Supplemental Figure 1. Immune phenotype of mcd19 targeted CAR T and dose titration of in vivo
|
|
|
- Bruce Rogers
- 5 years ago
- Views:
Transcription
1 Supplemental Materials Supplemental Methods Supplemental Figure 1. Immune phenotype of mcd19 targeted CAR T and dose titration of in vivo efficacy. Supplemental Figure 2. Gene expression of fluorescent-protein tagged CAR T cells. Supplemental Figure 3. Fluorescent protein tagged CAR T cells function similarly to non-tagged counterparts. Supplemental Figure 4. Transduction efficiency and immune phenotype of mcd19 targeted CAR T cells for survival study (Figure 2D). Supplemental Figure 5. Transduction efficiency and immune phenotype of CAR T cells used in irradiated CAR T study (Fig. 3B-C). Supplemental Figure 6. Differential gene expression of CD4+ m19-humbbz CAR T cells. Supplemental Figure 7. CAR expression and CD4/CD8 subsets of human CD19 targeted CAR T cells for Figure 5E-G. Supplemental Figure 8. Transduction efficiency and immune phenotype of mcd19 targeted wild type (WT) and TRAF1 -/- CAR T cells used for in vivo study (Figure 6D). Supplemental Figure 9. Mutated CAR T cells have increased NF-κB signaling, improved cytokine production, anti-apoptosis, and in vivo function. Supplemental Figure 1. TRAF and CAR co-expression in human CD19-targeted CAR T cells. Supplemental Figure 11. TRAF2 over-expressed h19bbz CAR T cells show similar in vivo efficacy to h19bbz CAR T cells in an aggressive leukemia model. S1
2 Supplemental Table 1. Probesets increased in m19z and vs CAR T cells. Supplemental Table 2. Probesets increased in vs m19z and CAR T cells. Supplemental Table 3. Probesets differentially expressed in m19z vs CAR T cells. Supplemental Table 4. Probesets differentially expressed in vs CAR T cells. S2
3 Supplemental Methods Gene Expression Microarray. For m19z, and comparison, three million CAR T cells were incubated with T3-mCD19 cells overnight. The next day live CAR T cells were sorted into Trizol (Thermo Fisher Scientific, Waltham, MA). RNA was isolated according to manufacturer s instructions and run on a MOE 43A 2. array Mouse Genechip (Affymetrix, Santa Clara, CA) at the Genomics Core Facility. Gene expression analyses and graphic representations were performed with the Partek Genomics Suite Software. RMA normalization was performed and values generated for each probeset for all samples. Differentially expressed genes were detected by ANOVA and probesets of statistical significance were defined by a-fold change > 2 and a FDR.5. RNA-SEQ. For m19z, and m19-humbbz comparison, three million CAR T cells were incubated with T3-mCD19 cells for 48 hr. Live CD4+ CAR T cells were sorted into Trizol. RNA was isolated according to manufacturer s instructions and evaluated for quality. The Genomic Core performed mrna enrichment and cdna library preparation using the Illumina Tru-seq stranded mrna sample prep kit. Final RNA-seq libraries were reviewed for size and quality on the Agilent TapeStation, followed by quantitative PCR-based quantitation with the Kapa Library Quantification Kit. The libraries sequenced on two NextSeq high-output 2x75 paired-end sequencing runs in order to generate approximately 4 million pairs of reads per sample. Sequence reads were aligned to the human reference genome in a splice-aware fashion using Tophat2 (1), allowing for accurate alignments of sequences across introns. Aligned reads were quantitated at the gene level using HTseq (2). Normalization, expression modeling, and difference testing were performed using DESeq (3). Quality control measures included custom scripts and RSeqC (4) to examine read count metrics, alignment fraction, chromosomal S3
4 alignment counts, expression distribution measures, and principle components analysis and hierarchical clustering. Differentially expressed genes were detected by ANOVA and probesets of statistical significance were defined by a-fold change > 4 and a FDR.1. Gene set enrichment analysis was performed (on the gene expression values) to analyze the enrichment of the gene sets using GSEA software ( C5 collection version v6. from the Molecular Signature Database (MSigDB v6. C5), which contains the expert-curated gene ontology (GO) gene sets, were used in the analysis. We used vertebrate homology resource ( to convert between homologues human and mouse genes. For all comparisons, data was collapsed to gene symbols. 1 permutations based on gene sets were performed. Gene sets were ranked according to false discovery rate (FDR) q-value. At the default FDR q-value cut-off within GSEA of.25, we identified 3 gene sets that are upregulated in m19z and 68 gene sets upregulated in. Microarray and RNA-SEQ data have been submitted to GEO (Gene Expression Omnibus) with the accession number GSE CD33-targeted CARs Anti-CD33 antibodies were developed at the Vanderbilt Antibody and Protein Resource using standard methods (5). Briefly, after completing a series of immunizations splenocytes of immunized mice were isolated and fused to a non-ig secreting myeloma cell line and grown in a semi-solid plate. Antibodysecreting clusters were identified in semi-solid plates and selected for clonal expansion in 96 well plates. S4
5 During expansion supernatant was collected and assayed for CD33 binding by ELISA as well as flow cytometry. Based on this screening hybridomas were selected for expansion and isolation of RNA, which was used to amplify IgH and IgL rearrangements. Based on the IgH and IgL rearrangements scfv were designed and cloned into the NcoI/NotI sites of our human CD19-targeted CAR in the SFG retroviral cassette. This allowed replacement of the anti-human CD19 scfv with anti-human CD33 scfv. These constructs were then used to produce gammaretroviral supernatant as described in Methods. In vivo NALM6 animal model of CD19-targeted CAR T cells The NALM6 leukemia mouse model has been described (6). Briefly, NALM6-GL cells were i.v. injected to NSG mice at 5x1 5 dose. Four days later, mice were treated with 3x1 5-1x1 6 human CD19 targeted CAR T cells. Human CD19 targeted CAR T cells with excess TRAF2 were made by CAR and mouse TRAF2 co-transduction or transduction with a bicistronic construct combining CAR and human TRAF2. Blood samples were collected weekly for flow cytometry. Leukemia burden was evaluated weekly using bioluminescence imaging on an IVIS system. Survival was monitored. Mice were sacrificed when they develop signs of progressive leukemia. S5
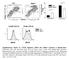
6 A m19 z CD4 CD44 CD8 CD62L B B cells/μl blood B6 m19 z (1E6) m19z (3E5) m19z (1E6) (3E5) (1E6) (3E5) (1E6) Thy1.1 T cells/μl blood B6 m19 z (1E6) m19z (3E5) m19z (1E6) (3E5) (1E6) (3E5) (1E6) Supplemental Figure 1. Immune phenotype of mcd19 targeted CAR T and dose titration of in vivo efficacy. (A) Immune phenotype of transduced T cells used in 5x1 6 dose in vivo study (Figure 1C&D). Cells were pre-gated on single live cells. (B) In vivo B cell killing and T cell persistence with T cell dose titration. After Eµ-ALL injection, different doses of T cells were given to 3 mg/kg CTX preconditioned mice. B (B22+CD19+) and T (CD3+Thy1.1+) cells in peripheral blood were quantitated 3 weeks after CAR T injection. Each dot indicates one mouse. Data are from one single experiment (n=34 total). S6
7 A C + 5 LTR GFP/ V H G/S V L EC TM G/S 3 LTR mcherry SD SA + 5 LTR GFP/ V H G/S V L EC TM CD3 G/S 3 LTR mcherry SD SA + GFP/ 5 LTR V H G/S V L EC TM CD28 CD3 G/S 3 LTR mcherry SD SA + 5 LTR V H G/S V EC TM m4-1bb CD3 G/S GFP/ L 3 LTR mcherry SD SA CD8 Leader CD8 B D CAR% 8% 6% 4% 2% % mcherry Protein L mcherry Protein L mcherry Protein L mcherry Protein L m19 z m19z E m19z m19-mbbz Supplemental Figure 2. Gene expression of fluorescent-protein tagged CAR T cells. (A) Schematic of genetic constructs for mcd19 targeted CARs. Shown are the long terminal repeats (LTR), packaging signal Ψ, splice donor (SD), splice acceptor (SA), VH and VL regions of the scfv (single-chain variable fragment), the extracellular hinge (EC), transmembrane (TM), and intracellular regions of the retroviral construct. G/S, (Gly4Ser1)3 linker sequence. (B) Comparison of fluorescence protein and Protein L as a method to evaluate CAR expression. One million T cells transduced with mcherry-tagged CARs were incubated with 1 µg Biotin-Protein L and then fluochrome-conjugated streptavidin. Cells were subjected to flow cytometry. Data are representative of 4 independent experiments. (C) Principal component S7
8 analysis (PCA) of mcd19-targeted CAR T cells stimulated with antigen. (D) Venn Diagram demonstrating the number of genes differentially expressed (n=25) in CAR T cells compared to both m19z and CAR T cells. (E) A heatmap of the 25 differentially expressed genes. The list of 25 genes is included in Supplemental Tables 1-4. For (C)-(E), CAR T cells with the m19z,, or CAR tagged to the fluorescent protein GFP were incubated with 3T3- mcd19 AAPC at 1:1 E:T ratio overnight, FACS-sorted, and lysed to isolate RNA. Each group of CAR T cells was transduced, stimulated, and sorted independently in triplicates. S8
9 A B C GFP Survival IFN (pg/ml) CD8 CD44 D % 5% CD3 CD4 CD62L % B cells/μl blood8 13 Days after leukemia WK 1 m19 z m19z **** **** WK 2 WK 4 *** ** **** *** **** WK 8 WK 12 TNF (pg/ml) m19 z m19z ** m19 z m19z m19δz m19z CAR T/μl blood * WK 1 **** ns B cells/femur6 15 WK 2 WK 4 WK 8 m19 z WK 12 m19z * Supplemental Figure 3. Fluorescent protein tagged CAR T cells function similarly to non-tagged counterparts. (A) Cytokines released by fluorescent protein tagged CAR T cells upon antigen stimulation. Day 4 CAR T cells were co-cultured with 3T3-mCD19 at 1:1 ratio for 24 hr. Supernatant was subjected to luminex assay for IFNγ and TNFα. (B) Immune phenotype of CAR T cells with a fluorescent protein tag. Day 4 CAR T cells were harvested, beads removed and subjected to flow cytometry. Cells were pre-gated on single live cells (top) or CD3+CAR+ cells (middle & bottom). (C) Survival (n=5), in vivo B cell killing and CAR T persistence in mice treated by CAR T cells with a fluorescent protein tag. Seven days after i.v. injection with 1X1 6 Eµ-ALL cells, mice were i.p. injected with CTX at 25 mg/kg and then one day later i.v. injected with 1X1 6 CAR T cells. Survival was monitored. BM was isolated 11 days after CAR T injection and subjected to flow cytometry. B (B22+CD19+) and CAR T (CD3+- CAR+) cells were counted using CountBright beads. Each dot indicates one mouse (n=3 per group). Data are from one single experiment. (D) B and CAR T cell counts over time in the blood after CAR T treatment. B6 mice were injected with CTX (25 mg/kg) and CAR T cells (3X1 5 ). Blood samples were collected over time and B and CAR T cell numbers were measured by flow cytometry (n=1 per group). Survival curve, logrank test; all other data, unpaired t test. *p<.5; **p<.1; ***p<.1; ****p<.1; ns, not significant. CAR T/femur m19 z m19z * S9
10 m19δz m19z m19-humbbz CD8 GFP CD44 CD4 CD62L Supplementary figure 4. Transduction efficiency and immune phenotype of mcd19 targeted CAR T cells for survival study (Figure 2D). Data are representative of four independent productions used to generate CAR T cells for the survival experiments of Figure 2D. For Supplemental Figure 4. Transduction efficiency and immune phenotype of mcd19 targeted CAR transduction efficiency (top panel), cells were pre-gated on single live cells. For immune Tphenotype cells for survival study (Figure 2D). Data representative four independent (middle and bottom panels), cells are were pre-gated onofsingle live CAR T productions used to generate CAR T cells for the survival experiments of Figure 2D. For transduction efficiency (top panel), (CD3+CAR+) cells. cells were pre-gated on single live cells. For immune phenotype (middle and bottom panels), cells were pre-gated on single live CAR T (CD3+CAR+) cells. S1
11 m19 z m19-humbbz CAR CD8 CD4 CD44 CD62L Supplemental Figure 5. Transduction efficiency and immune phenotype of CAR T cells used in irradiated CAR T study (Figure 3B-3C). Day 4 transduced cells were harvested, beads removed, stained with antibodies and subjected to flow cytometry. For transduction efficiency (Top panels), cells were pre-gated on single live cells. For immune phenotype (middle and bottom panels), cells were pregated on CD3+CAR+ cells. S11
12 Supplemental Figure 6. Differential gene expression of CD4+ m19-humbbz CAR T cells. T cells with the m19z,, or m19-humbbz CAR were incubated with 3T3-mCD19 AAPC at 1:1 E:T ratio, FACS-sorted, and lysed to isolate RNA. Each group of CAR T cells was transduced, stimulated, and sorted independently in biologic triplicates. (A) PCA of mouse CD19-targeted CAR T cells stimulated with antigen. (B) Venn Diagram demonstrating the number of genes differentially expressed in m19-humbbz CAR T cells compared to m19z and CAR T cells. (C) GSEA demonstrates gene sets correlating to NF-κ B regulatory pathways are differentially expressed in m19-humbbz CAR T cells versus m19z or CAR T cells. S12
13 Untrans h19bbz mut1 mut2 mut3 mut4 CD3 CD8 CAR CD4 Supplemental Figure 7. CAR expression and CD4/CD8 subsets of human CD19 targeted CAR T cells for Figure 5E-G. For CAR expression (top), cells were pre-gated on single live cells. For immune phenotype (bottom), cells were pre-gated on CD3+CAR+ cells. S13
14 m19 z WT WT TRAF1 -/- m19-humbbz WT m19-humbbz TRAF1-/- CD3 CD8 CD4 CD44 GFP CD62L Supplemental Figure 8. Transduction efficiency and immune phenotype of mcd19 targeted wild type (WT) and Traf1 -/- CAR T cells used for in vivo study (Figure 6D). Day 4 transduced cells were harvested, beads removed, stained with antibodies and subjected to flow cytometry. For transduction efficiency (top panel), cells were pre-gated on single live cells. For CD4/CD8 subsets (middle panel) and memory subsets (bottom panel) cells were pre-gated on single live CAR T (CD3+CAR+) cells. S14
15 A B C Luminescence **** **** m19 z m19-humbbz mut1 CD8+IFN + CAR T% 5% 4% 3% 2% 1% % m19 z ** * m19-humbbz mut1 CD8+Bclxl+ CAR T(%) 4% 3% 2% 1% % m19 z m19z m19-humbbz D m19-humbbz mut1 ns ** * mut1 CAR E F G 2 * 4 * ns ns 15 * ** 3 *** **** 1 2 B cell/μl blood CD3 5 m19-humbbz mut1 CAR T cell/μl blood 1 m19-humbbz mut1 Donor T cell/μl blood ns m19-humbbz ns ** *** mut1 Supplemental Figure 9. Mutated CAR T cells have increased NF-k B signaling, improved cytokine production, anti-apoptosis, and in vivo function. (A) NF-k B signaling in mcd19 targeted CAR T cells. CAR T cells derived from NF-k B-RE-luc transgenic mice were cocultured with 3T3-mCD19 for 4 hr. Cell lysates were prepared and subjected to a luciferase assay. Bioluminescence was measured and correlates to NF-k B signaling. Data are representative of three independent experiments. (B) Intracellular IFNg and (C) BCL-XL expression in CD8+ CAR T cells stimulated with 3T3-mCD19. Data are representative of two independent experiments done in triplicate. (D) CAR expression in T cells used for in vivo study below. Cells were pre-gated on single live cells. (E) B cells (CD19+B22+) in the blood 1 week after CAR T cells injection. (F) CAR T cells (CD3+GFP+) in the blood 1 week after CAR T cells injection. (G) Donor T cells (CD3+Thy1.1+) in the blood 1 week after CAR T cells injection. (E-G) Each dot indicates one mouse (n=1 per group). Bar graph shown as mean ± SD. All data, unpaired t test. *p<.5; **p<.1; ***p<.1; ****p<.1; ns, not significant. S15
16 A Untrans h19bbz +TRAF1 +TRAF2 +TRAF3 TRAF TRAF CAR B Untrans h19bbz +TRAF1 +TRAF2 +TRAF3 C IFN (pg/ml) CAR h19bbz +TF1 ns +TF2 +TF3 TNF pg/ml h19bbz +TF1 +TF2 +TF3 IL2 pg/ml h19bbz +TF1 +TF2 +TF3 IL6 pg/ml h19bbz +TF1 +TF2 +TF3 Supplemental Figure 1. TRAF and CAR co-expression in human CD19-targeted CAR T cells. (A) CAR and TRAF expression in hcd19 targeted CAR T cells before antigen stimulation for viability, proliferation, and cytotoxicity assays (Figure 7B-D). Cells were pre-gated on single live cells. Data are one representative of 3 healthy donors. (B) CAR and TRAF expression in hcd19 CAR T cells 3 days after stimulation with 3T3-hCD19. Cells were pre-gated on single live cells. Data are from one experiment in triplicate. Numbers indicate percentages of gated cells. (C) Cytokine production. CAR T cells were activated on 3T3-hCD19 at 1:1 ratio. After 24 hr supernatant were harvested and cytokines were measured by ELLA. Bar graphs shown as mean ± SD. Data are one representative of 3 different healthy donors in triplicate. All data, unpaired t test. ns, not significant. S16
17 A B C 6 1% Survival 5% % Days after NALM6-GL Untrans h19bbz +TF2 * ** ns 2 NALM6/μl blood1 Untrans Untrans h19bbz D E F 1% Untrans h19bbz 8 +TF2 6 5% 4 ns Survival % Days after NALM6-GL ** *** ns CD19+ cells/μl blood 4 2 h19bbz ns +TF2 +TF2 Total counts Total counts Untrans Untrans h19bbz h19bbz ns +TF2 * +TF2 Supplemental Figure 11. TRAF2 over-expressed h19bbz CAR T cells show similar in vivo efficacy to h19bbz CAR T cells in an aggressive leukemia model. NSG mice were implanted with leukemia by i.v. injecting 5x1 5 NALM6-GL cells. Four days later, mice were i.v. injected with 3x1 5-1x1 6 CAR T cells. Blood samples were collected weekly. Leukemia burden was evaluated weekly using bioluminescence imaging. Survival was monitored. For (A-C), TRAF2 was co-transduced to make CAR T cells, n=15 total, and data are from one experiment. For (D-F), a bicistronic construct expressing CAR and TRAF2 was transduced to make CAR T cells. Survival data are pooled from two independent experiments (n=26), and counts are from one experiment. Each dot indicates a mouse. Survival curve, logrank test; all other data, unpaired t test. *p<.5; **p<.1; ns, not significant. S17
18 Supplemental Table 1. Probesets increased in m19z and vs. CAR T cells Probeset ID Gene Symbol Fold- Change Probeset ID Gene Symbol Fold- Change _at Gzmf _at Serpinb _at Serpinb9b _at Plek _at Ccl _a_at Klrk _at Il _at Paqr _at Gstm _a_at P2ry _at Klra _a_at Ctbp _a_at Dhrs _a_at Metrnl _at Rgs _at Gpd _at Pdcd _at Ccr _at Gng _at Tmem _at Gzmg _at Gzmd _at Myo _at Sh3bp _at Irf _s_at S1a _at Serpine _at Swap _at Rgs _at Ccr _at Irf _a_at Bcat _at Nrgn _x_at Gzmd _at Hhex _at Prss _at Myo _at Spin _at Serpinb1b _a_at Ehd _at Ifng _at Spry _at Cd _at Upp _a_at Prf _at Tmem _at Klrg _at Sypl _at Havcr _s_at Pkp _at Gpx _at Sypl _a_at Lat _a_at Cdkn2a _at Serpinb6b _at Cd _at Klrb1c _at Adssl _a_at Nr4a _at Pkd _at Plek _at Myo1e _at Mcpt _s_at Sypl _at Spp _at Galnt _a_at Gem _x_at Myb _a_at Tnfrsf _at Cyp _at Scel _a_at Myb _at Sult2b _a_at Zbtb _x_at Gzme _a_at Myb _at Lhx _a_at Anxa2 2.3 S18
19 _at Gzmc _at Gipc _a_at Klf _at Ctbp _at Cd _at Nek _at _a_at Bcl2l _at Slc17a _at Rai _a_at Tjp _a_at Uxs _at LOC _at Kcna _a_at Ccl _at Exo _at Gzmd _at Tmem _a_at Rgs _a_at Bmp _at Epas _a_at Txk _at Slc17a _at Litaf _at Plek _at Litaf _at Fcer1g _a_at Dusp _at Oit _at Gm _at Tnfrsf _a_at Bspry _at Srgap _a_at Lyn _at Ccl _at Eomes _at Rgs _a_at Ypel _a_at Cd _s_at Ypel _at Lgals _at Cables _a_at Ier _a_at Rnf _a_at Slc37a _x_at Klra _at Tubb _s_at Klhdc _at Xcl _s_at Mt _at Bmp _a_at Gm _at Cd _a_at Abcb _at Dsc _at Pdcd1lg _at Tmem2 2.9 S19
20 Supplemental Table 2. Probesets increased in vs. m19z and CAR T cells Probeset ID Gene Symbol Fold- Change Probeset ID Gene Symbol Fold- Change 14231_at Fos _at Mbd _at Dusp _at Tlr _s_at Igfbp _at St8sia _at Cox7c _a_at Snord _at Btf _a_at Morc _a_at Zfp _at Cacnb _s_at Cox7c _at Gm _at Ubl _at Zfp _s_at Ighm _at Fam11b _at Klf _x_at Gm _at Actn _at Ppp1r15a _at Tcf _at Ndufa _a_at Igfbp _at Gm _at Nsg _at St8sia _at LOC _at Eif2s _a_at Slamf _at Itgb _a_at Ighm _at Rbm _at Cyp2s _at Actn _x_at H2-Aa _at Nfkbia _at Tcf _at Abca _at Fosb _at Cmah _at Klf _a_at Ssbp _at Itgae _at Itga _at Gm _at Gm _at Jun _at Adcy _at C _s_at Dbp _at Tdrp _at Srsf _a_at Ifi _x_at Stx1a _s_at Trib _at Psma _s_at Nfkbia _at Sfswap _at Ifit _at Tmem _at Itga _x_at Gm _at Nfkbia _x_at Snhg _at Slamf _at Cd _at Rbm _a_at Fbxl S2
21 Supplemental Table 3. Probesets differentially expressed in m19z vs. CAR T cells Probeset ID Gene Symbol Fold- Change Probeset ID Gene Symbol Fold- Change _x_at Sdc _at Crtam _a_at Ugt1a _at Smpdl3b _x_at LOC _x_at _x_at Ceacam _a_at Endod _at Tfpi _at Fdps _at Sdc _at Zeb _at Penk _at Pglyrp _s_at Klrb1b _at Tfrc _s_at Ugt1a _at Pdia _at Antxr _at Tmbim _at Gstt _s_at Ceacam _at Ccl _at Cyp _at Ccl _at Klhdc _a_at Klrc _s_at Ceacam _at Mycn _at Sema7a _at Khdc1a _at Phactr _at Coro2a _at Lrrk _at Vcl _at Napsa _a_at Rgs _at Endod _at Itih _at Car _a_at Chn _at Mtm _a_at Rgs _at Sdf2l _s_at Ceacam _x_at Vat _at Nrp _at Cd _x_at Ceacam _a_at Dusp _s_at Ltbp _at Plk _x_at Klra _at Cbx _at Pkp _x_at Endod _at Klrc _at Myo _a_at Tfrc _at Lag _at Nrp _a_at LOC _at Fam11c _a_at Hmgcs _at _at Mt _a_at Ugt1a _a_at Cd _at Scd _at Ndrg _x_at Ceacam _at Shmt _at Tg _at Uhrf1 2. X686_5_at _at Hbegf _a_at Tacc _at Cdkn2b 2. S21
22 _at Chn _at Ndc _at Klrb1b _at Hist3h2ba _at Lad _at Cd _at Alcam _at Taf _at Zfp _at Rapgef _at Slc25a _at Myl _at Ltbp _at 18132O8Rik _at Nrp _at Enah -2.3 X57349_5_at Tfrc _at Nfkbiz _at Klrb1a _at Pdlim _at Calcb 2.3 S22
23 Supplemental Table 4. Probesets differentially expressed in vs. CAR T cells Probeset ID Gene Symbol Fold- Change Probeset ID Gene Symbol Fold- Change _at Esm _a_at Shisa _s_at Eps _a_at Elmo _a_at Atp1b _at Ltb4r _at Ctsg _a_at Tnfsf _at Eps _at Alpk _at Sepn _a_at Pdzrn _a_at Lmna _at Tirap _at Clnk _a_at Cysltr _at Gzmk _a_at Naip _at Basp _at Pilrb _at Gcnt _a_at Eps _at E2Ri _at Tiam _at Stk32c _at Osbpl1a _at Tdgf _a_at Slc37a _a_at Rhoq _at Zak _a_at Tcf _x_at Cnn _at Tcf _a_at Cmpk _at Tspan _at Nfkbia _at Tyrobp _x_at Cnn _at Col6a _at Ldlrap _x_at Arhgap _a_at Capn _a_at Fndc _at Oas _a_at Fhl _a_at Rbm _at Shisa _at Sh3bp _at Marcks _at Ltb _s_at Gcnt _at Srsf _at Piwil _at Frat _x_at Marcks _a_at Hist1h1c _at Serpinb1a _at Thra _a_at Cma _at Mphosph _at Gcnt _at Ifit _at Tnfsf _at Klf _at Chpt _at Pou6f _at Cpa _s_at H2-Ab _a_at Ptger _x_at Tgm _at Hunk _at Tlr _at Marcks _x_at Nfix _a_at Frmd4b _at Ptgir _at Eomes _at S23
24 142364_at Gpr _x_at Tgm _x_at Tcf _a_at Enpp _at Aoah _a_at Sfswap _at Plxdc _at Abca _at Acvr _at Ncf _s_at Gcnt _a_at Serinc _at Dusp _at Setd1a _at Eps8l _at Slamf _at Ptgr _at Rabgap1l _at Prnp _at Gdap _at Tcf _at Sh3bp _at P2ry _at _a_at H2-Q _at Fas _at Chpt _a_at Cd _s_at Fam84a _a_at Slamf _at Dab2ip _at Dbp _at Tcf _at Ssbp _at S1a _at Il6ra _at Sox _at Thra _x_at Hba-a _a_at Idh _at Errfi _a_at Tspan _a_at Chpt _a_at Tnfrsf _at Plxdc _at Ccl _at Ell _at Ccr _x_at Marcks _at Cd _at Marcks _a_at Nfix _at Ccr _at Cd S24
25 SUPPLEMENTAL REFERENCES 1. Trapnell C, Pachter L, and Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 29;25(9): Anders S, Pyl PT, and Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 215;31(2): Anders S, and Huber W. Differential expression analysis for sequence count data. Genome Biol. 21;11(1):R Wang L, Wang S, and Li W. RSeQC: quality control of RNA-seq experiments. Bioinformatics. 212;28(16): Markham NO, Cooper T, Goff M, Gribben EM, Carnahan RH, and Reynolds AB. Monoclonal antibodies to DIPA: a novel binding partner of p12-catenin isoform 1. Hybridoma (Larchmt). 212;31(4): Zhao Z, Condomines M, van der Stegen SJC, Perna F, Kloss CC, Gunset G, Plotkin J, and Sadelain M. Structural Design of Engineered Costimulation Determines Tumor Rejection Kinetics and Persistence of CAR T Cells. Cancer Cell. 215;28(4): S25
CPM (x 10-3 ) Tregs +Teffs. Tregs alone ICOS CLTA-4
 A 2,5 B 4 Number of cells (x 1-6 ) 2, 1,5 1, 5 CPM (x 1-3 ) 3 2 1 5 1 15 2 25 3 Days of culture 1/1 1/2 1/4 1/8 1/16 1/32 Treg/Teff ratio C alone alone alone alone CD25 FoxP3 GITR CD44 ICOS CLTA-4 CD127
A 2,5 B 4 Number of cells (x 1-6 ) 2, 1,5 1, 5 CPM (x 1-3 ) 3 2 1 5 1 15 2 25 3 Days of culture 1/1 1/2 1/4 1/8 1/16 1/32 Treg/Teff ratio C alone alone alone alone CD25 FoxP3 GITR CD44 ICOS CLTA-4 CD127
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
CD4 + T cells recovered in Rag2 / recipient ( 10 5 ) Heart Lung Pancreas
 a CD4 + T cells recovered in Rag2 / recipient ( 1 5 ) Heart Lung Pancreas.5 1 2 4 6 2 4 6 Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas b Heart Lung Pancreas Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas
a CD4 + T cells recovered in Rag2 / recipient ( 1 5 ) Heart Lung Pancreas.5 1 2 4 6 2 4 6 Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas b Heart Lung Pancreas Ctla4 +/+ Ctla4 / Ctla4 / Lung Ctla4 / Pancreas
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Supplemental Figure 1. Protein L
 Supplemental Figure 1 Protein L m19delta T m1928z T Suppl. Fig 1. Expression of CAR: B6-derived T cells were transduced with m19delta (left) and m1928z (right) to generate CAR T cells and transduction
Supplemental Figure 1 Protein L m19delta T m1928z T Suppl. Fig 1. Expression of CAR: B6-derived T cells were transduced with m19delta (left) and m1928z (right) to generate CAR T cells and transduction
Supplemental Table 1: Enriched GO categories per cluster
 Supplemental Table 1: Enriched GO categories per cluster Cluster Enriched with #genes Raw p-value Corrected p-value Frequency in set (%) Cluster 1 immune response - GO:0006955 32 2.36E-14 0.001 13.5 Cluster
Supplemental Table 1: Enriched GO categories per cluster Cluster Enriched with #genes Raw p-value Corrected p-value Frequency in set (%) Cluster 1 immune response - GO:0006955 32 2.36E-14 0.001 13.5 Cluster
and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the
 Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Figure 1. Characterization of basophils after reconstitution of SCID mice
 Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplemental Methods RNA sequencing experiment
 Supplemental Methods RNA sequencing experiment Mice were euthanized as described in the Methods and the right lung was removed, placed in a sterile eppendorf tube, and snap frozen in liquid nitrogen. RNA
Supplemental Methods RNA sequencing experiment Mice were euthanized as described in the Methods and the right lung was removed, placed in a sterile eppendorf tube, and snap frozen in liquid nitrogen. RNA
Nature Medicine: doi: /nm.3922
 Title: Glucocorticoid-induced tumor necrosis factor receptor-related protein co-stimulation facilitates tumor regression by inducing IL-9-producing helper T cells Authors: Il-Kyu Kim, Byung-Seok Kim, Choong-Hyun
Title: Glucocorticoid-induced tumor necrosis factor receptor-related protein co-stimulation facilitates tumor regression by inducing IL-9-producing helper T cells Authors: Il-Kyu Kim, Byung-Seok Kim, Choong-Hyun
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Table S1. Differentially expressed transcripts between hepatic inkt cells. sorted form WT and plck-hcd1d tg mice. Differentially expressed
 Supplementary Information Table S1. Differentially expressed transcripts between hepatic inkt cells sorted form and plck-hcd1d tg mice. Differentially expressed transcripts identified by gene expression
Supplementary Information Table S1. Differentially expressed transcripts between hepatic inkt cells sorted form and plck-hcd1d tg mice. Differentially expressed transcripts identified by gene expression
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was
 Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
Supplementary Figure 1. mrna expression of chitinase and chitinase-like protein in splenic immune cells. Each splenic immune cell population was sorted by FACS. Surface markers for sorting were CD11c +
CD80 and PD-L2 define functionally distinct memory B cell subsets that are. Griselda V Zuccarino-Catania, Saheli Sadanand, Florian J Weisel, Mary M
 Supplementary Figures CD8 and PD-L define functionally distinct memory B cell subsets that are independent of antibody isotype Running title: Memory B Cell Subset Function Griselda V Zuccarino-Catania,
Supplementary Figures CD8 and PD-L define functionally distinct memory B cell subsets that are independent of antibody isotype Running title: Memory B Cell Subset Function Griselda V Zuccarino-Catania,
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured
 Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Cell-bound Cetuximab reduces EGFR staining intensity. Blood
 Antibody-mediated depletion of CD19-CAR T cells Supplemental 1 Supplemental Materials Supplemental Figure 1. Supplemental Figure 1. Cell-bound Cetuximab reduces EGFR staining intensity. Blood cells were
Antibody-mediated depletion of CD19-CAR T cells Supplemental 1 Supplemental Materials Supplemental Figure 1. Supplemental Figure 1. Cell-bound Cetuximab reduces EGFR staining intensity. Blood cells were
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Supplemental Information. Augmentation of Antitumor Immunity by Human. and Mouse CAR T Cells Secreting IL-18
 Cell Reports, Volume 20 Supplemental Information Augmentation of Antitumor Immunity by Human and Mouse CAR T Cells Secreting IL-18 Biliang Hu, Jiangtao Ren, Yanping Luo, Brian Keith, Regina M. Young, John
Cell Reports, Volume 20 Supplemental Information Augmentation of Antitumor Immunity by Human and Mouse CAR T Cells Secreting IL-18 Biliang Hu, Jiangtao Ren, Yanping Luo, Brian Keith, Regina M. Young, John
Supplementary Figure 1
 Supplementary Figure 1 Expression of apoptosis-related genes in tumor T reg cells. (a) Identification of FOXP3 T reg cells by FACS. CD45 + cells were gated as enriched lymphoid cell populations with low-granularity.
Supplementary Figure 1 Expression of apoptosis-related genes in tumor T reg cells. (a) Identification of FOXP3 T reg cells by FACS. CD45 + cells were gated as enriched lymphoid cell populations with low-granularity.
W/T Itgam -/- F4/80 CD115. F4/80 hi CD115 + F4/80 + CD115 +
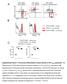
 F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
Supplementary. presence of the. (c) mrna expression. Error. in naive or
 Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
Figure 1. (a) Naive CD4 + T cells were activated in the presence of the indicated cytokines for 3 days. Enpp2 mrna expression was measured by qrt-pcrhr, infected with (b, c) Naive CD4 + T cells were activated
Nature Immunology: doi: /ni Supplementary Figure 1. Id2 and Id3 define polyclonal T H 1 and T FH cell subsets.
 Supplementary Figure 1 Id2 and Id3 define polyclonal T H 1 and T FH cell subsets. Id2 YFP/+ (a) or Id3 GFP/+ (b) mice were analyzed 7 days after LCMV infection. T H 1 (SLAM + CXCR5 or CXCR5 PD-1 ), T FH
Supplementary Figure 1 Id2 and Id3 define polyclonal T H 1 and T FH cell subsets. Id2 YFP/+ (a) or Id3 GFP/+ (b) mice were analyzed 7 days after LCMV infection. T H 1 (SLAM + CXCR5 or CXCR5 PD-1 ), T FH
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplemental Information. Aryl Hydrocarbon Receptor Controls. Monocyte Differentiation. into Dendritic Cells versus Macrophages
 Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
** *** population. (A) Representative FACS plots showing the gating strategy for T N
 Sca-1 (MFI) CD122 (MFI) CD44 A CD8 + gated: resting spleen CD8 + gated: T Mem -derived CD8 + gated: T N -derived, expanded alone B CD62L CD44 low CD62L +, resting CD44 high, T Mem -derived CD44 low CD62L
Sca-1 (MFI) CD122 (MFI) CD44 A CD8 + gated: resting spleen CD8 + gated: T Mem -derived CD8 + gated: T N -derived, expanded alone B CD62L CD44 low CD62L +, resting CD44 high, T Mem -derived CD44 low CD62L
X P. Supplementary Figure 1. Nature Medicine: doi: /nm Nilotinib LSK LT-HSC. Cytoplasm. Cytoplasm. Nucleus. Nucleus
 a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
Nature Medicine doi: /nm.3957
 Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1
 ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
ZH, Li et al, page 1 ECM1 controls T H 2 cell egress from lymph nodes through re-expression of S1P 1 Zhenhu Li 1,4,Yuan Zhang 1,4, Zhiduo Liu 1, Xiaodong Wu 1, Yuhan Zheng 1, Zhiyun Tao 1, Kairui Mao 1,
Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel
 Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
Supplementary Figures 1-8 Crucial role for human Toll-like receptor 4 in the development of contact allergy to nickel Marc Schmidt 1,2, Badrinarayanan Raghavan 1,2, Verena Müller 1,2, Thomas Vogl 3, György
Bioassays for Quality Control of Cell & Gene Therapy Products
 Bioassays for Quality Control of Cell & Gene Therapy Products Erik Rutjens, Cell & Gene Therapy, Novartis Pharma AG CASSS Bioassays, Silver Spring, March2015 CTL019 Introduction CARTs = Chimeric Antigen
Bioassays for Quality Control of Cell & Gene Therapy Products Erik Rutjens, Cell & Gene Therapy, Novartis Pharma AG CASSS Bioassays, Silver Spring, March2015 CTL019 Introduction CARTs = Chimeric Antigen
Supplementary Materials and Methods
 Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Materials and Methods Whole Mount X-Gal Staining Whole tissues were collected, rinsed with PBS and fixed with 4% PFA. Tissues were then rinsed in rinse buffer (100 mm Sodium Phosphate ph
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Bezzi et al., Supplementary Figure 1 *** Nature Medicine: doi: /nm Pten pc-/- ;Zbtb7a pc-/- Pten pc-/- ;Pml pc-/- Pten pc-/- ;Trp53 pc-/-
 Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Gr-1 Gr-1 Gr-1 Bezzi et al., Supplementary Figure 1 a Gr1-CD11b 3 months Spleen T cells 3 months Spleen B cells 3 months Spleen Macrophages 3 months Spleen 15 4 8 6 c CD11b+/Gr1+ cells [%] 1 5 b T cells
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5
 sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD-
 Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Protein SD Units (P-value) Cluster order
 SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human
 Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Supplemental Information. Checkpoint Blockade Immunotherapy. Induces Dynamic Changes. in PD-1 CD8 + Tumor-Infiltrating T Cells
 Immunity, Volume 50 Supplemental Information Checkpoint Blockade Immunotherapy Induces Dynamic Changes in PD-1 CD8 + Tumor-Infiltrating T Cells Sema Kurtulus, Asaf Madi, Giulia Escobar, Max Klapholz, Jackson
Immunity, Volume 50 Supplemental Information Checkpoint Blockade Immunotherapy Induces Dynamic Changes in PD-1 CD8 + Tumor-Infiltrating T Cells Sema Kurtulus, Asaf Madi, Giulia Escobar, Max Klapholz, Jackson
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1. Galectin-3 is present within tumors. (A) mrna expression levels of Lgals3 (galectin-3) and Lgals8 (galectin-8) in the four classes of cell lines as determined
Supplementary Figures Supplementary Fig. 1. Galectin-3 is present within tumors. (A) mrna expression levels of Lgals3 (galectin-3) and Lgals8 (galectin-8) in the four classes of cell lines as determined
Supplemental Figure 1. CD69 antigen-response curves of CAR engrafted Jurkat T cells. Supplemental Figure 2.
 Supplemental Figure 1. CD69 antigen-response curves of CAR engrafted Jurkat T cells. To evaluate the antigen sensitivity of mutant CARs transduced Jurkat T cells were stimulated with varying concentrations
Supplemental Figure 1. CD69 antigen-response curves of CAR engrafted Jurkat T cells. To evaluate the antigen sensitivity of mutant CARs transduced Jurkat T cells were stimulated with varying concentrations
AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification
 BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
supplemental Figure 1
 supplemental Figure 1 A T cell T1 anti-ny-eso-117-16/hla-a*:1 CDζ CH/CH scfv B T cell T1 anti-ny-eso-117-16/hla-a*:1 CDζ CH/CH scfv C T cell BW1/6 anti-cea CDζ CH/CH scfv supplemental Figure 1.79.9.87
supplemental Figure 1 A T cell T1 anti-ny-eso-117-16/hla-a*:1 CDζ CH/CH scfv B T cell T1 anti-ny-eso-117-16/hla-a*:1 CDζ CH/CH scfv C T cell BW1/6 anti-cea CDζ CH/CH scfv supplemental Figure 1.79.9.87
Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated
 1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
% of live splenocytes. STAT5 deletion. (open shapes) % ROSA + % floxed
 Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Supp. Figure 1. a 14 1 1 8 6 spleen cells (x1 6 ) 16 % of live splenocytes 5 4 3 1 % of live splenocytes 8 6 4 b 1 1 c % of CD11c + splenocytes (closed shapes) 8 6 4 8 6 4 % ROSA + (open shapes) % floxed
Activation of a PML/TP53 axis underlies acute promyelocytic. leukemia cure
 Activation of a PML/TP53 ais underlies acute promyelocytic leukemia cure Julien Ablain, Kim Rice, Hassane Soilihi, Aurélien de Reynies, Saverio Minucci & Hugues de Thé a PLZF-RARA vs PLZF-RARA pathway
Activation of a PML/TP53 ais underlies acute promyelocytic leukemia cure Julien Ablain, Kim Rice, Hassane Soilihi, Aurélien de Reynies, Saverio Minucci & Hugues de Thé a PLZF-RARA vs PLZF-RARA pathway
Supplementary Figure Legends. group) and analyzed for Siglec-G expression utilizing a monoclonal antibody to Siglec-G (clone SH2.1).
 Supplementary Figure Legends Supplemental Figure : Naïve T cells express Siglec-G. Splenocytes were isolated from WT B or Siglec-G -/- animals that have not been transplanted (n= per group) and analyzed
Supplementary Figure Legends Supplemental Figure : Naïve T cells express Siglec-G. Splenocytes were isolated from WT B or Siglec-G -/- animals that have not been transplanted (n= per group) and analyzed
Regulating the Regulators for Cancer Immunotherapy: LAG-3 Finally Catches Up. Drew Pardoll Sidney Kimmel Cancer Center Johns Hopkins
 Regulating the Regulators for Cancer Immunotherapy: LAG-3 Finally Catches Up Drew Pardoll Sidney Kimmel Cancer Center Johns Hopkins The hostile immune microenvironment within a tumor Stat3 Stat3 NK Lytic
Regulating the Regulators for Cancer Immunotherapy: LAG-3 Finally Catches Up Drew Pardoll Sidney Kimmel Cancer Center Johns Hopkins The hostile immune microenvironment within a tumor Stat3 Stat3 NK Lytic
TNFSF13B tumor necrosis factor (ligand) superfamily, member 13b NF-kB pathway cluster, Enrichment Score: 3.57
 Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplemental Table 1. Primer sequences for transcript analysis
 Supplemental Table 1. Primer sequences for transcript analysis Primer Sequence (5 3 ) Primer Sequence (5 3 ) Mmp2 Forward CCCGTGTGGCCCTC Mmp15 Forward CGGGGCTGGCT Reverse GCTCTCCCGGTTTC Reverse CCTGGTGTGCCTGCTC
Supplemental Table 1. Primer sequences for transcript analysis Primer Sequence (5 3 ) Primer Sequence (5 3 ) Mmp2 Forward CCCGTGTGGCCCTC Mmp15 Forward CGGGGCTGGCT Reverse GCTCTCCCGGTTTC Reverse CCTGGTGTGCCTGCTC
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Comparison of in vitro transfection efficiency of HBV plasmids between genotypes HBV expressing plasmids (5
 Supplemental Materials Figures legend Supplemental Table 1, 2 and 3 Supplemental Figure 1, Figure 2 and Figure 3 Supplemental Figure 1 Comparison of in vitro transfection efficiency of HBV plasmids between
Supplemental Materials Figures legend Supplemental Table 1, 2 and 3 Supplemental Figure 1, Figure 2 and Figure 3 Supplemental Figure 1 Comparison of in vitro transfection efficiency of HBV plasmids between
ILC1 and ILC3 isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the ILC1, ILC2 and
 Supplementary Methods and isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the, ILC2 and subsets. For this purpose we performed intracellular flow cytometry
Supplementary Methods and isolation and culture Following cell sorting, we confirmed that the recovered cells belonged to the, ILC2 and subsets. For this purpose we performed intracellular flow cytometry
Supplementary information. The proton-sensing G protein-coupled receptor T-cell death-associated gene 8
 1 Supplementary information 2 3 The proton-sensing G protein-coupled receptor T-cell death-associated gene 8 4 (TDAG8) shows cardioprotective effects against myocardial infarction 5 Akiomi Nagasaka 1+,
1 Supplementary information 2 3 The proton-sensing G protein-coupled receptor T-cell death-associated gene 8 4 (TDAG8) shows cardioprotective effects against myocardial infarction 5 Akiomi Nagasaka 1+,
Supplemental Information. Granulocyte-Monocyte Progenitors and. Monocyte-Dendritic Cell Progenitors Independently
 Immunity, Volume 47 Supplemental Information Granulocyte-Monocyte Progenitors and Monocyte-endritic ell Progenitors Independently Produce Functionally istinct Monocytes lberto Yáñez, Simon G. oetzee, ndre
Immunity, Volume 47 Supplemental Information Granulocyte-Monocyte Progenitors and Monocyte-endritic ell Progenitors Independently Produce Functionally istinct Monocytes lberto Yáñez, Simon G. oetzee, ndre
SUPPLEMENT Supplementary Figure 1: (A) (B)
 SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice
 Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Figure 1
 Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with
 Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
Supplementary Figure S1. PTPN2 levels are not altered in proliferating CD8+ T cells. Lymph node (LN) CD8+ T cells from C57BL/6 mice were stained with CFSE and stimulated with plate-bound α-cd3ε (10µg/ml)
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
Supplemental Figure 1. Activated splenocytes upregulate Serpina3g and Serpina3f expression.
 Relative Serpin expression 25 2 15 1 5 Serpina3f 1 2 3 4 5 6 8 6 4 2 Serpina3g 1 2 3 4 5 6 C57BL/6 DBA/2 Supplemental Figure 1. Activated splenocytes upregulate Serpina3g and Serpina3f expression. Splenocytes
Relative Serpin expression 25 2 15 1 5 Serpina3f 1 2 3 4 5 6 8 6 4 2 Serpina3g 1 2 3 4 5 6 C57BL/6 DBA/2 Supplemental Figure 1. Activated splenocytes upregulate Serpina3g and Serpina3f expression. Splenocytes
Venky Ramakrishna PhD Celldex Therapeutics, Hampton NJ, USA
 Antibody Targeting of Human CD27 with Varlilumab (CDX-1127) identifies genes and pathways related to inflammation Venky Ramakrishna PhD Celldex Therapeutics, Hampton NJ, USA www.celldextherapeutics.com
Antibody Targeting of Human CD27 with Varlilumab (CDX-1127) identifies genes and pathways related to inflammation Venky Ramakrishna PhD Celldex Therapeutics, Hampton NJ, USA www.celldextherapeutics.com
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/8/352/352ra110/dc1 Supplementary Materials for Spatially selective depletion of tumor-associated regulatory T cells with near-infrared photoimmunotherapy
www.sciencetranslationalmedicine.org/cgi/content/full/8/352/352ra110/dc1 Supplementary Materials for Spatially selective depletion of tumor-associated regulatory T cells with near-infrared photoimmunotherapy
Supplementary Data. Treg phenotype
 Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr
 Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr at day 0, 1, 4, 10 and 21 post- muscle injury. (b)
Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr at day 0, 1, 4, 10 and 21 post- muscle injury. (b)
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
HTG EdgeSeq Immuno-Oncology Assay Gene List
 A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
A2M ABCB1 ABCB11 ABCC2 ABCG2 ABL1 ABL2 ACTB ADA ADAM17 ADGRE5 ADORA2A AICDA AKT3 ALCAM ALO5 ANA1 APAF1 APP ATF1 ATF2 ATG12 ATG16L1 ATG5 ATG7 ATM ATP5F1 AL BATF BA BCL10 BCL2 BCL2L1 BCL6 BID BIRC5 BLNK
B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were
 Supplementary Methods Mice. B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were purchased from The Jackson Laboratory. Real-time PCR Total cellular RNA was extracted from the
Supplementary Methods Mice. B6.SJL (Ly5.2) mice were obtained from Taconic Farms. Rag1-deficient mice were purchased from The Jackson Laboratory. Real-time PCR Total cellular RNA was extracted from the
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and
 Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF
 CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
ACTR (Antibody Coupled T-cell Receptor): A universal approach to T-cell therapy
 ACTR (Antibody Coupled T-cell Receptor): A universal approach to T-cell therapy European Medicines Agency Workshop on Scientific and Regulatory Challenges of Genetically Modified Cell-based Cancer Immunotherapy
ACTR (Antibody Coupled T-cell Receptor): A universal approach to T-cell therapy European Medicines Agency Workshop on Scientific and Regulatory Challenges of Genetically Modified Cell-based Cancer Immunotherapy
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic
 Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figures
 Supplementary Figures Supplementary Fig. 1. Surface thiol groups and reduction of activated T cells. (a) Activated CD8 + T-cells have high expression levels of free thiol groups on cell surface proteins.
Supplementary Figures Supplementary Fig. 1. Surface thiol groups and reduction of activated T cells. (a) Activated CD8 + T-cells have high expression levels of free thiol groups on cell surface proteins.
BET bromodomain inhibition enhances T cell persistence and function in adoptive immunotherapy models
 BET bromodomain inhibition enhances T cell persistence and function in adoptive immunotherapy models Yuki Kagoya,, Cheryl H. Arrowsmith, Naoto Hirano J Clin Invest. 2016;126(9):3479-3494. https://doi.org/10.1172/jci86437.
BET bromodomain inhibition enhances T cell persistence and function in adoptive immunotherapy models Yuki Kagoya,, Cheryl H. Arrowsmith, Naoto Hirano J Clin Invest. 2016;126(9):3479-3494. https://doi.org/10.1172/jci86437.
Engineered T cells: Next-generation cancer immunotherapy
 Engineered T cells: Next-generation cancer immunotherapy Rahul Purwar, Assistant Professor Department of Biosciences & Bioengineering Indian Institute of Technology Bombay (IIT Bombay) Mumbai, India, 400076
Engineered T cells: Next-generation cancer immunotherapy Rahul Purwar, Assistant Professor Department of Biosciences & Bioengineering Indian Institute of Technology Bombay (IIT Bombay) Mumbai, India, 400076
Supplemental Methods. CD107a assay
 Supplemental Methods CD107a assay For each T cell culture that was tested, two tubes were prepared. One tube contained BCMA-K562 cells, and the other tube contained NGFR-K562 cells. Both tubes contained
Supplemental Methods CD107a assay For each T cell culture that was tested, two tubes were prepared. One tube contained BCMA-K562 cells, and the other tube contained NGFR-K562 cells. Both tubes contained
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Tbk1-TKO! DN cells (%)! 15! 10!
 a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
a! T Cells! TKO! B Cells! TKO! b! CD4! 8.9 85.2 3.4 2.88 CD8! Tbk1-TKO! 1.1 84.8 2.51 2.54 c! DN cells (%)! 4 3 2 1 DP cells (%)! 9 8 7 6 CD4 + SP cells (%)! 5 4 3 2 1 5 TKO! TKO! TKO! TKO! 15 1 5 CD8
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation
 Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
