Supplemental Information. Oncogenic RAS Signaling Promotes Tumor. Immunoresistance by Stabilizing PD-L1 mrna
|
|
|
- Shanon Antony Grant
- 5 years ago
- Views:
Transcription
1 Immunity, Volume 7 al Information Oncogenic RAS Signaling Promotes Tumor Immunoresistance by Stabilizing PD-L mrna Matthew A. Coelho, Sophie de Carné Trécesson, Sareena Rana, Davide Zecchin, Christopher Moore, Miriam Molina-Arcas, Philip East, Bradley Spencer- Dene, Emma Nye, Karin Barnouin, Ambrosius P. Snijders, Wi S. Lai, Perry J. Blackshear, and Julian Downward
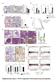
2 al Figure Legends Figure S, related to Figure. Cell-intrinsic Upregulation of PD-L through Oncogenic RAS Signalling (A) qpcr analysis of PD-L mrna expression at 6 h and h after addition of -OHT or EtOH vehicle in ER-HRAS GV MCFA cells. Mean ± SEM of two independent experiments. P<.; unpaired, two-tailed Student s t- tests. (B) Flow cytometry analysis of PD-L surface protein expression at h and four days after addition of -OHT or EtOH vehicle in ER-HRAS GV MCFA cells and four days in ER-HRAS GV HKE- cells. (C) Flow cytometry analysis of PD-L surface protein four days after addition of -OHT or EtOH vehicle to parental MCFA and HKE- cells. (D) Western blotting analysis of ER-KRAS GV type II pneumocytes treated with -OHT, MEK inhibitor or PIK inhibitor in starvation medium for h. (E) qpcr analysis of PD-L expression in H and H79 cells h after addition of MEK inhibitor or PIK inhibitor or the combination. Mean ± SEM of three (H) or two (H79) independent experiments. P<.5, *P<.5; unpaired, two-tailed Student s t-tests. (F) qpcr analysis of PD-L expression in H58 and H cells h after addition of the ERK/ inhibitor SCH7798 (5 nm). Mean ± SD of biological duplicates. P<.; unpaired, two-tailed Student s t-tests. (G) qpcr analysis of PD-L expression in H79 cells treated with PMA for h following a min pre-treatment with or MEK inhibitor. Data represent the mean ± SEM of two independent experiments. P<.; unpaired, two-tailed Student s t-test. (H) Surface expression of PD-L was measured by flow cytometry. MFI values are adjusted for the isotype control in each condition. Mean ± SEM of biological duplicates. (I) qpcr analysis of transcripts encoding antigen processing and presentation machinery in ER-KRAS GV type II pneumocytes simulated with -OHT for
3 h in starvation medium. Data represent the mean ± SEM of four independent experiments. P<.; paired, two-tailed Student s t-test. MFI, Mean Fluorescence Intensity. -OHT, nm. IFN-γ, ng/ml. MEK inhibitor GSK, 5 nm. PIK inhibitor GDC-9, 5 nm. PMA, nm.
4 Figure S, relating to Figure. RAS Signalling Increases PD-L mrna Stability through AU-rich Elements in the UTR (A) Normalised luciferase signal from the indicated human CD7 promoter region reporter constructs in H58 cells treated for 6.5 h with medium only, PMA ( nm) or IFN-γ ( ng/ml). Numbering corresponds to the GRCh8 assembly. Data are representative of two independent experiments. P<.5, two-way ANOVA. (B) Stability of murine PD-L mrna measured by qpcr after the addition of actinomycin D (5 μg/ml) and or PIK inhibitor GDC-9 (5 nm). KPB6 cells were pre-treated with or PIK inhibitor for min before actinomycin D addition. Data represent the mean ± SEM and are normalised to the h time point when actinomycin D was added, and are representative of two independent experiments. (C) Stability of murine Tusc mrna (left panel) and Ptgs mrna (right panel) measured by qpcr after the addition of actinomycin D (5 μg/ml) and or MEK inhibitor GSK (5 nm). KPB6 cells were pre-treated with or MEK inhibitor for min before actinomycin D addition. Data represent the mean ± SEM and are normalised to the h time point when actinomycin D was added.
5 Figure S, relating to Figure. AU-rich element Binding Proteins TTP and KSRP are Negative Regulators of PD-L Expression (A-C) qpcr analysis of knockdown efficiency 8 h after sirna transfections, relative to rambled control. Data represent the mean ± SD of triplicates. (D) qpcr analysis of PD-L expression h after transfection with the indicated constructs. Data represent the mean ± SEM of two independent experiments. P<.; P<.; unpaired, two-tailed Student s t-test. (E) qpcr analysis of PD-L expression in H cells 8 h after transfection with sirnas targeting AU-rich binding proteins (AU-BPs), relative to rambled () control. Data represent the mean ± SEM of two independent experiments. (F) qpcr analysis of knock-down efficiency in H cells 8 h after sirna transfections, relative to rambled control. Data represent the mean ± SEM of two independent experiments. (G) Western blotting analysis of TTP expression in H, A7 and H58 cells 8 h after sirna transfection with sirna pools against TTP relative to rambled. Overexpression of Myc-TTP serves as a positive control for immunodetection. (H) qpcr analysis of PD-L and TTP expression in H and H58 cells 8 h after sirna transfection with sirna pools or single sirnas against TTP relative to rambled. Data represent the mean ± SEM of biological duplicates.
6 Figure S, relating to Figure. RAS Regulates PD-L Expression through TTP (A) qpcr analysis of PD-L and KSRP expression in H58 cells following sirna mediated knock-down of KSRP ( h) followed by MEK inhibition ( h) with GSK (5 nm). Data represent the mean ± SEM of two independent experiments. P<.; *P<.; P<.5; *P<.5; n.s; not significant. (B) qpcr analysis of PD-L mrna from RNA immunoprecipitates using IgG control or anti-ttp antibody, or anti-ksrp antibody, using KPB6 cells pretreated with or MEK inhibitor GSK for 5.5 h (5 nm). Data represent the mean ± SD of biological triplicate IPs. (C) qpcr analysis of Gapdh mrna (control, lacking AU-rich elements) from RNA immunoprecipitates using IgG control or anti-ttp antibody, or anti- KSRP antibody, using KPB6 cells pre-treated with or MEK inhibitor GSK for 5.5 h (5 nm). Data represent the mean ± SD of biological triplicate IPs. (D) qpcr analysis of PD-L expression in H58 cells after transfection with empty, wild-type KSRP or phospho-mutant KSRP S9A constructs. Data represent the mean ± SEM of two independent experiments. P<.; *P<.; NS, not significant. Unless otherwise stated, data were compared using unpaired, two-tailed Student s t-tests. 5
7 Figure S5, relating to Figure 5. RAS-ROS-p8 Signalling Controls TTP Activity (A) MS/MS spectra for phosphopeptides STphSLVEGR (S5) and QSIphSFSGLPSGR (S78). -98 indicates the loss of H PO. (B) Flow cytometry analysis of PD-L surface protein, and intracellular ROS measured by staining with H DCFDA, in MCFA ER-HRASGV cells treated with -OHT or vehicle ± NAC ( mm) for h. The same dataset is represented as a dot-plot and a histogram and data are representative of two independent experiments. (C) Western blotting analysis of MCFA cells harbouring an inducible version of the kinase domain of MEKK (ER- MEKK), h after the addition of - OHT ( nm) or vehicle. (D) qpcr analysis of PD-L mrna expression h after treatment with NAC ( mm), reduced glutathione ( mm) or MK inhibitor III ( μm). Data represent the mean ± SEM of two independent experiments. P<., *P<.5, P<.; two-tailed Student s t-tests comparing to control condition. (E) Western blotting analysis of CT6 TTP KO cells harbouring doxycyclineinducible WT or phospho-mutant Myc-TTP constructs, treated with doxycycline or vehicle for h. Arrow indicates Myc-TTP. 6
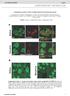
8 Figure S6, relating to Figure 6. RAS Pathway Activation is Associated with PD-L Upregulation in Human Cancers (A) Heat maps showing LUAD and COAD samples from the TCGA dataset clustered into RAS high or low pathway activity groups using RNA sequencing expression data and published RAS activity gene expression signatures (Loboda et al., ; Sweet-Cordero et al., 5). KRAS mutation status (codons, and 6) is shown for each sample. (B) qpcr analysis of PD-L and IFNGR expression in ER-KRAS GV type II pneumocytes h after treatment with vehicle, -OHT ( nm), or -OHT with IFN-γ blocking antibody ( μg/ml) or with ruxolitinib (5 nm). Mean ± SEM of two independent experiments. The panel on the right shows a qpcr analysis of PD-L expression in ER-KRAS GV type II pneumocytes following a h treatment with IFN-γ ( ng/ml) and IFN-γ with IFN-γ blocking antibody ( μg/ml) or with ruxolitinib (5 nm) to verify blocking of IFN-γ IFGR signalling. (C) TTP mrna expression in human patient lung and colon normal tissue versus adenocarcinoma, from publically available datasets (Selamat et al., ; Skrzypczak et al., ). Wilcoxon signed-rank test. (D) qpcr analysis of PD-L and TTP expression in FACS purified CD5 - CD - DAPI - EpCAM + cells derived from lung tumours or matched normal adjacent lung tissue from Kras LSL-GD/+ ; Trp5 F/F mice. Each point represents data from an individual mouse and is normalised to the matched normal lung tissue. *P<.5; unpaired, two-tailed Student s t-tests. (E) Flow cytometry analysis of PD-L expression on CD5-CD-DAPI- cells derived from macroscopically dissected lung tumours or normal adjacent lung tissue from Kras LSL-GD/+ ; Trp5 F/F mice. Each point represents data from an individual mouse and is normalised to the matched normal lung tissue. Data are pooled from two independent experiments. MFI; Mean Fluorescence Intensity. *P<.5; unpaired, two-tailed Student s t-test. 7
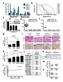
9 Figure S7, relating to Figure 7. Restoration of Tumour Cell TTP Expression Enhances Anti-tumour Immunity (A) Stability of murine PD-L mrna measured by qpcr analysis. CT6 TTP (teton) cells were pretreated with doxycycline (Dox.; μg/ml) or vehicle for 6 h and then MEK inhibitor (GSK, trametinib; 5 nm) for an additional min before actinomycin D (ActD; μg/ml) was added. Data are normalised to time h when ActD was added and represent the mean ± SEM of two independent experiments. (B) Western blotting analysis of stable MC8 cell lines expressing Myctagged, mouse TTP under a tetracycline-inducible promoter (TTP tet-on). Cells were treated with doxycycline (Dox.,. μg/ml or μg/ml) or vehicle in starvation medium for h before analysis. Arrow indicates Myc-TTP. (C) qpcr analysis of PD-L expression from stable MC8 (TTP tet-on) cell lines treated with doxycycline (Dox., μg/ml) or vehicle in starvation medium for h before analysis. Data represent the mean ± SEM of biological duplicates. (D) Confluency was measured using IncuCyte for CT6 stable derivative cell lines treated with the indicated concentrations of doxycycline or vehicle at t = h. Data represent the mean ± SD of biological triplicates and are representative of two independent experiments. (E) Confluency was measured using IncuCyte for MC8 stable derivative cell lines treated with the indicated concentrations of doxycycline or vehicle at t = h. Data represent the mean ± SD of biological triplicates. (F) Representative histograms from flow cytometry analysis of PD-L surface expression in TTP (teton) CT6 stable cells lines expressing endogenous PD-L, PD-L + wild-type UTR cdna, or PD-L UTR cdna, after treatment with doxycycline (Dox., μg/ml) or vehicle for 7 h. Data are representative of two independent experiments and also form part of Figure 7B. (G) Tumour growth curves for the indicated CT6-derived cell lines shown in Figure S7C, subcutaneously transplanted into BALB/c mice (n = 5 WT UTR + Dox and UTR + H O; n = WT UTR + H O and UTR + Dox; n = 8
10 empty + H O and empty + Dox). Vehicle or doxycycline (Dox., 5 mg/kg) was administered by oral gavage and commenced from day three after tumour cell injection. Mean ± SEM. P<., *P<., *P<.5; two-way ANOVA. (H) Representative histological analysis of CD+ cells in subcutaneous tumours at the end-point from the experiment described in Figure 7C, and quantified in Figure 7G. Scale bar is 5 µm. (I) Proposed molecular model. Oncogenic RAS activity leads to hyperphosphorylation, whereas PPA activity promotes hypophosphorylation of TTP (Bourcier et al., ; Deleault et al., 8; Essafi-Benkhadir et al., 7; Hardle et al., 5; Rahman et al., 5; Sun et al., 7), constituting a rapid switch controlling TTP activity. Low TTP expression and activity in tumour cells represents a permissive context for PD-L expression and immune evasion. 9
11 Table S, relating to Figure 5 and Figure S5. TTP phosphopeptides. Identified mouse TTP phosphopeptides from MS analyses pooled from two independent biological experiments. Identifications are % FDR controlled. PEP indicates the probability that the identification is incorrect. Phosphosite assignment probabilities are indicated in parenthesis. PEP Score Position Modified sequence Phospho (STY) Probabilities Number of Phospho (STY).67 5 _STS(ph)LVEGR_ STS()LVEGR.E- 7 8 _PGPELS(ph)PSPT(ph)SPTATPTTSSR_ PGPELS(.998)PS(.6)PT(.77)S(.)PT(.68)AT(.)P T(.)T(.)S(.)S(.)R.E _PGPELSPS(ph)PTSPTATPTTSSR_ PGPELS(.6)PS(.9)PT(.)S(.9)PT(.)AT(.)P TTSSR.7E _PGPELSPSPTS(ph)PTATPTTSSR_ PGPELSPSPT(.8)S(.857)PT(.)ATPTTSSR. 7 5 _TYS(ph)ESGRCR_ TYS()ESGRCR 6.9E-8 78 _QSIS(ph)FSGLPSGR_ QSIS()FSGLPSGR RS(.7)S(.767)PPPPGFS(.95)GPS(.6)LS(.)S(.).5E _RSS(ph)PPPPGFS(ph)GPSLSSCSFSPSSSPPPP CS(.)FS(.)PS(.9)S(.85)S(.8)PPPPGDLPLSPSAFSA GDLPLSPSAFSAAPGTPVTR_ APGTPVTR.5E _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_.5E _RSS(ph)PPPPGFS(ph)GPS(ph)LSSCSFSPSSSP PPPGDLPLSPSAFSAAPGTPVTR_ 9.68E-5 69 _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E-5 69 _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E-5 69 _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ 9.68E-5 69 _RSSPPPPGFS(ph)GPS(ph)LSSCSFSPSSSPPPP GDLPLSPSAFSAAPGTPVTR_ RS(.5)S(.5)PPPPGFS(.67)GPS(.7)LS(.69)S(.67) CS(.66)FS(.65)PS(.65)S(.6)S(.6)PPPPGDLPLSPSA FSAAPGTPVTR RS(.7)S(.68)PPPPGFS(.79)GPS(.)LS(.)S(.97)C S(.9)FS(.8)PS(.7)S(.7)S(.65)PPPPGDLPLSPSAFSA APGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR RS(.)S(.77)PPPPGFS(.)GPS(.)LS(.)S(. )CS(.)FS(.)PS(.)S(.)S(.)PPPPGDLPLSPSA FSAAPGTPVTR _S(ph)TTPSTIWGPLGGLAR_ S(.)T(.)T(.)PSTIWGPLGGLAR _S(ph)TTPSTIWGPLGGLAR_ S(.)T(.)T(.)PSTIWGPLGGLAR.9E _STT(ph)PSTIWGPLGGLAR_ ST(.)T(.997)PSTIWGPLGGLAR 6.7E _S(ph)PSAHSLGSDPDDYASSGSSLGGSDSPVFE 6.7E _S(ph)PSAHSLGSDPDDYASSGSSLGGSDSPVFE 5.7E _SPSAHS(ph)LGSDPDDYASSGSSLGGSDSPVFE.5E _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 79 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 8 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 8 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 8 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 87 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE.5 89 _SPSAHSLGS(ph)DPDDYASSGSSLGGSDSPVFE S(.6)PS(.)AHS(.87)LGS(.)DPDDY(.)AS(. 5)S(.5)GS(.5)S(.5)LGGS(.5)DS(.5)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.65)LGS(.6)DPDDY(.)AS(.) S(.)GS(.)S(.)LGGS(.)DS(.)PVFEAGVFGP PQTPAPPR S(.7)PS(.7)AHS(.7)LGS(.)DPDDY(.)AS(. )S(.)GS(.)S(.)LGGS(.)DS(.)PVFEAGVF GPPQTPAPPR S(.9)PS(.9)AHS(.78)LGS(.)DPDDY(.7)AS(.7 8)S(.78)GS(.78)S(.78)LGGS(.78)DS(.78)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR S(.)PS(.)AHS(.)LGS(.9)DPDDY(.)AS(. 9)S(.9)GS(.9)S(.9)LGGS(.9)DS(.9)PVFEAGVF GPPQTPAPPR
12 Table S, relating to STAR Methods, Method Details. Cell lines and growth conditions. Cell line Normal medium Starvation medium H58 RPMI + % FCS N/A A7 RPMI + % FCS N/A H79 RPMI + % FCS N/A KPB6 DMEM + % FCS N/A Type II pneumocytes DCCM- + % FCS +.5 % FCS for qpcr and FACS + % FCS for mrna half-life analysis SW87 DMEM + % FCS N/A H RPMI + % FCS N/A 9FT DMEM + % FCS N/A TTP KO and TTP DMEM + % FCS +.5 % FCS WT MEFs MCFA F:DMEM mix (:) + 5 % horse serum and 5 % horse serum, ng/ml EGF, µg/ml insulin, ng/ml cholera toxin,.5 µg/ml hydrocortisone CT6 RPMI + % FCS N/A A59 DMEM + % FCS N/A MC8 RPMI + % FCS +.5 % FCS N/A, not applicable.
13 Figure S A B PD-L expression realtive MCFA-ER HRAS GV EtOH -OHT 6 h h % of maximum 8 6 MCFA-ER HRAS GV 5 PD-L - PE Unstained Isotype EtOH h -OHT h EtOH d -OHT d 8 6 HKE--ER HRAS GV 5 PD-L - PE Unstained Isotype EtOH -OHT C % of maximum 8 6 MCFA - parental 8 6 HKE- - parental Unstained Isotype EtOH -OHT D -OHT EtOH MEK i PIK i p-erk ERK p-akt (S7) AKT Vinculin 5 5 PD-L - PE E PD-L expression relative..5. MEK i H * PIK i MEK i + PIK i PD-L expression relative..5. MEK i H79 * PIK i MEK i + PIK i F PD-L expression relative..5. H58 ERKi PD-L expression relative..5. H ERKi G PD-L expression relative H79 PMA PMA + MEKi H PD-L (relative MFI) 5 Human Mouse A7 H58 H79 68T MEK i PIK i MEK i + PIK i I Expression relative Type II pneumocytes - ER-KRAS GV ns BM HLA-A HLA-B HLA-C TAP TAP TAPBP Et-OH -OHT
14 Figure S A pgl Firefly luciferase reporters: SV FF luciferase SV enh. SV (Control) GRCh8 Homo sapiens Chromosome 9 FF luciferase CD7 pro. FF luciferase Basic (Empty) kb pro. 587 CD7 pro FF luciferase kb pro CD7 pro. FF luciferase CD7 pro. FF luciferase CD7 enh. kb pro. kb pro. + enh. ø PMA IFN- 6 8 Firefly luciferase normalised to Renilla B KPB6 PD-L expression relative to Gapdh Time (h) + Act D (5 ug/ml) PIK i + Act D (5 ug/ml) C KPB6 KPB6 Tusc expression relative to Gapdh Time (h) Ptgs expression relative to Gapdh Time (h) + Act D (5 ug/ml) MEK i + Act D (5 ug/ml)
15 Figure S A H58 sirna knock-down efficiency D H58 - h Expression relative..5. * * siauf siksrp sihur sittp PD-L expression relative Empty + Emtpy Empty + TTP KSRP + Empty KSRP + TTP B Expression relative..5. A7 sirna knock-down efficiency * siauf siksrp sihur sittp E PD-L expression relative 6 sibrf- H sibrf- * sittp C F H sirna knock-down efficiency H sirna knock-down efficiency Expression relative..5. siauf siksrp sihur sittp Expression relative..5. sibrf- sibrf- sittp G H A7 H58 H58 kda sittp sittp sittp Empty Myc-TTP 5 TTP 8 Vinculin H H58 H Expression relative 6 5 sittp # sittp # sittp # sittp # sittp pool Expression relative sittp # sittp # sittp # sittp # sittp pool PD-L TTP
16 Figure S A PD-L expression relative H58 - PD-L * MEKi n.s MEKi KSRP expression relative H58 - KSRP * n.s MEKi MEKi siksrp siksrp B KPB6 KPB6 6 RNA IP - PD-L n.s *.9.5 MEKi 5 RNA IP - PD-L n.s.78 * 7.9 MEKi % Input.. IgG TTP IgG TTP % Input IgG KSRP IgG KSRP C RNA IP - Gapdh (5.5 h) RNA IP - Gapdh (5.5 h) % Input 6 n.s n.s.9... IgG TTP IgG TTP MEKi % Input n.s * * IgG KSRP IgG KSRP MEKi D. H58 - h * n.s PD-L expression relative egfp-empty egfp-ksrp egfp-ksrp S9A
17 Figure S5 A MS/MS spectrum for STphSLVEGR MS/MS spectrum for QSIphSFSGLPSGR B 5 MCFA-ER HRAS GV Unstained ROS - DCF % of maximum 8 6 % of maximum 8 6 Isotype EtOH -OHT -OHT + NAC 5 PD-L - PE 5 ROS - DCF 5 PD-L - PE C MCFA-ER MEKK D A7 H kda 5 8 EtOH -OHT p-p8 (T8, Y8) Vinculin PD-L expression relative..5. NAC * Glut. MK i III PD-L expression relative..5. NAC * Glut. MK i III E CT6 TTP KO H79 H58 kda 5 8 TTP WT HO Dox. S5A S78A TTP HO Dox. Myc Vinculin PD-L expression relative NAC Glut. MK i III PD-L expression relative NAC Glut. MK i III
18 Figure S6 A Loboda-Watters signature Lung Loboda-Watters signature Colon CD7 FRL IL8R IFNGR SLCA LCP CDC SOCS PSEN ILRB PTPRC TP5 TIGIT CDD IL8RAP PSEN CDE RPS6 EOMES Expression IFNGR CD7 SOCS SLCA IL8RAP FRL RPS6 TIGIT PTPRC ILRB LCP CDE IL8R PSEN CDC CDD EOMES PSEN TP5 Expression High KRAS mutation NO KRAS mutation NO RAS pathway activity YES YES Low p<.5 p-value p<.5 p-value B Expression relative Type II pneumocytes - ER-KRAS GV n.s * n.s EtOH -OHT -OHT + IFN-g block -OHT + Rux. n.s EtOH -OHT -OHT + IFN-g block -OHT + Rux. PD-L IFNGR PD-L expression relative Type II pneumocytes - ER-KRAS GV ø IFN- IFN- + Rux. IFN- + blocking ab. C TTP expression (log) Selamat - Lung P =.5 x Skrzypczak - Colon P = 9. x - D E mrna expression relative to control KrasLSL-GD/+; Trp5F/F mouse lung: CD5 - CD - DAPI - EpCAM + PD-L TTP KrasLSL/+; Trp5 F/F mouse lung: CD5 - CD - DAPI - Normal adjacent Tumour * * Lung (n = 58) Lung adenocarcinoma (n = 58) Colon (n = ) Colorectal adenocarcinoma (n = 5) PD-L (relative MFI) Normal adjacent Tumours
19 Figure S7 A CT6 TTP (teton) B MC8 TTP (teton) C MC8 TTP (teton) PD-L expression relative to Gapdh..5. Time (h) ActD + H O MEKi Dox. Dox. + MEKi kda 5 8 HO Dox. Dox. Myc Vinculin PD-L expression relative to Gapdh..5. HO Dox. D CT6 TTP (teton) E MC8 TTP (teton) Confluency (%) Time (h) Empty: TTP (teton) Empty: TTP (teton) + H O Empty: TTP (teton) + Dox.. g/ml Empty: TTP (teton) + Dox. g/ml PD-L 'UTR: TTP (teton) PD-L 'UTR: TTP (teton) + H O PD-L 'UTR: TTP (teton) + Dox.. g/ml PD-L 'UTR: TTP (teton) + Dox. g/ml % Confluency Time (h) TTP WT ø TTP WT H O TTP WT Dox.. µg/ml TTP WT Dox. µg/ml % of maximum F 8 6 CT6 Unstained Isotype TTP (teton): Empty + HO TTP (teton): Empty + Dox. TTP (teton): PD-L WT UTR + HO TTP (teton): PD-L WT UTR + Dox. TTP (teton): PD-L UTR + HO Tumour volume (mm ) G 8 6 CT6 - BALB/c * n.s * TTP (teton): Empty + HO TTP (teton): Empty + Dox. TTP (teton): PD-L WT UTR + HO TTP (teton): PD-L WT UTR + Dox. TTP (teton): PD-L UTR + HO TTP (teton): PD-L UTR + Dox. 5 PD-L - PE TTP (teton): PD-L UTR + Dox. 5 5 Time (days) H TTP (teton): Empty TTP (teton): Empty + H O + Dox. TTP (teton): PD-L 'UTR TTP (teton): PD-L 'UTR + H O + Dox. I ACTIVE MK OA ERK INACTIVE H&E TTP PPA Trametinib TTP P P P PD-L LOW PD-L HIGH CD
Oncogenic RAS Signaling Promotes Tumor Immunoresistance by Stabilizing PD-L1 mrna
 Article Oncogenic RAS Signaling Promotes Tumor Immunoresistance by Stabilizing PD-L1 mrna Graphical Abstract Authors Matthew A. Coelho, Sophie de Carné Trécesson, Sareena Rana,..., Wi S. Lai, Perry J.
Article Oncogenic RAS Signaling Promotes Tumor Immunoresistance by Stabilizing PD-L1 mrna Graphical Abstract Authors Matthew A. Coelho, Sophie de Carné Trécesson, Sareena Rana,..., Wi S. Lai, Perry J.
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung
 Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Figure 1
 Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplementary material. Supplementary Figure legends
 Supplementary material Supplementary Figure legends Supplementary Figure 1: Senescence-associated proliferation stop in response to oncogenic N-RAS expression Proliferation of NHEM cells without (ctrl.)
Supplementary material Supplementary Figure legends Supplementary Figure 1: Senescence-associated proliferation stop in response to oncogenic N-RAS expression Proliferation of NHEM cells without (ctrl.)
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
SUPPLEMENTARY FIGURE LEGENDS. atypical adenomatous hyperplasias (AAH); Grade II: adenomas; Grade III: adenocarcinomas;
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure S1: Tumor grades in Ras G12D ; p53 / lung tumors. Representative histology (H&E) of K-Ras G12D ; p53 / lung tumors 13 weeks after tumor initiation. Grade
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure S1: Tumor grades in Ras G12D ; p53 / lung tumors. Representative histology (H&E) of K-Ras G12D ; p53 / lung tumors 13 weeks after tumor initiation. Grade
Supplementary Figure 1. BMS enhances human T cell activation in vitro in a
 Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
(A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and a
 Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary Figure 1. Deletion of Smad3 prevents B16F10 melanoma invasion and metastasis in a mouse s.c. tumor model.
 A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
A B16F1 s.c. Lung LN Distant lymph nodes Colon B B16F1 s.c. Supplementary Figure 1. Deletion of Smad3 prevents B16F1 melanoma invasion and metastasis in a mouse s.c. tumor model. Highly invasive growth
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Nature Medicine: doi: /nm.4078
 Supplementary Figure 1. Cetuximab induces ER stress response in DiFi cells. (a) Scheme of SILAC proteome. (b) MS-base read out of SILAC experiment. The histogram of log 2 -transformed normalized H/L ratios
Supplementary Figure 1. Cetuximab induces ER stress response in DiFi cells. (a) Scheme of SILAC proteome. (b) MS-base read out of SILAC experiment. The histogram of log 2 -transformed normalized H/L ratios
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 EGFR inhibition activates signaling pathways (a-b) EGFR inhibition activates signaling pathways (a) U251EGFR cells were treated with erlotinib (1µM) for the indicated times followed
Supplementary Figure 1 EGFR inhibition activates signaling pathways (a-b) EGFR inhibition activates signaling pathways (a) U251EGFR cells were treated with erlotinib (1µM) for the indicated times followed
Supplemental Figure 1. Western blot analysis indicated that MIF was detected in the fractions of
 Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Integrin CD11b negatively regulates TLR-triggered inflammatory responses by. activating Syk and promoting MyD88 and TRIF degradation via cbl-b
 Integrin CD11b negatively regulates TLR-triggered inflammatory responses by activating Syk and promoting MyD88 and TRIF degradation via cbl-b Chaofeng Han, Jing Jin, Sheng Xu, Haibo Liu, Nan Li, and Xuetao
Integrin CD11b negatively regulates TLR-triggered inflammatory responses by activating Syk and promoting MyD88 and TRIF degradation via cbl-b Chaofeng Han, Jing Jin, Sheng Xu, Haibo Liu, Nan Li, and Xuetao
Supplemental Table S1
 Supplemental Table S. Tumorigenicity and metastatic potential of 44SQ cell subpopulations a Tumorigenicity b Average tumor volume (mm ) c Lung metastasis d CD high /4 8. 8/ CD low /4 6./ a Mice were injected
Supplemental Table S. Tumorigenicity and metastatic potential of 44SQ cell subpopulations a Tumorigenicity b Average tumor volume (mm ) c Lung metastasis d CD high /4 8. 8/ CD low /4 6./ a Mice were injected
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table File Name: Peer Review File Description: Supplementary Table 1 Primers and taqman probes used were the following:
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table File Name: Peer Review File Description: Supplementary Table 1 Primers and taqman probes used were the following:
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary Materials and Methods
 Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Supplementary Figure 1. Characterization of basophils after reconstitution of SCID mice
 Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL-
 Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis
 Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
Supplementary Figure 1: STAT3 suppresses Kras-induced lung tumorigenesis (a) Immunohistochemical (IHC) analysis of tyrosine 705 phosphorylation status of STAT3 (P- STAT3) in tumors and stroma (all-time
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with
 Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
Supplementary Materials
 Supplementary Materials Figure S1. MTT Cell viability assay. To measure the cytotoxic potential of the oxidative treatment, the MTT [3-(4,5-dimethylthiazol- 2-yl)-2,5-diphenyl tetrazolium bromide] assay
Supplementary Materials Figure S1. MTT Cell viability assay. To measure the cytotoxic potential of the oxidative treatment, the MTT [3-(4,5-dimethylthiazol- 2-yl)-2,5-diphenyl tetrazolium bromide] assay
Supplementary Information and Figure legends
 Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information
 Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Increased ABHD5 expression in human colon cancer associated macrophages. (a) Murine peritoneal macrophages were treated with regular culture medium (Ctrl) or
Supplementary Figures Supplementary Figure 1 Increased ABHD5 expression in human colon cancer associated macrophages. (a) Murine peritoneal macrophages were treated with regular culture medium (Ctrl) or
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. ETBF activate Stat3 in B6 and Min mice colons
 Supplementary Figure 1 ETBF activate Stat3 in B6 and Min mice colons a pstat3 controls Pos Neg ETBF 1 2 3 4 b pstat1 pstat2 pstat3 pstat4 pstat5 pstat6 Actin Figure Legend: (a) ETBF induce predominantly
Supplementary Figure 1 ETBF activate Stat3 in B6 and Min mice colons a pstat3 controls Pos Neg ETBF 1 2 3 4 b pstat1 pstat2 pstat3 pstat4 pstat5 pstat6 Actin Figure Legend: (a) ETBF induce predominantly
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
X P. Supplementary Figure 1. Nature Medicine: doi: /nm Nilotinib LSK LT-HSC. Cytoplasm. Cytoplasm. Nucleus. Nucleus
 a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
Spleen. mlns. E Spleen 4.1. mlns. Spleen. mlns. Mock 17. Mock CD8 HIV-1 CD38 HLA-DR. Ki67. Spleen. Spleen. mlns. Cheng et al. Fig.
 C D E F Mock 17 Mock 4.1 CD38 57 CD8 23.7 HLA-DR Ki67 G H I Cheng et al. Fig.S1 Supplementary Figure 1. persistent infection leads to human T cell depletion and hyper-immune activation. Humanized mice
C D E F Mock 17 Mock 4.1 CD38 57 CD8 23.7 HLA-DR Ki67 G H I Cheng et al. Fig.S1 Supplementary Figure 1. persistent infection leads to human T cell depletion and hyper-immune activation. Humanized mice
c Ischemia (30 min) Reperfusion (8 w) Supplementary Figure bp 300 bp Ischemia (30 min) Reperfusion (4 h) Dox 20 mg/kg i.p.
 a Marker Ripk3 +/ 5 bp 3 bp b Ischemia (3 min) Reperfusion (4 h) d 2 mg/kg i.p. 1 w 5 w Sacrifice for IF size A subset for echocardiography and morphological analysis c Ischemia (3 min) Reperfusion (8
a Marker Ripk3 +/ 5 bp 3 bp b Ischemia (3 min) Reperfusion (4 h) d 2 mg/kg i.p. 1 w 5 w Sacrifice for IF size A subset for echocardiography and morphological analysis c Ischemia (3 min) Reperfusion (8
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads (Miltenyi Biotec) for depletion of CD25 +
 Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplemental Information. NRF2 Is a Major Target of ARF. in p53-independent Tumor Suppression
 Molecular Cell, Volume 68 Supplemental Information NRF2 Is a Major Target of ARF in p53-independent Tumor Suppression Delin Chen, Omid Tavana, Bo Chu, Luke Erber, Yue Chen, Richard Baer, and Wei Gu Figure
Molecular Cell, Volume 68 Supplemental Information NRF2 Is a Major Target of ARF in p53-independent Tumor Suppression Delin Chen, Omid Tavana, Bo Chu, Luke Erber, Yue Chen, Richard Baer, and Wei Gu Figure
Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as
 Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as well as their downstream effectors across a panel of ESCC
Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as well as their downstream effectors across a panel of ESCC
Supplementary Figure 1: Digitoxin induces apoptosis in primary human melanoma cells but not in normal melanocytes, which express lower levels of the
 Supplementary Figure 1: Digitoxin induces apoptosis in primary human melanoma cells but not in normal melanocytes, which express lower levels of the cardiac glycoside target, ATP1A1. (a) The percentage
Supplementary Figure 1: Digitoxin induces apoptosis in primary human melanoma cells but not in normal melanocytes, which express lower levels of the cardiac glycoside target, ATP1A1. (a) The percentage
K-Ras EGFR pegfr Tubulin. pplcγ 1. Mock EGF
 shcttl shkras EGF + + 15 17 17 KRas EGFR pegfr 13 pplcγ 1 Level of pegfr (A.U.) 1 8 6 4 2 shkras Mock EGF Supplementary Figure S1. KRasdeficient pancreatic cancer cells retain normal RTK signalling. (a)
shcttl shkras EGF + + 15 17 17 KRas EGFR pegfr 13 pplcγ 1 Level of pegfr (A.U.) 1 8 6 4 2 shkras Mock EGF Supplementary Figure S1. KRasdeficient pancreatic cancer cells retain normal RTK signalling. (a)
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Fang et al. NMuMG. PyVmT unstained Anti-CCR2-PE MDA-MB MCF MCF10A
 A NMuMG PyVmT 16.5+.5 47.+7.2 Fang et al. unstained Anti-CCR2-PE 4T1 Control 37.6+6.3 56.1+.65 MCF1A 16.1+3. MCF-7 3.1+5.4 MDA-M-231 42.1+5.5 unstained Secondary antibody only Anti-CCR2 SUPPLEMENTAL FIGURE
A NMuMG PyVmT 16.5+.5 47.+7.2 Fang et al. unstained Anti-CCR2-PE 4T1 Control 37.6+6.3 56.1+.65 MCF1A 16.1+3. MCF-7 3.1+5.4 MDA-M-231 42.1+5.5 unstained Secondary antibody only Anti-CCR2 SUPPLEMENTAL FIGURE
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1
 Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/430/ra57/dc1 Supplementary Materials for The 4E-BP eif4e axis promotes rapamycinsensitive growth and proliferation in lymphocytes Lomon So, Jongdae Lee, Miguel
www.sciencesignaling.org/cgi/content/full/9/430/ra57/dc1 Supplementary Materials for The 4E-BP eif4e axis promotes rapamycinsensitive growth and proliferation in lymphocytes Lomon So, Jongdae Lee, Miguel
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. NKT ligand-loaded tumour antigen-presenting B cell- and monocyte-based vaccine induces NKT, NK and CD8 T cell responses. (A) The cytokine profiles of liver
Supplementary Figures Supplementary Figure 1. NKT ligand-loaded tumour antigen-presenting B cell- and monocyte-based vaccine induces NKT, NK and CD8 T cell responses. (A) The cytokine profiles of liver
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figure 1
 Supplementary Figure 1 how HFD how HFD Epi WT p p Hypothalamus p p Inguinal WT T Liver Lean mouse adipocytes p p p p p p Obese mouse adipocytes Kidney Muscle Spleen Heart p p p p p p p p Extracellular
Supplementary Figure 1 how HFD how HFD Epi WT p p Hypothalamus p p Inguinal WT T Liver Lean mouse adipocytes p p p p p p Obese mouse adipocytes Kidney Muscle Spleen Heart p p p p p p p p Extracellular
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table Title of file for HTML: Peer Review File Description: Innate Scavenger Receptor-A regulates
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs
 Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
Supplementary Fig. 1 p38 MAPK negatively regulates DC differentiation. (a) Western blot analysis of p38 isoform expression in BM cells, immature DCs (idcs) and mature DCs (mdcs). A myeloma cell line expressing
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
SUPPLEMENTARY INFORMATION
 Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Supplemental Figure 1. Furin is efficiently deleted in CD4 + and CD8 + T cells. a, Western blot for furin and actin proteins in CD4cre-fur f/f and fur f/f Th1 cells. Wild-type and furin-deficient CD4 +
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep
 SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
SUPPLEMENTARY INFORMATION The autoimmune disease-associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness Jinyi Zhang, Naima
Expanded View Figures
 PEX13 functions in selective autophagy Ming Y Lee et al Expanded View Figures Figure EV1. PEX13 is required for Sindbis virophagy. A, B Quantification of mcherry-capsid puncta per cell (A) and GFP-LC3
PEX13 functions in selective autophagy Ming Y Lee et al Expanded View Figures Figure EV1. PEX13 is required for Sindbis virophagy. A, B Quantification of mcherry-capsid puncta per cell (A) and GFP-LC3
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts
 Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
Supplementary Figure 1 Induction of cellular senescence and isolation of exosome. a to c, Pre-senescent primary normal human diploid fibroblasts (TIG-3 cells) were rendered senescent by either serial passage
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
Supplementary Figure S1 Supplementary Figure S2
 Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma
 Supplementary information for: Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma Rintaro Hashizume 1, Noemi Andor 2, Yuichiro Ihara 2, Robin Lerner 2, Haiyun
Supplementary information for: Pharmacologic inhibition of histone demethylation as a therapy for pediatric brainstem glioma Rintaro Hashizume 1, Noemi Andor 2, Yuichiro Ihara 2, Robin Lerner 2, Haiyun
Supplemental Figures:
 Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplemental Materials for. Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to. FTY720 during neuroinflammation
 Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic
 Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Tbk1-TKO! DN cells (%)! 15! 10!
 a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Supplemental information
 Supplemental information PI(3)K p11δ controls the sucellular compartmentalization of TLR4 signaling and protects from endotoxic shock Ezra Aksoy, Salma Taoui, David Torres, Sandrine Delauve, Aderrahman
Supplemental information PI(3)K p11δ controls the sucellular compartmentalization of TLR4 signaling and protects from endotoxic shock Ezra Aksoy, Salma Taoui, David Torres, Sandrine Delauve, Aderrahman
Bezzi et al., Supplementary Figure 1 *** Nature Medicine: doi: /nm Pten pc-/- ;Zbtb7a pc-/- Pten pc-/- ;Pml pc-/- Pten pc-/- ;Trp53 pc-/-
 Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured
 Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β
 Supplementary Figures Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β and LPS. Non-parenchymal liver cells were isolated and treated with or without TGF-β
Supplementary Figures Supplementary Figure 1. Expression of CUGBP1 in non-parenchymal liver cells treated with TGF-β and LPS. Non-parenchymal liver cells were isolated and treated with or without TGF-β
Complexity of intra- and inter-pathway loops in colon cancer and melanoma: implications for targeted therapies
 Complexity of intra- and inter-pathway loops in colon cancer and melanoma: implications for targeted therapies René Bernards The Netherlands Cancer Institute Amsterdam The Netherlands Molecular versus
Complexity of intra- and inter-pathway loops in colon cancer and melanoma: implications for targeted therapies René Bernards The Netherlands Cancer Institute Amsterdam The Netherlands Molecular versus
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
a surface permeabilized
 a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5
 sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
