Supplementary references
|
|
|
- Pearl Montgomery
- 6 years ago
- Views:
Transcription
1 Gene expression, pathway and network analyses Gene expression associated with CHI-D was determined using GSE32610 from the Gene Expression Omnibus (GEO: (1). The transcriptomic datasets of CHI and control pancreas [Human 58K oligonucleotide array (GPL14670)] were downloaded into Qlucore Omics Explorer 2.3 (Qlucore, Lund, Sweden), normalized by multi-dimensional scaling (Isomapping) (2) and compared by ANOVA with a significance cut-off of P<0.05. Identification of biological pathways and functions associated with gene expression changes was undertaken using a right-sided Fisher s exact test within Ingenuity Pathway Analysis (IPA) software. Network analysis was also used to identify and prioritize key functional elements. In brief, to derive an interactome model differentially expressed genes were used as seeds and all known protein:protein and protein:gene interactions between the seeds and their inferred immediate neighbours were calculated to generate a biological network using the Biogrid model of the human Interactome ( ) (3). Network generation and processing was performed using Cytoscape Clustering and community structure within interactome models, known to be associated with function (4-6), was prioritized using the ModuLand plug-in for Cytoscape to determine overlapping modules (7) and to identify hierarchical structure (8). Clusters were confirmed using MCODE algorithm (9) and module centrality was used to derive a priority index for genes of biological importance. Supplementary references 1. Michelsen NV, Brusgaard K, Tan Q, Thomassen M, Hussein K, Christesen HT: Investigation of Archived Formalin-Fixed Paraffin-Embedded Pancreatic Tissue with Whole-Genome Gene Expression Microarray. ISRN Pathology 2011;2011:Article ID Tenenbaum JB, de Silva V, Langford JC: A global geometric framework for nonlinear dimensionality reduction. Science 2000;290: Chatr-Aryamontri A, Breitkreutz BJ, Heinicke S, Boucher L, Winter A, Stark C, Nixon J, Ramage L, Kolas N, O'Donnell L, Reguly T, Breitkreutz A, Sellam A, Chen D, Chang C, Rust J, Livstone M, Oughtred R, Dolinski K, Tyers M: The BioGRID interaction database: 2013 update. Nucleic Acids Res 41:D Sun J, Zhao Z: A comparative study of cancer proteins in the human protein-protein interaction network. BMC Genomics 2010;11 Suppl 3:S5 5. Ravasz E, Barabasi AL: Hierarchical organization in complex networks. Phys Rev E Stat Nonlin Soft Matter Phys 2003;67: Yu H, Kim PM, Sprecher E, Trifonov V, Gerstein M: The importance of bottlenecks in protein networks: correlation with gene essentiality and expression dynamics. PLoS Comput Biol 2007;3:e59 7. Szalay-Beko M, Palotai R, Szappanos B, Kovacs IA, Papp B, Csermely P: ModuLand plug-in for Cytoscape: determination of hierarchical layers of overlapping network modules and community centrality. Bioinformatics 28: Kovacs IA, Palotai R, Szalay MS, Csermely P: Community landscapes: an integrative approach to determine overlapping network module hierarchy, identify key nodes and predict network dynamics. PLoS One 2010;5 9. Bader GD, Hogue CW: An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 2003;4:2 10. Piper K, Brickwood S, Turnpenny LW, Cameron IT, Ball SG, Wilson DI, Hanley NA: Beta cell differentiation during early human pancreas development. J Endocrinol 2004;181:11-23
2 Supplementary Table 1. CHI-D patient information. Ten cases undergoing pancreatectomy for sustained hypoglycemia unresponsive to medical treatment. Mode of inheritance for gene defects; AR=autosomal recessive, AD=autosomal dominant. Details of mutations; p.? = intronic mutation resulting in unknown protein size. Patient Age at Surgery Sex Birth Weight (kg) Gene Defect Inheritance AD/AR Paternal Mutation CHI-D1 2m AR p.? M 6.7 ABCC8 (c g>a) CHI-D2 2m M AR p.q299r 4.3 KCNJ11 (c.896a>g) CHI-D3 M 4.4 ABCC8 AR p.a30v 2m (c.89c>t) CHI-D4 F 3.5 ABCC8 AR p.s581t 2m (c.1741t>a) CHI-D5 3m M 4.5 AR p.? ABCC8 (c.1818-?_1923+?del) CHI-D6 4m M 2.9 ABCC8 AR p.h36r (c.107a>g) CHI-D7 5m M 3.5 AR p.l610p ABCC8 (C.1829T>C) CHI-D8 5m M 1.9 ABCC8 AD p.i1512t (c.4535t>c) CHI-D9 6m M AR p.? 2.9 ABCC8 (c.148+1g>a) CHI-D10 13m AR p.? F 4.6 ABCC8 (c g>t) Maternal Mutation p.? (c g>a) p.q299r (c.896a>g) p.a30v (c.89c>t) p.? (c g>a) p.t172fs c.512dup p.? (c g>t) p.l610p (C.1829T>C) - p.? (c.148+1g>a) p.a4v (c.11c>t)
3 Supplementary Table 2. Primary antibodies The antibodies developed by O.D. Madsen (anti-nkx6.1) and T.M. Jessell (anti-nkx2.2 and anti-isl1) were obtained from the Developmental Studies Hybridoma Bank developed under the auspices of the NICHD and maintained by the University of Iowa, Department of Biology, Iowa City, IA Primary Antibody Raised In Dilution Supplier Polyclonal anti-insulin Rabbit 1:1000 Abcam Polyclonal anti-insulin Guinea Pig 1:100 Zymed Polyclonal anti-glucagon Rabbit 1:50 Zymed Polyclonal anti-somatostatin Rabbit 1:50 Zymed Polyclonal anti-pp Rabbit 1:50 Zymed Polyclonal anti-ghrelin Goat 1:500 Abcam Polyclonal anti-gastrin Rabbit 1:200 Cell Marque Monoclonal anti-nkx6.1 Mouse 1:1000 DSHB Monoclonal anti-nkx2.2 Mouse 1:75 DSHB Monoclonal anti-ki67 Mouse 1:100 Novocastra Monoclonal anti-phh3 Mouse 1:200 Polyclonal anti-phospho- -H2AX Rabbit 1:200 Cell Signaling Cell Signaling Polyclonal anti-sox9 Rabbit 1:5000 Millipore Polyclonal anti-gata4 Goat 1:450 Abcam Polyclonal anti-cdk6 Rabbit 1:500 Abcam Polyclonal anti-ck19 Mouse 1:100 Novocastra Polyclonal anti- P27Kip1 Rabbit 1:200 Santa Cruz Polyclonal anti-prb Rabbit 1:500 Abcam Polyclonal anti-foxa2 Goat 1:800 R & D Polyclonal anti-pdx1 Guinea pig 1:500 Abcam Monoclonal anti-isl1 Mouse 1:200 DSHB
4 Supplementary Table 3. qrt-pcr primers Forward and reverse primer sequences are given for each gene along with the accession number and product size. All primer pairs / products were intron-spanning wherever possible. Gene Accession No. Forward Primer Reverse Primer Size (bp) -ACTIN NM_ CCAACCGCGAGAAGATGA CCAGAGGCGTACAGGGATAG 138 HPRT NM_ TGACCTTGATTTATTTTGCATACC CGAGCAAGACGTTCAGTCCT 102 PDX1 NM_ AAAGGCCAGTGGGCAGGCGG GCGCGGCCGTGAGATGTACT 135 FOXA2 NM_ GAAGATGGAAGGGCACGAGC GTACGTGTTCATGCCGTTCA 115 SOX9 NM_ GTACCCGCACTTGCACAAC TCGCTCTCGTTCAGAAGTCTC 72 NKX6.1 NM_ GGCCTGTACCCCTCATCAAG TCCGGAAAAAGTGGGTCTCG 79 NEUROD1 NM_ GAGGCCCCAGGGTTATGAGA TGGTCATGTTTCGATTTCCTTTGT 70 MAFA NM_ AGAGCGAGAAGTGCCAACTC GTACAGGTCCCGCTCTTTGG 83 NKX2.2 NM_ ATGTCGCTGACCAACACAAAG GATGTCCTTGACCGAAAACCC 45 INSULIN NM_ TGGCTTCTTCTACACACCCA TCTAGTTGCAGTAGTTCTCCA 984 PAX4 NM_ AAGAAAGCAGCTTGCGTTGAC GGGCGTGAGACAGAGGTATC 165 PAX6 NM_ CCCGGCAGAAGATTGTAGAG GCTAGCCAGGTTGCGAAGAA 323 ARX NM_ ATTCGGCAGGCTCTTTTCCA ATGTTGGAGTTGGAGCGAGG 502 MAFB NM_ TGAACTTTGCGCGTTAAGCC CACGCAGCCGCCGGAGTTTC 436 CDKN1B NM_ TCCGGCTAACTCTGAGGACA GAAGAATCGTCGGTTGCAGG 120
5 Supplementary Table 4. Top 20 functional modules following network analysis ranked by the index of centrality of their most central gene The most central gene in the top 20 modules is shown in bold. In parentheses are the other genes in the top ten for each module. Module Most central gene in the module (others genes in the top 10) rank 1 EP300 (UBD, SIRT7, KAT2B, TP53, FN1, PRMT1, ARRB1, VCAM1, TRIM28) 2 PML (UBD, DAXX, SUMO2, TRIM28, FN1, SUMO1, PIAS1, ARRB1, VCAM1) 3 CDKN1B (CDK2, CDK7, CDK4, SKP2, CCT8, UBD, CCNE1, COPS5, VCAM1) 4 NCOA2 (ESR1, EP300, AR, GNAI2, PRMT1, FN1, SIRT7, SAFB, ACTR2) 5 CAND1 (CUL1, COPS3, FN1, CUL3, PHKG2, CUL4A, RNF7, SIRT7, UBD) 6 PIK3R1 (CRK, FYN, CBL, VAV1, ITSN1, ERBB3, GRB2, EGFR, BCAR1) 7 TBP (TAF9, TAF6, TAF10, TAF5, TCEA1, TAF1, TAF4, ELAVL1, GTF2B) 8 HDAC5 (AR, UBD, TBL1XR1, NCOR1, PHKG2, ZBTB16, ARRB1, HNF4A, HSP90AA1) 9 CDK7 (TCEA1, CCNH, GTF2H2, POLR2A, SIRT7, ERCC3, MNAT1, GTF2H1, APP) 10 COPS3 (COPS5, COPS6, ERCC8, TK1, SIRT7, CUL3, DDB1, CUL4B, TOR1AIP2) 11 MRE11A (SIRT7, RAD50, NBN, DDX1, TERF1, BRCA1, NRF1, HNRNPD, VCAM1) 12 ZBTB16 (HDAC1, HDAC5, TRIM28, FN1, PHB, DNMT3B, TK1, EHMT2, VCAM1) 13 SMURF1 (SMAD1, SMAD7, SMAD5, STRAP, NEDD4, APP, DCTN2, PHKG2, UBE2D3) 14 FN1 (SIRT7, HNRNPD, VCAM1, GAPDH, VHL, TK1, PHKG2, CCT8, ELAVL1) 15 BAG1 (HSPA8, HSPA4, TTC1, ARRB1, NRF1, PHKG2, STUB1, TERF1, ACTR2) 16 NEDD4 (UBE4B, UBE2D2, UBE2D3, UBE2L3, LAPTM5, MKRN3, MGRN1, ARRB1, UBE2D1) 17 UBE2M (APP, NEDD8, PDIA3, NRF1, RBX1, CLU, NDUFS6, DCUN1D1, UBA3) 18 PSMA6 (PSMA5, UBD, VCAM1, PHKG2, FKBP8, FN1, PSMA2, PSMA3, STK4) 19 HEXIM1 (BRD4, CDK9, CCNT1, EAF1, TERF1, RN7SK, NRF1MED12, STK4) 20 RING1 (BMI1, PHC1, RNF2, PCGF2, PHC2, INTS6, CBX4, TERF1, APP)
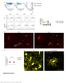
6 Supplementary Figure 1. -cells, -cells and -cells in CHI-D and control pancreas during the early postnatal period Consecutive 5 µm sections from three examples of CHI-D and age-matched control pancreas are stained for insulin ( -cells), glucagon ( -cells) and somatostatin ( -cells) (all brown) counter-stained with toluidine blue. (A- B) 2-3 months (mo); (C-D) 5-6 months; and (E-F) months. Arrowheads in each glucagon panel point to an islet at each age that is also apparent in sections either side stained for insulin and somatostatin. At 2-3 months, note the extra-islet, scattered hormone-positive cells in both CHI-D and controls. At all ages, note that glucagonpositive cells are more evenly distributed throughout the islets in CHI-D compared to the peripheral -cells in control islets. The consistently central insulin staining and, in older CHI-D samples, more peripheral somatostatin in adjacent sections exemplifies that this feature was not due to the plane of sectioning (also see Fig. 1 in main text). Gastrin was not observed in CHI-D or age-matched control pancreas compared to a positive control of human fetal intestine at 10 weeks post-conception (image below). Scale bar represents 200 µm.
7 Size bar = 100 m
8 Supplementary Figure 2. Endocrine lineages and islet composition in CHI-D and age-matched controls Dual confocal immunofluorescence of consecutive sections from CHI-D and age-matched control pancreas counterstained with DAPI. (A-B), 2-3 months (mo); (C-D), 5-6 months; and (E-F), months. Insets demonstrate PP staining elsewhere within the same section. At early stages α-cells and δ-cells are more diffusely scattered throughout islets rather than arranged as a mantle. Ghrelin staining was located peripherally along with occasional PP-cells in both CHI-D and control Islet structure remains less organized and compact in CHI-D. Scale bars represent 100 µm.
9 Supplementary Figure 3. Localization of insulin and glucagon within the same endocrine cells in CHI-D Dual confocal immunofluorescence for insulin and glucagon counterstained with DAPI in tissue sections from two cases of CHI-D (A-B) and human fetal pancreas (C) at 10 weeks post-conception (wpc). In CHI-D the arrowheads point to cells which contain both insulin and glucagon but where each hormone localizes to a discrete area of the cell. In contrast, detection of the two hormones completely overlaps in human fetal pancreas (10). The associated supplementary video confirms the cellular co-detection by moving through a cell on confocal Z-stack. Scale bar represents 50 µm.
10 Supplementary Figure 4. SOX9, FOXA2, NKX6.1 and GATA4 in CHI-D and control pancreas Brightfield immunohistochemistry counterstained with toluidine blue shows SOX9 (duct cells, A-B), FOXA2 (duct cells and -cells, C-D), NKX6.1 ( -cells, E-F) and GATA4 (acinar cells, G-H) in an example of CHI-D pancreas and its aged-matched control. The staining profile was identical in all CHI-D samples (2-13 months). Note examples of nucleomegaly amongst the FOXA2 and NKX6.1 staining in CHI-D but not CHI-D duct or acinar cells or any of the control cells. NKX6.1 also illustrates the diffuse nature of β-cells in CHI-D compared to control pancreas. Note also the increased GATA4 staining in CHI-D compared to control consistent with the major acinar cell proliferation observed in CHI-D (Fig. 6C, main text). Along with satisfactory tissue morphology, the sensitive detection of nuclear transcription factors argues against significant pancreatic autolysis. Scale bars represent 200 µm.
11 Supplementary Figure 5. Nucleomegaly in CHI-D affects δ-cells as well as β-cells (A) Nuclear diameter of islet cells in CHI-D samples compared to age-matched controls. CHI-D cells had larger nuclei on average with a subset having especially large nuclei beyond a normal distribution (nucleomegaly; marked on graph). Diameter was measured using digitization data obtained from histological slides using the HistoQuant and Pannoramic Viewer software. (B) Brightfield immunohistochemistry for transcription factors (stained brown) in CHI-D pancreas demonstrating examples of nucleomegaly (arrowheads). Not all nucleomegalic cells stained for PDX1 (e.g. white arrowhead). Size bar represents 20 μm. (C) Immunofluorescence counterstained with DAPI. Arrows demonstrate examples of nucleomegalic β-cells (stained for insulin) and δ-cells (stained for somatostatin). Scale bar represents 10 μm.
12 Supplementary Figure 6. Relative detection of PHH3 in CHI-D pancreas. (A) PHH3 was predominantly detected in acinar cells (arrows), rather than in ducts (asterisk) or islets (red hatched circles). (B) Higher magnification example to show PHH3 detection within an islet. (C) Summary data of PHH3- positive cells in two cases of CHI-D. Approximately 90% of PHH3-positive cells were acinar, 7% were islet and 3% were duct (n=162).
13 Supplementary Figure 7. Clusters of genes identified as functional modules within a network model of CHI (A) Interactome model derived from the BioGRID database of significantly altered gene expression in CHI tissue (1). Cluster modules (coloured) were identified with the interactome model using the Moduland algorithm. (B) Network of the cluster modules showing the most central gene (top 20 ranking shown in Supplementary Table 4).
14 Supplementary Figure 8. Correlation between cell-cycle markers and -cell nucleomegaly (A) Ki67-positive nuclei in CHI-D are slightly larger. Frequency histograms showing the range of nuclear diameters recorded in Ki67-positive cells from each of the five cases of CHI-D (blue curves) and the mean diameter from Ki67-negative cells from the same cases (black curve). Comparing nuclear diameter in Ki67- positive and Ki67-negative populations was significant at P<0.01. (B) Example of nuclear Ki67 staining (arrow) showing not all nucleomegalic cells are proliferative (arrowhead). (C) Dual immunofluorescence for CDK6, retinoblastoma protein (prb) and P27 proteins with insulin counterstained with DAPI showing not all nucleomegalic cells are positively stained. Scale bar represents 20 m. Supplementary Video. Insulin and glucagon colocalization in CHI-D Animation captured at x630 magnification moving through a series of cross-sectional z-stack images for a CHI-D cell demonstrating codetection of insulin (green) and glucagon (red).
SUPPLEMENTARY DATA. Supplementary Table 2. Antibodies used for Immunofluoresence. Supplementary Table 3. Real-time PCR primer sequences.
 Supplementary Table 2. Antibodies used for Immunofluoresence. Antibody Dilution Source Goat anti-pdx1 1:100 R&D Systems Rabbit anti-hnf6 1:100 Santa Cruz Biotechnology Mouse anti-nkx6.1 1:200 Developmental
Supplementary Table 2. Antibodies used for Immunofluoresence. Antibody Dilution Source Goat anti-pdx1 1:100 R&D Systems Rabbit anti-hnf6 1:100 Santa Cruz Biotechnology Mouse anti-nkx6.1 1:200 Developmental
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
islets scored 1 week month months
 Supplementary Table 1. Sampling parameters for the morphometrical analyses Time (post- DT) Control mice (age-matched) α-cells mice pancreatic surface (mm 2 ) scored DT-treated mice islets scored mice pancreatic
Supplementary Table 1. Sampling parameters for the morphometrical analyses Time (post- DT) Control mice (age-matched) α-cells mice pancreatic surface (mm 2 ) scored DT-treated mice islets scored mice pancreatic
Impact of Sox9 Dosage and Hes1-mediated Notch Signaling in Controlling the Plasticity of Adult Pancreatic Duct Cells in Mice
 Impact of Sox9 Dosage and Hes1-mediated Notch Signaling in Controlling the Plasticity of Adult Pancreatic Duct Cells in Mice Shinichi Hosokawa 1,3,Kenichiro Furuyama 1,3, Masashi Horiguchi 1,3,Yoshiki
Impact of Sox9 Dosage and Hes1-mediated Notch Signaling in Controlling the Plasticity of Adult Pancreatic Duct Cells in Mice Shinichi Hosokawa 1,3,Kenichiro Furuyama 1,3, Masashi Horiguchi 1,3,Yoshiki
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1: Neuregulin 1 increases the growth of mammary organoids compared to EGF. (a) Mammary epithelial cells were freshly isolated,
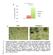 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1: Neuregulin 1 increases the growth of mammary organoids compared to EGF. (a) Mammary epithelial cells were freshly isolated, embedded in matrigel and exposed
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1: Neuregulin 1 increases the growth of mammary organoids compared to EGF. (a) Mammary epithelial cells were freshly isolated, embedded in matrigel and exposed
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment
 Supplementary Information Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment Robin A. Kimmel, Stefan Dobler, Nicole Schmitner, Tanja Walsen, Julia
Supplementary Information Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment Robin A. Kimmel, Stefan Dobler, Nicole Schmitner, Tanja Walsen, Julia
Supplementary Table 1. Characterization of HNSCC PDX models established at MSKCC
 Supplementary Table 1. Characterization of HNSCC PDX models established at MSKCC Supplementary Table 2. Drug content and loading efficiency estimated with F-NMR and UV- Vis Supplementary Table 3. Complete
Supplementary Table 1. Characterization of HNSCC PDX models established at MSKCC Supplementary Table 2. Drug content and loading efficiency estimated with F-NMR and UV- Vis Supplementary Table 3. Complete
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 )
 770 771 Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 ) Human CXCL1 GCGCCCAAACCGAAGTCATA ATGGGGGATGCAGGATTGAG PF4 CCCCACTGCCCAACTGATAG TTCTTGTACAGCGGGGCTTG
770 771 Supplementary Table 1. The primers used for quantitative RT-PCR. Gene name Forward (5 > 3 ) Reverse (5 > 3 ) Human CXCL1 GCGCCCAAACCGAAGTCATA ATGGGGGATGCAGGATTGAG PF4 CCCCACTGCCCAACTGATAG TTCTTGTACAGCGGGGCTTG
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Figure legends Supplemental Fig.1. Glucose-induced insulin secretion and insulin content of islets. Supplemental Fig. 2.
 Figure legends Supplemental Fig.. Glucose-induced insulin secretion and insulin content of islets. Insulin secretory responses to.,., and. mm glucose (A) (n = 7-), and the insulin content in the islets
Figure legends Supplemental Fig.. Glucose-induced insulin secretion and insulin content of islets. Insulin secretory responses to.,., and. mm glucose (A) (n = 7-), and the insulin content in the islets
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Inhibition of Cdk5 Promotes β-cell Differentiation from Ductal Progenitors
 Inhibition of Cdk5 Promotes β-cell Differentiation from Ductal Progenitors Ka-Cheuk Liu, Gunter Leuckx, Daisuke Sakano, Philip A. Seymour, Charlotte L. Mattsson, Linn Rautio, Willem Staels, Yannick Verdonck,
Inhibition of Cdk5 Promotes β-cell Differentiation from Ductal Progenitors Ka-Cheuk Liu, Gunter Leuckx, Daisuke Sakano, Philip A. Seymour, Charlotte L. Mattsson, Linn Rautio, Willem Staels, Yannick Verdonck,
Supplementary Figure 1
 Supplementary Figure 1 Control Pancreatitis Supplementary Figure 2 A Panc Liver SI Spleen H 2 O B EZH2 fl/fl C EZH2 fl/fl 37bp EZH2 ERK2 D E 5 EZH2 fl/fl Fasting Glucose (mg/dl) 2 18 16 14 12 1 8 6 4 2
Supplementary Figure 1 Control Pancreatitis Supplementary Figure 2 A Panc Liver SI Spleen H 2 O B EZH2 fl/fl C EZH2 fl/fl 37bp EZH2 ERK2 D E 5 EZH2 fl/fl Fasting Glucose (mg/dl) 2 18 16 14 12 1 8 6 4 2
Supplemental Information. Otic Mesenchyme Cells Regulate. Spiral Ganglion Axon Fasciculation. through a Pou3f4/EphA4 Signaling Pathway
 Neuron, Volume 73 Supplemental Information Otic Mesenchyme Cells Regulate Spiral Ganglion Axon Fasciculation through a Pou3f4/EphA4 Signaling Pathway Thomas M. Coate, Steven Raft, Xiumei Zhao, Aimee K.
Neuron, Volume 73 Supplemental Information Otic Mesenchyme Cells Regulate Spiral Ganglion Axon Fasciculation through a Pou3f4/EphA4 Signaling Pathway Thomas M. Coate, Steven Raft, Xiumei Zhao, Aimee K.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Advances in pancreatic islet monolayer culture on glass surfaces enable superresolution microscopy and insights into beta cell ciliogenesis and proliferation Edward A. Phelps,
SUPPLEMENTARY INFORMATION Advances in pancreatic islet monolayer culture on glass surfaces enable superresolution microscopy and insights into beta cell ciliogenesis and proliferation Edward A. Phelps,
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Hippocampal sections from new-born Pten+/+ and PtenFV/FV pups were stained with haematoxylin and eosin (H&E) and were imaged at (a) low and (b) high
marker. DAPI labels nuclei. Flies were 20 days old. Scale bar is 5 µm. Ctrl is
 Supplementary Figure 1. (a) Nos is detected in glial cells in both control and GFAP R79H transgenic flies (arrows), but not in deletion mutant Nos Δ15 animals. Repo is a glial cell marker. DAPI labels
Supplementary Figure 1. (a) Nos is detected in glial cells in both control and GFAP R79H transgenic flies (arrows), but not in deletion mutant Nos Δ15 animals. Repo is a glial cell marker. DAPI labels
NEW IHC A n t i b o d i e s
 NEW IHC Antibodies TABLE OF CONTENTS NEW IHC ANTIBODIES from Cell Marque CITED1 (5H6).... 1 Claudin 7 (5D10F3).... 1 GATA1 (4F5).... 1 Transgelin (2A10C2).... 1 NEW IHC ANTIBODIES using RabMAb Technology
NEW IHC Antibodies TABLE OF CONTENTS NEW IHC ANTIBODIES from Cell Marque CITED1 (5H6).... 1 Claudin 7 (5D10F3).... 1 GATA1 (4F5).... 1 Transgelin (2A10C2).... 1 NEW IHC ANTIBODIES using RabMAb Technology
Supplementary Figure 1: Validation of labeling specificity of immature OSNs and presynaptic terminals. (A) (B) (C) (D) (E)
 Supplementary Figure 1: Validation of labeling specificity of immature OSNs and presynaptic terminals. (A) Confocal images of septal olfactory epithelium of an adult Gγ8-sypGFP-tdTom mouse showing colocalization
Supplementary Figure 1: Validation of labeling specificity of immature OSNs and presynaptic terminals. (A) Confocal images of septal olfactory epithelium of an adult Gγ8-sypGFP-tdTom mouse showing colocalization
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Fig. S4. Current-voltage relations of iglurs. A-C: time courses of currents evoked by 100 ms pulses
 Fig. S1. Immunohistochemical detection of iglur2 protein in single islet cells. A: α cells identified using glucagon-specific antibody express the iglur2 subtype of AMPA receptor. 24 out of 26 identified
Fig. S1. Immunohistochemical detection of iglur2 protein in single islet cells. A: α cells identified using glucagon-specific antibody express the iglur2 subtype of AMPA receptor. 24 out of 26 identified
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Figure (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were
 Supplementary Figure 1. Gd@C 82 (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were treated with PBS, Gd@C 82 (OH) 22, C 60 (OH) 22 or GdCl
Supplementary Figure 1. Gd@C 82 (OH) 22 nanoparticles did not affect cell viability and apoposis. MDA-MB-231, MCF-7, MCF-10A and BT549 cells were treated with PBS, Gd@C 82 (OH) 22, C 60 (OH) 22 or GdCl
Supplementary Figure 1.
 Supplementary Figure 1. Increased β cell mass and islet diameter in βtsc2 -/- mice up to 35 weeks A: Reconstruction of multiple anti-insulin immunofluorescence images showing differences in β cell mass
Supplementary Figure 1. Increased β cell mass and islet diameter in βtsc2 -/- mice up to 35 weeks A: Reconstruction of multiple anti-insulin immunofluorescence images showing differences in β cell mass
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2610 Figure S1 FSMCs derived from MSLN CLN transgenic mice express smooth muscle-specific proteins. Beta-galactosidase is ubiquitously expressed within cultured FSMCs derived from MSLN
DOI: 10.1038/ncb2610 Figure S1 FSMCs derived from MSLN CLN transgenic mice express smooth muscle-specific proteins. Beta-galactosidase is ubiquitously expressed within cultured FSMCs derived from MSLN
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Materials. for Garmy-Susini, et al, Integrin 4 1 signaling is required for lymphangiogenesis and tumor metastasis
 Supplementary Materials for Garmy-Susini, et al, Integrin 4 1 signaling is required for lymphangiogenesis and tumor metastasis 1 Supplementary Figure Legends Supplementary Figure 1: Integrin expression
Supplementary Materials for Garmy-Susini, et al, Integrin 4 1 signaling is required for lymphangiogenesis and tumor metastasis 1 Supplementary Figure Legends Supplementary Figure 1: Integrin expression
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h)
 Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Supplemental Information. 3D-CLEM Reveals that a Major Portion. of Mitotic Chromosomes Is Not Chromatin
 Molecular Cell, Volume 64 Supplemental Information 3D-CLEM Reveals that a Major Portion of Mitotic Chromosomes Is Not Chromatin Daniel G. Booth, Alison J. Beckett, Oscar Molina, Itaru Samejima, Hiroshi
Molecular Cell, Volume 64 Supplemental Information 3D-CLEM Reveals that a Major Portion of Mitotic Chromosomes Is Not Chromatin Daniel G. Booth, Alison J. Beckett, Oscar Molina, Itaru Samejima, Hiroshi
Report on Pathology. Study: The effect of Compound X on pancreatic islets in rhesus macaques
 Report on Pathology Study: The effect of Compound X on pancreatic islets in rhesus macaques Prepared for: Client Name Client Address January 1, 2013 Prepared by: Charter Preclinical Services 21 Main St.,
Report on Pathology Study: The effect of Compound X on pancreatic islets in rhesus macaques Prepared for: Client Name Client Address January 1, 2013 Prepared by: Charter Preclinical Services 21 Main St.,
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Supplemental Information. Tissue Myeloid Progenitors Differentiate. into Pericytes through TGF-b Signaling. in Developing Skin Vasculature
 Cell Reports, Volume 18 Supplemental Information Tissue Myeloid Progenitors Differentiate into Pericytes through TGF-b Signaling in Developing Skin Vasculature Tomoko Yamazaki, Ani Nalbandian, Yutaka Uchida,
Cell Reports, Volume 18 Supplemental Information Tissue Myeloid Progenitors Differentiate into Pericytes through TGF-b Signaling in Developing Skin Vasculature Tomoko Yamazaki, Ani Nalbandian, Yutaka Uchida,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
15. Supplementary Figure 9. Predicted gene module expression changes at 24hpi during HIV
 Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
Supplemental Information. Induction of Expansion and Folding. in Human Cerebral Organoids
 Cell Stem Cell, Volume 20 Supplemental Information Induction of Expansion and Folding in Human Cerebral Organoids Yun Li, Julien Muffat, Attya Omer, Irene Bosch, Madeline A. Lancaster, Mriganka Sur, Lee
Cell Stem Cell, Volume 20 Supplemental Information Induction of Expansion and Folding in Human Cerebral Organoids Yun Li, Julien Muffat, Attya Omer, Irene Bosch, Madeline A. Lancaster, Mriganka Sur, Lee
Suppl Video: Tumor cells (green) and monocytes (white) are seeded on a confluent endothelial
 Supplementary Information Häuselmann et al. Monocyte induction of E-selectin-mediated endothelial activation releases VE-cadherin junctions to promote tumor cell extravasation in the metastasis cascade
Supplementary Information Häuselmann et al. Monocyte induction of E-selectin-mediated endothelial activation releases VE-cadherin junctions to promote tumor cell extravasation in the metastasis cascade
Supplemental Figure 1
 Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
MANUSCRIPT TITLE: Protein kinase C δ signaling is required for dietary prebiotic-induced strengthening of intestinal epithelial barrier function
 MANUSCRIPT TITLE: Protein kinase C δ signaling is required for dietary prebiotic-induced strengthening of intestinal epithelial barrier function Authors: Richard Y. Wu 1,2, Majd Abdullah 1, Pekka Määttänen
MANUSCRIPT TITLE: Protein kinase C δ signaling is required for dietary prebiotic-induced strengthening of intestinal epithelial barrier function Authors: Richard Y. Wu 1,2, Majd Abdullah 1, Pekka Määttänen
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature11463 %Sox17(+) 9 8 7 6 5 4 3 2 1 %Sox17(+) #Sox17(+) d2 d4 d6 d8 d1 d12 d14 d18 25 2 15 1 5 Number of Sox17(+) cells X 1 Supplementary Figure 1: Expression of
SUPPLEMENTARY INFORMATION doi:1.138/nature11463 %Sox17(+) 9 8 7 6 5 4 3 2 1 %Sox17(+) #Sox17(+) d2 d4 d6 d8 d1 d12 d14 d18 25 2 15 1 5 Number of Sox17(+) cells X 1 Supplementary Figure 1: Expression of
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Endocannabinoid-activated Nlrp3 inflammasome in infiltrating macrophages mediates β- cell loss in type 2 diabetes
 Endocannabinoid-activated Nlrp3 inflammasome in infiltrating macrophages mediates β- cell loss in type 2 diabetes T Jourdan, G Godlewski, R Cinar, A Bertola, G Szanda, J Liu, J Tam, T Han, B Mukhopadhyay,
Endocannabinoid-activated Nlrp3 inflammasome in infiltrating macrophages mediates β- cell loss in type 2 diabetes T Jourdan, G Godlewski, R Cinar, A Bertola, G Szanda, J Liu, J Tam, T Han, B Mukhopadhyay,
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation
 Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
The window period of NEUROGENIN3 during human gestation
 The window period of NEUROGENIN3 during human gestation Rachel J Salisbury, Jennifer Blaylock, Andrew A Berry, Rachel E Jennings, Ronald De Krijger, Karen Piper Hanley, and Neil A Hanley QUERY SHEET This
The window period of NEUROGENIN3 during human gestation Rachel J Salisbury, Jennifer Blaylock, Andrew A Berry, Rachel E Jennings, Ronald De Krijger, Karen Piper Hanley, and Neil A Hanley QUERY SHEET This
Preferential reduction of β cells derived from Pax6 MafB pathway in MafB deficient mice
 Available online at www.sciencedirect.com Developmental Biology 314 (2008) 443 456 www.elsevier.com/developmentalbiology Genomes & Developmental Control Preferential reduction of β cells derived from Pax6
Available online at www.sciencedirect.com Developmental Biology 314 (2008) 443 456 www.elsevier.com/developmentalbiology Genomes & Developmental Control Preferential reduction of β cells derived from Pax6
GATA4 and GATA6 control mouse pancreas organogenesis
 GATA4 and GATA6 control mouse pancreas organogenesis Manuel Carrasco,, Francisco Martín, Anabel Rojas J Clin Invest. 2012;122(10):3504-3515. https://doi.org/10.1172/jci63240. Research Article Development
GATA4 and GATA6 control mouse pancreas organogenesis Manuel Carrasco,, Francisco Martín, Anabel Rojas J Clin Invest. 2012;122(10):3504-3515. https://doi.org/10.1172/jci63240. Research Article Development
Supplementary Figure 1. Identification of tumorous sphere-forming CSCs and CAF feeder cells. The LEAP (Laser-Enabled Analysis and Processing)
 Supplementary Figure 1. Identification of tumorous sphere-forming CSCs and CAF feeder cells. The LEAP (Laser-Enabled Analysis and Processing) platform with laser manipulation to efficiently purify lung
Supplementary Figure 1. Identification of tumorous sphere-forming CSCs and CAF feeder cells. The LEAP (Laser-Enabled Analysis and Processing) platform with laser manipulation to efficiently purify lung
Immunostaining was performed on tumor biopsy samples arranged in a tissue-microarray format or on
 Supplemental Methods Immunohistochemical Analyses Immunostaining was performed on tumor biopsy samples arranged in a tissue-microarray format or on prostatectomy sections obtained post-study. Briefly,
Supplemental Methods Immunohistochemical Analyses Immunostaining was performed on tumor biopsy samples arranged in a tissue-microarray format or on prostatectomy sections obtained post-study. Briefly,
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary Methods: IGFBP7 Drives Resistance to Epidermal Growth Factor Receptor Tyrosine Kinase Inhibition in Lung Cancer
 S1 of S6 Supplementary Methods: IGFBP7 Drives Resistance to Epidermal Growth Factor Receptor Tyrosine Kinase Inhibition in Lung Cancer Shang-Gin Wu, Tzu-Hua Chang, Meng-Feng Tsai, Yi-Nan Liu, Chia-Lang
S1 of S6 Supplementary Methods: IGFBP7 Drives Resistance to Epidermal Growth Factor Receptor Tyrosine Kinase Inhibition in Lung Cancer Shang-Gin Wu, Tzu-Hua Chang, Meng-Feng Tsai, Yi-Nan Liu, Chia-Lang
Supplementary Information
 Supplementary Information Title Degeneration and impaired regeneration of gray matter oligodendrocytes in amyotrophic lateral sclerosis Authors Shin H. Kang, Ying Li, Masahiro Fukaya, Ileana Lorenzini,
Supplementary Information Title Degeneration and impaired regeneration of gray matter oligodendrocytes in amyotrophic lateral sclerosis Authors Shin H. Kang, Ying Li, Masahiro Fukaya, Ileana Lorenzini,
Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III
 Supplementary Materials: Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III patient samples. Genomic DNA samples extracted from punch biopsies from either FFPE or frozen tumor
Supplementary Materials: Supplementary Figure 1. IDH1 and IDH2 mutation site sequences on WHO grade III patient samples. Genomic DNA samples extracted from punch biopsies from either FFPE or frozen tumor
Supplementary Table 1. Primer sequences for conventional RT-PCR on mouse islets
 Supplementary Table 1. Primer sequences for conventional RT-PCR on mouse islets Gene 5 Forward 3 5 Reverse 3.T. Product (bp) ( C) mnox1 GTTCTTGGGCTGCCTTGG GCTGGGGCGGCGG 60 300 mnoxa1 GCTTTGCCGCGTGC GGTTCGGGTCCTTTGTGC
Supplementary Table 1. Primer sequences for conventional RT-PCR on mouse islets Gene 5 Forward 3 5 Reverse 3.T. Product (bp) ( C) mnox1 GTTCTTGGGCTGCCTTGG GCTGGGGCGGCGG 60 300 mnoxa1 GCTTTGCCGCGTGC GGTTCGGGTCCTTTGTGC
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Nanoneedle-Mediated Stimulation of Cell Mechanotransduction Machinery Catherine S Hansel, Spencer W Crowder, Samuel Cooper, Sahana Gopal, Maria João Pardelha da Cruz, Leonardo
SUPPLEMENTARY INFORMATION Nanoneedle-Mediated Stimulation of Cell Mechanotransduction Machinery Catherine S Hansel, Spencer W Crowder, Samuel Cooper, Sahana Gopal, Maria João Pardelha da Cruz, Leonardo
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the
 Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
Supplementary Figure 1. HOPX is hypermethylated in NPC. (a) Methylation levels of HOPX in Normal (n = 24) and NPC (n = 24) tissues from the genome-wide methylation microarray data. Mean ± s.d.; Student
stomach p63/krt5 CDH17 CEACAM5 CEACAM7 ANXA13 ANXA10 CLDN18 VSIG1 TFF2 VIL1 GDA
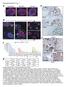 Supplementary Fig. 1 a esophagus Barrett s e FABP1 Ecad stomach TFF3 b p63/krt5 CDH17 REG4 CDH17 f c signal intensity p63/krt5 EsoSC 5000 4000 3000 2000 CDH17 1500 1000 Barrett s KRT14 KRT6A KRT13 KRT5
Supplementary Fig. 1 a esophagus Barrett s e FABP1 Ecad stomach TFF3 b p63/krt5 CDH17 REG4 CDH17 f c signal intensity p63/krt5 EsoSC 5000 4000 3000 2000 CDH17 1500 1000 Barrett s KRT14 KRT6A KRT13 KRT5
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3200 Supplementary Figure 1 Expression analysis of stomach markers in gutlike structure. (a) Differentiation scheme of gut-like structure formation from embryonic stem cells. (b) RT-PCR
DOI: 10.1038/ncb3200 Supplementary Figure 1 Expression analysis of stomach markers in gutlike structure. (a) Differentiation scheme of gut-like structure formation from embryonic stem cells. (b) RT-PCR
Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at
 Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at E10.5 were double-stained for TUNEL (red) and PECAM-1 (green).
Supplementary Figure 1. EC-specific Deletion of Snail1 Does Not Affect EC Apoptosis. (a,b) Cryo-sections of WT (a) and Snail1 LOF (b) embryos at E10.5 were double-stained for TUNEL (red) and PECAM-1 (green).
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Erzsebet Kokovay, Susan Goderie, Yue Wang, Steve Lotz, Gang Lin, Yu Sun, Badrinath Roysam, Qin Shen,
 Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.
 Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2638 Figure S1 Morphological characteristics of fetal testes and ovaries from 6.5-20 developmental weeks. Representative images of Hematoxylin and Eosin staining of testes and ovaries over
DOI: 10.1038/ncb2638 Figure S1 Morphological characteristics of fetal testes and ovaries from 6.5-20 developmental weeks. Representative images of Hematoxylin and Eosin staining of testes and ovaries over
Cell Therapy for Diabetes: Generating Functional Islets from Human Exocrine Tissue
 Cell Therapy for Diabetes: Generating Functional Islets from Human Exocrine Tissue Kevin Docherty University of Aberdeen ELRIG Drug Discovery 2016 ACC Liverpool 14 th October 2016 Autoimmune destruction
Cell Therapy for Diabetes: Generating Functional Islets from Human Exocrine Tissue Kevin Docherty University of Aberdeen ELRIG Drug Discovery 2016 ACC Liverpool 14 th October 2016 Autoimmune destruction
Polycomb protein Ezh2 regulates pancreatic β-cell Ink4a/Arf expression and regeneration in streptozotocin-induced diabetes mellitus
 Chen et al. 1 Polycomb protein Ezh2 regulates pancreatic β-cell Ink4a/Arf expression and regeneration in streptozotocin-induced diabetes mellitus Hainan Chen 1, Xueying Gu 1, I-hsin Su 2, Rita Bottino
Chen et al. 1 Polycomb protein Ezh2 regulates pancreatic β-cell Ink4a/Arf expression and regeneration in streptozotocin-induced diabetes mellitus Hainan Chen 1, Xueying Gu 1, I-hsin Su 2, Rita Bottino
Supporting Information
 Supporting Information Guney et al. 1.173/pnas.117218 SI Materials and Methods PCR and Genotyping. Genotyping of CTGF e2coin/e2coin mice was performed by PCR on N isolated from ear punches using the following
Supporting Information Guney et al. 1.173/pnas.117218 SI Materials and Methods PCR and Genotyping. Genotyping of CTGF e2coin/e2coin mice was performed by PCR on N isolated from ear punches using the following
PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65
 SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
Supplemental figure 1. PDGFRα is expressed dominantly by stromal cells surrounding mammary ducts and alveoli. A) IHC staining of PDGFRα in
 Supplemental figure 1. PDGFRα is expressed dominantly by stromal cells surrounding mammary ducts and alveoli. A) IHC staining of PDGFRα in nulliparous (left panel) and InvD6 mouse mammary glands (right
Supplemental figure 1. PDGFRα is expressed dominantly by stromal cells surrounding mammary ducts and alveoli. A) IHC staining of PDGFRα in nulliparous (left panel) and InvD6 mouse mammary glands (right
SUPPLEMENTARY DATA. Supplementary experimental procedures
 Supplementary experimental procedures Glucose tolerance, insulin secretion and insulin tolerance tests Glucose tolerance test (GTT) and glucose stimulated insulin secretion (GSIS) was measured in over-night
Supplementary experimental procedures Glucose tolerance, insulin secretion and insulin tolerance tests Glucose tolerance test (GTT) and glucose stimulated insulin secretion (GSIS) was measured in over-night
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell and patient-derived tumor organoids
 Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell and patient-derived tumor organoids Ling Huang 1, Audrey Holtzinger 1,2,11, Ishaan Jagan 1,11, Michael BeGora 1, Ines
Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell and patient-derived tumor organoids Ling Huang 1, Audrey Holtzinger 1,2,11, Ishaan Jagan 1,11, Michael BeGora 1, Ines
Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease
 1 Supplemental Materials 2 3 Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease 4 5 6 Authors: Madhumita Basu, 1 Jun-Yi Zhu, 2 Stephanie LaHaye 1,3, Uddalak
1 Supplemental Materials 2 3 Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease 4 5 6 Authors: Madhumita Basu, 1 Jun-Yi Zhu, 2 Stephanie LaHaye 1,3, Uddalak
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10188 Supplementary Figure 1. Embryonic epicardial genes are down-regulated from midgestation stages and barely detectable post-natally. Real time qrt-pcr revealed a significant down-regulation
doi:10.1038/nature10188 Supplementary Figure 1. Embryonic epicardial genes are down-regulated from midgestation stages and barely detectable post-natally. Real time qrt-pcr revealed a significant down-regulation
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY INFORMATION
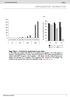 b 350 300 250 200 150 100 50 0 E0 E10 E50 E0 E10 E50 E0 E10 E50 E0 E10 E50 Number of organoids per well 350 300 250 200 150 100 50 0 R0 R50 R100 R500 1st 2nd 3rd Noggin 100 ng/ml Noggin 10 ng/ml Noggin
b 350 300 250 200 150 100 50 0 E0 E10 E50 E0 E10 E50 E0 E10 E50 E0 E10 E50 Number of organoids per well 350 300 250 200 150 100 50 0 R0 R50 R100 R500 1st 2nd 3rd Noggin 100 ng/ml Noggin 10 ng/ml Noggin
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with
 Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
Supplementary Figure 1. Repression of hepcidin expression in the liver of mice treated with DMN Immunohistochemistry for hepcidin and H&E staining (left). qrt-pcr assays for hepcidin in the liver (right).
Avrahami et al., Suppression of p57kip2 allows successful replication of adult human -cells
 Avrahami et al., Suppression of p57kip2 allows successful replication of adult human -cells Legends to Supplementary Figures Supplementary Figure 1. A. Suppression of p57 Kip2 in human islets does not
Avrahami et al., Suppression of p57kip2 allows successful replication of adult human -cells Legends to Supplementary Figures Supplementary Figure 1. A. Suppression of p57 Kip2 in human islets does not
Inhibition of DYRK1A stimulates human beta-cell proliferation
 Inhibition of DYRK1A stimulates human beta-cell proliferation Ercument Dirice 1,, Deepika Walpita 2,, Amedeo Vetere 2, Bennett C. Meier 2,5, Sevim Kahraman 1, Jiang Hu 1, Vlado Dančík 2, Sean M. Burns
Inhibition of DYRK1A stimulates human beta-cell proliferation Ercument Dirice 1,, Deepika Walpita 2,, Amedeo Vetere 2, Bennett C. Meier 2,5, Sevim Kahraman 1, Jiang Hu 1, Vlado Dančík 2, Sean M. Burns
Supplementary Figure 1
 Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
Supplementary Figure 1. SDS-FRL localization of CB 1 in the distal CA3 area of the rat hippocampus. (a-d) Axon terminals (t) in stratum pyramidale
 Supplementary Figure 1. SDS-FRL localization of CB 1 in the distal CA3 area of the rat hippocampus. (a-d) Axon terminals (t) in stratum pyramidale (b) show stronger immunolabeling for CB 1 than those in
Supplementary Figure 1. SDS-FRL localization of CB 1 in the distal CA3 area of the rat hippocampus. (a-d) Axon terminals (t) in stratum pyramidale (b) show stronger immunolabeling for CB 1 than those in
Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse
 Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
Transcription factor expression in the developing human fetal endocrine pancreas
 Diabetologia (8) 5:9 8 DOI.7/s5-8--z ARTICLE Transcription factor expression in the developing human fetal endocrine pancreas B. M. Lyttle & J. Li & M. Krishnamurthy & F. Fellows & M. B. Wheeler & C. G.
Diabetologia (8) 5:9 8 DOI.7/s5-8--z ARTICLE Transcription factor expression in the developing human fetal endocrine pancreas B. M. Lyttle & J. Li & M. Krishnamurthy & F. Fellows & M. B. Wheeler & C. G.
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Subnuclear Distribution of Progesterone Receptors A and B in Normal and Malignant Endometrium
 0021-972X/04/$15.00/0 The Journal of Clinical Endocrinology & Metabolism 89(3):1429 1442 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/jc.2003-031111 Subnuclear Distribution of
0021-972X/04/$15.00/0 The Journal of Clinical Endocrinology & Metabolism 89(3):1429 1442 Printed in U.S.A. Copyright 2004 by The Endocrine Society doi: 10.1210/jc.2003-031111 Subnuclear Distribution of
Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the
 Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the location of the transmembrane (TM), FRM binding (FB)
Supplementary Figure 1 Expression of Crb3 in mouse sciatic nerve: biochemical analysis (a) Schematic of Crb3 isoforms, ERLI and CLPI, indicating the location of the transmembrane (TM), FRM binding (FB)
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Table 1. Antibodies and dilutions used in the immunohistochemical study
 Supplementary Table 1. Antibodies and dilutions used in the immunohistochemical study Antigen Species Clone Source Dilution Insulin Guinea pig - Dako, Carpinteria, 1:200 Glucagon Rabbit - Dako, Carpinteria,
Supplementary Table 1. Antibodies and dilutions used in the immunohistochemical study Antigen Species Clone Source Dilution Insulin Guinea pig - Dako, Carpinteria, 1:200 Glucagon Rabbit - Dako, Carpinteria,
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung
 Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supplementary Figure 1. A. Bar graph representing the expression levels of the 19 indicated genes in the microarrays analyses comparing human lung immortalized broncho-epithelial cells (AALE cells) expressing
Supporting Information
 Supporting Information Pang et al. 10.1073/pnas.1322009111 SI Materials and Methods ELISAs. These assays were performed as previously described (1). ELISA plates (MaxiSorp Nunc; Thermo Fisher Scientific)
Supporting Information Pang et al. 10.1073/pnas.1322009111 SI Materials and Methods ELISAs. These assays were performed as previously described (1). ELISA plates (MaxiSorp Nunc; Thermo Fisher Scientific)
