malate dehydrogenase, cytoplasmic gi IgM heavy chain constant region gi F-box only protein 50 gi protein S100A2 gi
32 cystatin-a gi cathelicidin-1 precursor gi nucleoside diphosphate kinase B gi transitional endoplasmic reticulum ATPase 3 gi serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 12-like gi lactate dehydrogenase-a gi S100 calcium binding protein A11 gi beta actin gi L-lactate dehydrogenase B chain gi serotransferrin precursor gi pancreatic adenocarcinoma upregulated factor-like gi proteasome subunit beta type-2 isoform 2 gi mammalian 20s proteasome gi protein DJ-1 gi lactoperoxidase precursor gi alpha-s2-casein precursor gi casein para kappaa gi IgG1 heavy chain constant region, partial gi thioredoxin gi junction plakoglobin isoform X1 gi apolipoprotein A-I precursor gi histone H2B gi serpin B3-like gi
33 serpin B3 gi hypothetical protein BOS_17317 gi keratin, type I cytoskeletal 16 gi keratin, type II cytoskeletal 78 gi keratin, type I cytoskeletal 24 gi proteasome subunit alpha type-7 isoform X3 gi protein S100A14 gi polyubiquitin-c gi cathepsin O preproprotein-like gi heat shock cognate 71 kda protein gi protein sigma gi zymogen granule protein 16 homolog B gi galectin-7 gi tropomyosin alpha-3 chain isoform 8 gi heat shock 27kDa protein 1 gi cyclophilin I gi
34 Table C5: Normalised total SpCs of 25 proteins showing two-fold change in protein abundance between late-lactating and 14 d involution TCL (p < 0.05) Late-lactating TCL 14 d involution TCL Identified proteins LL Ave d14 Ave p-value Fc lactoferrin vimentin cofilin glyceraldehyde-3-phosphate dehydrogenase fatty acid binding protein, epithelial repetin transgelin purine nucleoside phosphorylase histone H2A type 1-B/E tubulin alpha-1b chain histone H2B type 1-D collagen, type VI, alpha 3-like isoform nucleoside diphosphate kinase B L-lactate dehydrogenase A chain peroxiredoxin
35 Table B5 continued: protein sigma Junction plakoglobin insulin-degrading enzyme precursor protein zeta peptidylprolyl isomerase A aspartic peptidase, retroviral-like galectin heat shock protein beta heat shock cognate 71 kda protein beta-actin Fc, fold count
36 Supplementary Data D Microbiological methods and proteomic GeLC-MS/MS analysis of the bovine teat canal lining 24 h post bacterial challenge
37 D1 Preparation of cultures for teat canal inoculation The extent of turbidity, or optical density (OD), was used to measure the cell biomass indirectly as the bacteria grew in liquid culture. Two cultures were made for each of the three microbes. One culture for measuring OD and the other culture for glycerol stocks. All three bacterial cultures exhibited standard, in vitro, temporal bacterial growth curves with characteristic lag, exponential and stationary phases (Figure D1). Both the E.coli and S. aureus cultures were stopped and harvested for glycerol-stock storage at late-log phase when their absorbance reading reached an OD of 0.8. S. uberis not only grew slower than the other two cultures, but its stationary phase was also lower (OD of 0.4). Based on this growth curve, the S. uberis culture was stopped and 0.5 ml aliquots harvested for glycerol-stock storage at late-log phase when its absorbance reading reached an OD of Glycerol stocks of bacteria were stored at -80 C until required. In preparation for teat infusion, bacteria were recovered from frozen glycerol stock solutions, washed in 1 ml of one-quarter strength ringer s solution containing 0.1 % (w/v) proteose-peptone for 1 h at 37 C. One hundred microlitres of this suspension was diluted into 9.9 ml of sterile skim milk. After 16 h growth at 37 C, ten-fold serial dilutions were performed to estimate the bacterial concentration in the skim milk suspension (See Methods Figure 2.2). The suspensions were then adjusted by dilution to give final inoculum concentrations of 2.5 x 10 8 cfu/ml (~500,000 cfu in 2 µl of skim milk), 1.25 x 10 6 cfu/ml (~2500 cfu in 2 µl of skim milk), and 1.25 x 10 6 cfu/ml (~2500 cfu in 2 µl of skim milk), for E. coli, S. aureus, and S, uberis, respectively. Minimal cell death (< 3 %) was observed when the stock skim milk suspensions were stored overnight at 4 C (data not shown). Stock skim milk suspensions were therefore made two days preceding the on-farm inoculations, allowing for a 16 h growth phase and then a 24 h storage phase. This storage phase enabled an accurate bacterial count for each inoculum to be determined prior to administration (Table D1). Dilutions of the inoculums were also replated after the on-farm inoculations. The results from this analysis showed negligible change (< 5.0 %) from the preceding dilution series (data not shown).
38 Figure D1: Growth curves for E.coli, S. aureus and S. uberis in brain heart infusion broth.
39 Table D1: Bacterial concentrations for teat canal inoculations. Day 1 Colony counts prior to teat canal inoculations Bacteria Number of colonies Average Dilution cfu a /ml SD b E. coli TMTC c TMTC TMTC 1 x 10-3 TMTC TMTC TMTC 1 x x x x x 10-6 S. uberis TMTC TMTC TMTC 1 x x x x x x 10-5 S. aureus TMTC TMTC TMTC 1 x x x x x x 10-5 Day 2 Colony counts prior to teat canal inoculations Bacteria Number of colonies Average Dilution cfu/ml SD E. coli TMTC TMTC TMTC 1 x x x x x x 10-7 S. uberis TMTC TMTC TMTC 1 x x x x x x 10-5 S. aureus TMTC TMTC TMTC 1 x x x x x x 10-5 a) cfu, colony forming units; b) SD, standard deviation; c) TMTC, Too many to count
40 Table D2: Significantly altered 2-DE proteins spots from Control verses E. coli infection groups SSP Con-1 Con-2 Con-3 E. coli-1 E. coli-2 E. coli-3 (Ave) Con SD (Ave) E. coli SD Fc p-value (n = 3) Identified protein caspase lactoglobulin keratin actin heat shock protein beta spot not identified serpin B3-like serpin B3-like serum albumin S100A serpin B3-like keratin 4/ keratin keratin 6A ubiquitin hemoglobin subunit beta spot not identified
41 Table D3: Significantly altered 2-DE proteins spots from Control verses S. aureus infection groups SSP Con-1 Con-2 Con-3 S.aureus-1 S.aureus-2 S.aureus-3 (Ave) Con SD (Ave) S. aureus SD Fc p-value Identified protein caspase spot not identified heat shock cognate 71 kda protein keratin 4/ keratin serum albumin spot not identified serpin B3-like keratin ubiquitin beta hemoglobin spot not identified spot not identified keratin 6A non-specific cytotoxic cell receptor protein 1 homolog
42 Table D4: Significantly altered 2-DE proteins spots from Control verses S. uberis infection groups (Ave) (Ave) SSP Con-1 Con-2 Con-3 S.uberis-1 S.uberis-2 S.uberis-3 SD SD Fc p-value Identified protein Con S. uberis spot not identified spot not identified spot not identified spot not identified caspase spot not identified lactoglobulin keratin S100A serum albumin spot not identified keratin 4/ keratin keratin spot not identified ubiquitin S100A spot not identified spot not identified annexin A non-specific cytotoxic cell receptor protein 1 homolog keratin 1
43 Table D5: Summary of total MS/MS spectra from each teat canal lining tryptic digest fraction from the bacterial challenge study group of 4 cows # Treatment Con EC SA SU Con EC SA SU Quarter FL BL FR BR FL BR FR BL Fraction Fraction Fraction Fraction Fraction Fraction Fraction Fraction Total # Treatment Con EC SA SU Con EC SA SU Quarter FR BR FL BL FR BR BL FL Fraction Fraction Fraction Fraction Fraction Fraction Fraction Fraction Total Fraction 1 Fraction 2 Fraction 3 Fraction 4 Fraction 5 Fraction 6 Fraction 7 Fraction 8 Average SD % CV SD, standard deviation; CV, coefficient of variation
44 Table D6: Normalised raw SpC data for experimentally induced bacterial challenge study # Protein Acc. No. Normalised SpCs Control E. coli S. aureus S. uberis Ratio Protein Name Mwt gi keratin, type II cytoskeletal 1 63 kda gi keratin, type I cytoskeletal kda gi keratin, type II cytoskeletal 6A 61 kda gi cytokeratin type II, component IV isoform 2 59 kda gi keratin, type II cytoskeletal 3 65 kda gi keratin, type II cytoskeletal 4 58 kda gi keratin, type II cytoskeletal 5 63 kda gi keratin, type II cytoskeletal kda gi profilaggrin-like 56 kda gi keratin, type II cytoskeletal 2 (epidermal) 64 kda gi keratin, type I cytoskeletal kda gi filaggrin-like 180 kda gi S100-A9 16 kda gi S100-A12 11 kda gi serpin B3-like 44 kda gi keratin, type I cytoskeletal kda gi alpha S1 casein, partial 24 kda gi serpin B3-like 44 kda gi serpin B3-like 42 kda gi serpin B4 44 kda gi hornerin 175 kda gi S100-A8 10 kda gi hemoglobin beta-a chain 16 kda gi caspase kda gi junction plakoglobin 82 kda gi beta-lactoglobulin 18 kda E. coli /Con S. aureus /Con S. uberis /Con
45 27 gi desmoplakin 332 kda gi serum albumin precursor 69 kda gi S100-A7 12 kda gi S100-A7 isoform X2 12 kda gi nucleoside diphosphate kinase B 17 kda gi hemoglobin subunit alpha 15 kda gi hornerin-like 164 kda gi glyceraldehyde-3-phosphate dehydrogenase 36 kda gi fatty acid binding protein, epidermal 15 kda gi protein sigma 28 kda gi kappa casein 18 kda gi gasdermin-a 50 kda gi beta casein 25 kda gi galectin-7-like 15 kda gi alpha-s2-casein precursor 26 kda gi immunoglobulin mu heavy chain constant region 50 kda gi IGL@ protein 25 kda gi fatty acid-binding protein, adipocyte 15 kda gi polyubiquitin-b 34 kda gi histone H2A type 1-like 34 kda gi protein zeta/delta 28 kda gi retroviral-like aspartic protease 1 34 kda gi cystatin-a 11 kda gi S100-A14 11 kda gi histone H2B type 1-D 14 kda gi heat shock protein beta-1 22 kda gi S100-A2 11 kda gi 74 annexin I 39 kda gi beta-actin 42 kda gi malate dehydrogenase, cytoplasmic 36 kda gi proteasome subunit beta type-2 23 kda
46 58 gi F-box only protein kda gi ganglioside GM2 activator precursor 21 kda gi L-lactate dehydrogenase A chain 37 kda gi protein DJ-1 20 kda gi annexin A2 39 kda gi keratin, type II cytoskeletal kda gi pancreatic adenocarcinoma upregulated factor-like 17 kda gi alpha-actinin kda gi peroxiredoxin 1 22 kda gi peptidylprolyl isomerase A 17 kda gi thioredoxin 12 kda gi repetin 78 kda gi serpin B12 46 kda gi insulin-degrading enzyme precursor 118 kda gi desmoglein-1 precursor 112 kda gi transitional endoplasmic reticulum ATPase 89 kda gi triosephosphate isomerase 27 kda gi heat shock cognate 71 kda protein 71 kda gi PSMA7 protein 27 kda gi tropomyosin alpha-3 chain isoform 4 29 kda gi L-lactate dehydrogenase B chain 37 kda gi profilin 15 kda gi beta2-microglobulin 12 kda gi AHNAK nucleoprotein kda gi serotransferrin precursor 78 kda gi ezrin 69 kda gi myosin light polypeptide 6 isoform 1 17 kda gi cornulin-like 87 kda gi fibrinogen gamma-b chain isoform X1 49 kda
47 87 gi peptidoglycan recognition protein 1 21 kda gi ras-related protein Rab kda gi ATP-dependent RNA helicase DDX3X 73 kda gi creatine kinase M chain 43 kda gi prelamin-a/c isoform X1 74 kda gi S ribosomal protein S20 isoform 2 13 kda gi putative RNA-binding protein 3 10 kda gi protein POF1B 57 kda gi cathelicidin-1 precursor 18 kda gi myoglobin 17 kda gi lactoferrin 78 kda gi elongation factor 2 95 kda gi calmodulin-like isoform 2 17 kda gi xanthine dehydrogenase 147 kda gi histone H kda gi glutathione S-transferase P 24 kda gi apolipoprotein A-I precursor 30 kda gi S ribosomal protein L11 20 kda gi keratin, type I cytoskeletal kda gi heat shock protein HSP 90- alpha isoform X1 89 kda gi tripeptidyl-peptidase kda gi ribosomal protein L12 16 kda gi pyruvate kinase isozymes M1/M2 58 kda gi S ribosomal protein S7 22 kda gi cofilin-1 19 kda gi AHNAK nucleoprotein 587 kda gi plastin-3 72 kda gi eukaryotic translation elongation factor 1 alpha 1-like 50 kda gi transthyretin precursor 16 kda gi transgelin-2 22 kda gi tubulin alpha-1b chain 50 kda
48 118 gi ATP-citrate synthase isoform X1 123 kda gi proteasome subunit beta type-3 23 kda gi phosphatidylethanolaminebinding protein 1 21 kda gi short-chain dehydrogenase/ reductase family 9C member 7 35 kda gi epiplakin 376 kda gi annexin A5 36 kda gi myosin kda gi filamin A, alpha 276 kda gi actin, alpha 1, skeletal muscle 42 kda gi clathrin heavy chain kda gi enolase, alpha 47 kda gi vimentin 54 kda gi malate dehydrogenase, mitochondrial 30 kda gi purine nucleoside phosphorylase 32 kda gi collagen, type VI, alpha 3-like isoform kda gi annexin A8 37 kda gi collagen alpha-2(vi) chain isoform X1 109 kda gi kallikrein-10 precursor 31 kda gi myosin kda gi annexin IV 36 kda gi proteasome subunit beta type-4 precursor 29 kda gi peroxiredoxin-2 22 kda gi proteasome subunit alpha type-3 isoform X1 28 kda
49 Table D7: Normalised total, EmPAI and NSAF ratios for significantly regulated teat canal lining proteins Normalised total ratio Average normalised SpC E. coli S. aureus S. uberis p-value verses control # Protein name Control E.coli S. aureus S. uberis /Control /Control /Control E.coli S. aureus S. uberis 28 serum albumin precursor glyceraldehyde-3-phosphate dehydrogenase heat shock protein beta pancreatic adenocarcinoma upregulated factor-like serotransferrin precursor lactoferrin apolipoprotein A-I precursor EmPAI ratio Average EmPAI value E. coli S. aureus S. uberis p-value verses control # Protein name Control E.coli S. aureus S. uberis /Con /Con /Con E.coli S. aureus S. uberis 28 serum albumin precursor glyceraldehyde-3-phosphate dehydrogenase heat shock protein beta pancreatic adenocarcinoma upregulated factor-like serotransferrin precursor * 41.3* 17.0* lactoferrin apolipoprotein A-I precursor NSAF ratio Average NSAF value E. coli S. aureus S. uberis p-value verses control # Protein name Control E.coli S. aureus S. uberis /Con /Con /Con E.coli S. aureus S. uberis 28 serum albumin precursor glyceraldehyde-3-phosphate dehydrogenase heat shock protein beta pancreatic adenocarcinoma upregulated factor-like serotransferrin precursor * 6.63* 2.40* lactoferrin apolipoprotein A-I precursor * For serotransferrin fold calculations, the control value was set to 0.01 and to calculate EmPAI and NSAF ratios, respectively.
50 Table D8: Quickscore summary of MHC class II signal in teat-end tissue regions from 24 h bacteria challenge cows. # Treatment Tissue region Control E. coli S. aureus S. uberis Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Immunofluorescent signal intensities in the cryosections were subjectively graded as weakly positive (+), mildly positive (++), moderately positive (+++) and strongly positive (++++). Table D9: Quickscore summary of CH138A signal in teat-end tissue regions from 24 h bacteria challenge cows. # Treatment Tissue region Control E. coli S. aureus S. uberis Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Teat canal Fürstenberg s rosette Teat sinus Immunofluorescent signal intensities in the cryosections were subjectively graded as weakly positive (+), mildly positive (++), moderately positive (+++) and strongly positive (++++).
51 (A) Con EC SA SU Teat canal Con EC SA SU Teat sinus Con EC SA SU Fürstenberg's rosette (B) Con EC SA SU Teat canal Con EC SA SU Teat sinus Con EC SA SU Fürstenberg's rosette Figure D2: Semi-quantitative comparison of the proportion CD205 + and CD3 + cells in 24 h challenged teat-end tissues. Positively stained CD205 expressing cells associated with the epithelial bilayer (A), and CD3 expressing cells located within the epithelial layer (B) were counted from seven nonadjacent fields per slide at 200 x magnification and then averaged. The results are cumulative data from four latelactating cows and are expressed as a proportion of the total number of nuclei in each field. Con; Control inoculation, EC; E. coli, SA; S. aureus, SU; S. uberis. Error bar: ± SEM.
SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS
 SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS No. Protein 1 serum albumin precursor gi 30794280 2 annexin A2 gi 27807289 3 Phosphatidylethanolamine-binding
SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS No. Protein 1 serum albumin precursor gi 30794280 2 annexin A2 gi 27807289 3 Phosphatidylethanolamine-binding
Biological processes. Mitochondrion Metabolic process Catalytic activity Oxidoreductase
 Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Protein Name. IFLENVIR,DSVTYTEHAK,TV TALDVVYALK,KTVTALDVV YALK,TVTALDVVYALKR,IF LENVIRDSVTYTEHAK gi Histone H2B
 Table 1. A functional category list of proteins (Lentinula edodes) identified by 1-DGE and nesi-lc-ms/ms. The table lists indicated fraction numbers, matching peptides, scores, accession numbers, protein
Table 1. A functional category list of proteins (Lentinula edodes) identified by 1-DGE and nesi-lc-ms/ms. The table lists indicated fraction numbers, matching peptides, scores, accession numbers, protein
Supplemental Information. Host-Microbiota Interactions in the Pathogenesis. of Antibiotic-Associated Diseases
 Cell Reports, Volume Supplemental Information Host-Microbiota Interactions in the Pathogenesis of -Associated Diseases Joshua S. Lichtman, Jessica A. Ferreyra, Katharine M. Ng, Samuel A. Smits, Justin
Cell Reports, Volume Supplemental Information Host-Microbiota Interactions in the Pathogenesis of -Associated Diseases Joshua S. Lichtman, Jessica A. Ferreyra, Katharine M. Ng, Samuel A. Smits, Justin
Sequence Coverage (%) Profilin-1 P UD 2
 Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
Fig. S1. Summary of the altered metabolism pathways in alcoholic fatty liver disease using MetPA analysis (panel A).
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Fig. S1. Summary of the altered metabolism pathways in alcoholic fatty liver disease
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Fig. S1. Summary of the altered metabolism pathways in alcoholic fatty liver disease
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking
 5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
CD82/ CD63 expression in human thyroid tissues
 - 34-4 RESULTS 4.1 CD82/ CD63 expression in human thyroid tissues 4.1.1 Transcriptional analysis CD82 All investigated benign goiter tissues showed strong transcriptional level for CD82. Thyroid carcinoma
- 34-4 RESULTS 4.1 CD82/ CD63 expression in human thyroid tissues 4.1.1 Transcriptional analysis CD82 All investigated benign goiter tissues showed strong transcriptional level for CD82. Thyroid carcinoma
Quantitative Proteomics. Quantitative Proteomics
 Quantitative Proteomics Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 216 Quantitative Proteomics Quantitation in proteomics has become a popular area
Quantitative Proteomics Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 216 Quantitative Proteomics Quantitation in proteomics has become a popular area
Table S1 Differentially expressed genes showing > 2 fold changes and p <0.01 for 0.01 mm NS-398, 0.1mM ibuprofen and COX-2 RNAi.
 Table S1 Differentially expressed genes showing > 2 fold changes and p
Table S1 Differentially expressed genes showing > 2 fold changes and p
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/2/e1602038/dc1 Supplementary Materials for Mitochondrial metabolic regulation by GRP78 Manoj Prasad, Kevin J. Pawlak, William E. Burak, Elizabeth E. Perry, Brendan
advances.sciencemag.org/cgi/content/full/3/2/e1602038/dc1 Supplementary Materials for Mitochondrial metabolic regulation by GRP78 Manoj Prasad, Kevin J. Pawlak, William E. Burak, Elizabeth E. Perry, Brendan
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Supporting Information
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Supporting Information Covalent functionalization of graphene oxide with
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Supporting Information Covalent functionalization of graphene oxide with
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma
 Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler
 Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
Supplementary data Table S3. GO terms, pathways and networks enriched among the significantly correlating genes using Tox-Profiler DR CALUX Boys Girls Database Systemic lupus erythematosus 4.4 0.0021 6.7
REGULATION OF ENZYME ACTIVITY. Medical Biochemistry, Lecture 25
 REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
Protein carbonylation: a marker of oxidative stress damage
 Protein carbonylation: a marker of oxidative stress damage ITN-TREATMENT Metabolic Dysfunctions associated with Pharmacological Treatment of Schizophrenia TREATMENT Protein carbonyl formation: Metal-Catalyzed
Protein carbonylation: a marker of oxidative stress damage ITN-TREATMENT Metabolic Dysfunctions associated with Pharmacological Treatment of Schizophrenia TREATMENT Protein carbonyl formation: Metal-Catalyzed
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
the nature and importance of biomacromolecules in the chemistry of the cell: synthesis of biomacromolecules through the condensation reaction lipids
 the nature and importance of biomacromolecules in the chemistry of the cell: synthesis of biomacromolecules through the condensation reaction lipids and their sub-units; the role of lipids in the plasma
the nature and importance of biomacromolecules in the chemistry of the cell: synthesis of biomacromolecules through the condensation reaction lipids and their sub-units; the role of lipids in the plasma
The source of protein structures is the Protein Data Bank. The unit of classification of structure in SCOP is the protein domain.
 UNIT 14 PROTEINS DEFINITION A large molecule composed of one or more chains of amino acids in a specific order; the order is determined by the base sequence of nucleotides in the gene that codes for the
UNIT 14 PROTEINS DEFINITION A large molecule composed of one or more chains of amino acids in a specific order; the order is determined by the base sequence of nucleotides in the gene that codes for the
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array.
 Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Protein name Location Family MW (kda)
 Supplementary Information, Table S1. Proteomic results from LC-MS/MS analysis of MC/9 exosomes. Proteins of MC/9 exosomes were collected in the stacking parts of an SDS-PAGE, cut out, trypsinated and analysed
Supplementary Information, Table S1. Proteomic results from LC-MS/MS analysis of MC/9 exosomes. Proteins of MC/9 exosomes were collected in the stacking parts of an SDS-PAGE, cut out, trypsinated and analysed
Quiz #1. BIO200 January 11, point each
 Quiz #1 January 11, 2013 1. The primary amine group of an amino acid has a pka of 10 and the carboxylic acid group of an amino acid has a pka of 2. The side chain of the amino acid alanine is a methyl
Quiz #1 January 11, 2013 1. The primary amine group of an amino acid has a pka of 10 and the carboxylic acid group of an amino acid has a pka of 2. The side chain of the amino acid alanine is a methyl
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
SwissProt/ TrEmbl Acc. No. 1 Description. Found in Other Studies 5. MW (kda) 2 pi 3 Function 4
 P02774 Vitamin D-binding protein precursor 52.95 5.4 Cell communication c, a,b S,C P08833 Insulin-like growth factor binding protein 1 precursor 27.88 5.1 Cell communication c, a P02760 AMBP protein precursor
P02774 Vitamin D-binding protein precursor 52.95 5.4 Cell communication c, a,b S,C P08833 Insulin-like growth factor binding protein 1 precursor 27.88 5.1 Cell communication c, a P02760 AMBP protein precursor
Loss of protein association causes cardiolipin degradation in Barth syndrome
 SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
Nature Methods: doi: /nmeth.4257
 Supplementary Figure 1 Screen for polypeptides that affect cellular actin filaments. (a) Table summarizing results from all polypeptides tested. Source shows organism, gene, and amino acid numbers used.
Supplementary Figure 1 Screen for polypeptides that affect cellular actin filaments. (a) Table summarizing results from all polypeptides tested. Source shows organism, gene, and amino acid numbers used.
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2014-15 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2014-15 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Characterization of the use of wastewater from hydrothermal liquefaction as a nitrogen source by the potential algal biofuel strain Picochlorum sp.
 Characterization of the use of wastewater from hydrothermal liquefaction as a nitrogen source by the potential algal biofuel strain Picochlorum sp. Shuyi Wang, Xinguo Shi, Fatima Foflonker, Debashish Bhattacharya,
Characterization of the use of wastewater from hydrothermal liquefaction as a nitrogen source by the potential algal biofuel strain Picochlorum sp. Shuyi Wang, Xinguo Shi, Fatima Foflonker, Debashish Bhattacharya,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Supplemental Information:
 SupplementalInformation: Content: FigureS1.AccessibilityoftheL.interrogansproteomebyLC MSanalysis. FigureS2.Functionalannotationoftheidentifiedproteins. FigureS3.DistributionofMS1 featureintensities. FigureS4.ComparisonoftheDDA
SupplementalInformation: Content: FigureS1.AccessibilityoftheL.interrogansproteomebyLC MSanalysis. FigureS2.Functionalannotationoftheidentifiedproteins. FigureS3.DistributionofMS1 featureintensities. FigureS4.ComparisonoftheDDA
Supplementary Figure 1. Comparative analysis of thermal denaturation of enzymatically
 Supplementary Figures Supplementary Figure 1. Comparative analysis of thermal denaturation of enzymatically synthesized polyubiquitin chains of different length. a, Differential scanning calorimetry traces
Supplementary Figures Supplementary Figure 1. Comparative analysis of thermal denaturation of enzymatically synthesized polyubiquitin chains of different length. a, Differential scanning calorimetry traces
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Dense and Dynamic Polyethylene Glycol Shells Cloak Nanoparticles. from Uptake by Liver Endothelial Cells for Long Blood Circulation
 Dense and Dynamic Polyethylene Glycol Shells Cloak Nanoparticles from Uptake by Liver Endothelial Cells for Long Blood Circulation Hao Zhou, Zhiyuan Fan, Peter Y. Li, Junjie Deng,, Dimitrios C. Arhontoulis,
Dense and Dynamic Polyethylene Glycol Shells Cloak Nanoparticles from Uptake by Liver Endothelial Cells for Long Blood Circulation Hao Zhou, Zhiyuan Fan, Peter Y. Li, Junjie Deng,, Dimitrios C. Arhontoulis,
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis?
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
Supplementary figure legends
 Supplementary figure legends Fig. S1. Lineweaver-Burk plot of putrescine uptake by YeeF. An overnight culture of SK629 was inoculated in 100-mL LBG medium in 500-mL Erlenmeyer flasks. The medium was supplemented
Supplementary figure legends Fig. S1. Lineweaver-Burk plot of putrescine uptake by YeeF. An overnight culture of SK629 was inoculated in 100-mL LBG medium in 500-mL Erlenmeyer flasks. The medium was supplemented
Helen Kim, Ph.D. and John Cutts. Dept of Pharmacology & Toxicology University of Alabama at Birmingham
 Understanding the actions of a dietary anti-oxidant at the protein and small molecule level using top-down proteomics, enzyme assays and mass spectrometry elen Kim, Ph.D. and John Cutts Mar 9, 2012 UAB
Understanding the actions of a dietary anti-oxidant at the protein and small molecule level using top-down proteomics, enzyme assays and mass spectrometry elen Kim, Ph.D. and John Cutts Mar 9, 2012 UAB
Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences
 Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Electron Transport Chain and Oxidative phosphorylation
 Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
! Proteins are involved functionally in almost everything: " Receptor Proteins - Respond to external stimuli. " Storage Proteins - Storing amino acids
 Proteins Most structurally & functionally diverse group! Proteins are involved functionally in almost everything: Proteins Multi-purpose molecules 2007-2008 Enzymatic proteins - Speed up chemical reactions!
Proteins Most structurally & functionally diverse group! Proteins are involved functionally in almost everything: Proteins Multi-purpose molecules 2007-2008 Enzymatic proteins - Speed up chemical reactions!
MOLECULAR WEIGHT OF DIFFERENT PROTEINS PRESENT IN ZEBRAFISH EMBRYO DURING GASTRULATION PERIOD
 MOLECULAR WEIGHT OF DIFFERENT PROTEINS PRESENT IN ZEBRAFISH EMBRYO DURING GASTRULATION PERIOD 1) 37% molecular weight of about 97KDa 2) 14.6% molecular weight of about 45 KDa 3+4) 27.4% molecular weight
MOLECULAR WEIGHT OF DIFFERENT PROTEINS PRESENT IN ZEBRAFISH EMBRYO DURING GASTRULATION PERIOD 1) 37% molecular weight of about 97KDa 2) 14.6% molecular weight of about 45 KDa 3+4) 27.4% molecular weight
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
Nature Structural & Molecular Biology: doi: /nsmb.3218
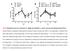
 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Unit 1: Biochemistry
 Name: Date: Carbohydrates, lipids, proteins, and enzymes 1. All living things contain which element? A. helium B. sodium C. copper D. carbon 4. Which of the following elements is best able to combine with
Name: Date: Carbohydrates, lipids, proteins, and enzymes 1. All living things contain which element? A. helium B. sodium C. copper D. carbon 4. Which of the following elements is best able to combine with
Symposium on Protein Fractionation
 Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 20122013 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO1A14 Time allowed: 2 hours Answer ALL questions in Section A,
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 20122013 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO1A14 Time allowed: 2 hours Answer ALL questions in Section A,
Biochemistry 330 September 6, 2011
 Biochemistry 330 September 6, 2011 Introductions Course Outline and Expectations Adverse Childhood Experiences Dynamic reach of biochemistry Intro to Biomolecular Structure BioChem 330 - Course Outline
Biochemistry 330 September 6, 2011 Introductions Course Outline and Expectations Adverse Childhood Experiences Dynamic reach of biochemistry Intro to Biomolecular Structure BioChem 330 - Course Outline
CHAPTER 3 Amino Acids, Peptides, Proteins
 CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
SUPPLEMENTAL DATA AGING, April 2013, Vol.5 No.4
 SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
Immunology - Lecture 2 Adaptive Immune System 1
 Immunology - Lecture 2 Adaptive Immune System 1 Book chapters: Molecules of the Adaptive Immunity 6 Adaptive Cells and Organs 7 Generation of Immune Diversity Lymphocyte Antigen Receptors - 8 CD markers
Immunology - Lecture 2 Adaptive Immune System 1 Book chapters: Molecules of the Adaptive Immunity 6 Adaptive Cells and Organs 7 Generation of Immune Diversity Lymphocyte Antigen Receptors - 8 CD markers
AP Biology Cells: Chapters 4 & 5
 AP Biology Cells: Chapters 4 & 5 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. The was the first unifying principle of biology. a. spontaneous generation
AP Biology Cells: Chapters 4 & 5 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. The was the first unifying principle of biology. a. spontaneous generation
WT siz1-2 siz1-3. WT siz1-2 siz1-3. -Pi, 0.05 M IAA. -Pi, 2.5 M NPA
 A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
number Done by Corrected by Doctor Nayef Karadsheh
 number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
Integration of Metabolism
 Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ).
 Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
1- Which of the following statements is TRUE in regards to eukaryotic and prokaryotic cells?
 Name: NetID: Exam 3 - Version 1 October 23, 2017 Dr. A. Pimentel Each question has a value of 4 points and there are a total of 160 points in the exam. However, the maximum score of this exam will be capped
Name: NetID: Exam 3 - Version 1 October 23, 2017 Dr. A. Pimentel Each question has a value of 4 points and there are a total of 160 points in the exam. However, the maximum score of this exam will be capped
Figure 1 Original Advantages of biological reactions being catalyzed by enzymes:
 Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
NBCE Mock Board Questions Biochemistry
 1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
FIG S1 Replication rates of S. suis strain 10, 10Δsly, and 10cpsΔEF on mono- and virus pre-infected porcine PCLS.
 A strain 10 10cps EF B strain 10 H1N1 + strain 10 10cps EF H1N1 + 10cps EF 10 8 10 sly 10 7 H3N2 + strain 10 H3N2 + 10cps EF CFU/ml media 10 7 10 6 10 5 10 4 CFU/ml media 10 6 10 5 10 4 10 3 0 2 4 8 12
A strain 10 10cps EF B strain 10 H1N1 + strain 10 10cps EF H1N1 + 10cps EF 10 8 10 sly 10 7 H3N2 + strain 10 H3N2 + 10cps EF CFU/ml media 10 7 10 6 10 5 10 4 CFU/ml media 10 6 10 5 10 4 10 3 0 2 4 8 12
FUNCTIONAL INTERACTIONS BETWEEN ERYTHROID KRUPPEL-LIKE FACTOR (EKLF/KLF1) AND PROTEIN PHOSPHATASE PPM1B/PP2Cß
 SUPPLEMENTAL INFORMATION FOR FUNCTIONAL INTERACTIONS BETWEEN ERYTHROID KRUPPEL-LIKE FACTOR (EKLF/KLF1) AND PROTEIN PHOSPHATASE PPM1B/PP2Cß Yvette Y. Yien and James J. Bieker FIGURE S1. Induction of EKLF
SUPPLEMENTAL INFORMATION FOR FUNCTIONAL INTERACTIONS BETWEEN ERYTHROID KRUPPEL-LIKE FACTOR (EKLF/KLF1) AND PROTEIN PHOSPHATASE PPM1B/PP2Cß Yvette Y. Yien and James J. Bieker FIGURE S1. Induction of EKLF
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12864 Supplementary Table 1 1 2 3 4 5 6 7 Peak Gene code Screen Function or Read analysis AMP reads camp annotation reads minor Tb927.2.1810 AMP ISWI Confirmed
SUPPLEMENTARY INFORMATION doi:10.1038/nature12864 Supplementary Table 1 1 2 3 4 5 6 7 Peak Gene code Screen Function or Read analysis AMP reads camp annotation reads minor Tb927.2.1810 AMP ISWI Confirmed
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) (b) (c) (d) (e) (f) (g) .
 Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Lecture 13 (10/13/17)
 Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
glpx Rv1100 Probe 4.4 kb 75 kda 50 kda 37 kda 25 kda 20 kda
 PvuI 2.0 k PvuI PvuI Rv1101c Rv1100 glpx fum 4.4 k PvuI PvuI ΔglpX Rv1101c Rv1100 HygR fum ΔglpX 5 k 4 k 3 k c 75 kda 50 kda 37 kda GLPX Δ Δ/C GLPX 2 k 1.5 k 25 kda 20 kda PRCB 1 k 0.5 k Supplementary
PvuI 2.0 k PvuI PvuI Rv1101c Rv1100 glpx fum 4.4 k PvuI PvuI ΔglpX Rv1101c Rv1100 HygR fum ΔglpX 5 k 4 k 3 k c 75 kda 50 kda 37 kda GLPX Δ Δ/C GLPX 2 k 1.5 k 25 kda 20 kda PRCB 1 k 0.5 k Supplementary
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
Chapter 6 Reading Guide
 Chapter 6 Reading Guide 1. Describe the structure of an amino acid. 2. What s the difference between an amino acid and a protein? Where do dipeptides, tripeptides and polypeptides fit in? 3. How many amino
Chapter 6 Reading Guide 1. Describe the structure of an amino acid. 2. What s the difference between an amino acid and a protein? Where do dipeptides, tripeptides and polypeptides fit in? 3. How many amino
BIOCHEMICAL EXAMINATION OF ACUTE MYOCARDIAL INFARCTION. Written by Lenka Fialová, translated by Jan Pláteník
 BIOCHEMICAL EXAMINATION OF ACUTE MYOCARDIAL INFARCTION 1 Structure of heart muscle Written by Lenka Fialová, translated by Jan Pláteník Heart muscle (myocardium) is a particular form of striated muscle,
BIOCHEMICAL EXAMINATION OF ACUTE MYOCARDIAL INFARCTION 1 Structure of heart muscle Written by Lenka Fialová, translated by Jan Pláteník Heart muscle (myocardium) is a particular form of striated muscle,
N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis
 JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This
 SUPPLEMENTAL FIGURE LEGEND Fig. S1. Generation and characterization of. (A) Coomassie staining of soluble hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This protein was expressed
SUPPLEMENTAL FIGURE LEGEND Fig. S1. Generation and characterization of. (A) Coomassie staining of soluble hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This protein was expressed
Chapter 3. Biological Molecules Great and Small
 Chapter 3 Biological Molecules Great and Small Chapter Goal: Understanding how cells use small building blocks to build larger molecules and how some of those molecules then fold into 3-D shapes Key Questions
Chapter 3 Biological Molecules Great and Small Chapter Goal: Understanding how cells use small building blocks to build larger molecules and how some of those molecules then fold into 3-D shapes Key Questions
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Biological Molecules. Carbohydrates, Proteins, Lipids, and Nucleic Acids
 Biological Molecules Carbohydrates, Proteins, Lipids, and Nucleic Acids Organic Molecules Always contain Carbon (C) and Hydrogen (H) Carbon is missing four electrons Capable of forming 4 covalent bonds
Biological Molecules Carbohydrates, Proteins, Lipids, and Nucleic Acids Organic Molecules Always contain Carbon (C) and Hydrogen (H) Carbon is missing four electrons Capable of forming 4 covalent bonds
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Proteins. AP Biology. Proteins. Proteins. Proteins. Effect of different R groups: Nonpolar amino acids. Amino acids H C OH H R. Structure.
 2008-2009 Most structurally & functionally diverse group : involved in almost everything (pepsin, DNA polymerase) (keratin, collagen) (hemoglobin, aquaporin) (insulin & other hormones) (antibodies) (actin
2008-2009 Most structurally & functionally diverse group : involved in almost everything (pepsin, DNA polymerase) (keratin, collagen) (hemoglobin, aquaporin) (insulin & other hormones) (antibodies) (actin
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding
 SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
Proteomics for Breast Cancer Programming and the Response to Endocrine Disruptors and Soy Isoflavones
 Proteomics for Breast Cancer Programming and the Response to Endocrine Disruptors and Soy Isoflavones Coral Lamartiniere, Ph.D. Department of Pharmacology and Toxicology UAB Comprehensive Cancer Center
Proteomics for Breast Cancer Programming and the Response to Endocrine Disruptors and Soy Isoflavones Coral Lamartiniere, Ph.D. Department of Pharmacology and Toxicology UAB Comprehensive Cancer Center
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
- 1 - Cell types Monocytes THP-1 cells Macrophages. LPS Treatment time (Hour) IL-6 level (pg/ml)
 Supplementary Table ST1: The dynamic effect of LPS on IL-6 production in monocytes and THP-1 cells after GdA treatment. Monocytes, THP-1 cells and macrophages (5x10 5 ) were incubated with 10 μg/ml of
Supplementary Table ST1: The dynamic effect of LPS on IL-6 production in monocytes and THP-1 cells after GdA treatment. Monocytes, THP-1 cells and macrophages (5x10 5 ) were incubated with 10 μg/ml of
Ovarian Cancer-Biomarkers
 Ovarian Cancer-Biomarkers Bobby Graham Deputy Director of the Stoller Biomarker Discovery Centre Discovery Validation Stoller Biomarker Discovery Centre Arginine R Glutamic acid E Leucine L Serine S Threonine
Ovarian Cancer-Biomarkers Bobby Graham Deputy Director of the Stoller Biomarker Discovery Centre Discovery Validation Stoller Biomarker Discovery Centre Arginine R Glutamic acid E Leucine L Serine S Threonine
Boucher et al NCOMMS B
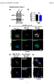 1 Supplementary Figure 1 (linked to Figure 1). mvegfr1 constitutively internalizes in endothelial cells. (a) Immunoblot of mflt1 from undifferentiated mouse embryonic stem (ES) cells with indicated genotypes;
1 Supplementary Figure 1 (linked to Figure 1). mvegfr1 constitutively internalizes in endothelial cells. (a) Immunoblot of mflt1 from undifferentiated mouse embryonic stem (ES) cells with indicated genotypes;
COURSE: Medical Microbiology, MBIM 650/720 - Fall TOPIC: Antigen Processing, MHC Restriction, & Role of Thymus Lecture 12
 COURSE: Medical Microbiology, MBIM 650/720 - Fall 2008 TOPIC: Antigen Processing, MHC Restriction, & Role of Thymus Lecture 12 FACULTY: Dr. Mayer Office: Bldg. #1, Rm B32 Phone: 733-3281 Email: MAYER@MED.SC.EDU
COURSE: Medical Microbiology, MBIM 650/720 - Fall 2008 TOPIC: Antigen Processing, MHC Restriction, & Role of Thymus Lecture 12 FACULTY: Dr. Mayer Office: Bldg. #1, Rm B32 Phone: 733-3281 Email: MAYER@MED.SC.EDU
Name: Date: Block: Biology 12
 Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
A. Major parts 1. Nucleus 2. Cytoplasm a. Contain organelles (see below) 3. Plasma membrane (To be discussed in Cellular Transport Lecture)
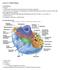 Lecture 5: Cellular Biology I. Cell Theory Concepts: 1. Cells are the functional and structural units of living organisms 2. The activity of an organism is dependent on both the individual and collective
Lecture 5: Cellular Biology I. Cell Theory Concepts: 1. Cells are the functional and structural units of living organisms 2. The activity of an organism is dependent on both the individual and collective
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions.
 Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
Lecture 1, Fall 2014 The structure of the eukaryotic cell, as it relates to cells chemical and biological functions. There are several important themes that transcends just the chemistry and bring the
