Supplemental Figure S1. Sequence feature and phylogenetic analysis of GmZF351. (A) Amino acid sequence alignment of GmZF351, AtTZF1, SOMNUS, AtTZF5,
|
|
|
- Carmella Adams
- 5 years ago
- Views:
Transcription
1 Supplemental Figure S. Sequence feature and phylogenetic analysis of GmZF35. (A) Amino acid sequence alignment of GmZF35, AtTZF, SOMNUS, AtTZF5, and OsTZF. Characters with black background indicate conserved amino acids. () Phylogenetic analysis of GmZF35 and Arabidopsis TZF proteins.
2 6 3 Gm mzf35 expression (rela ative to At4g59) 4 Gm mzf35 expression (rela ative to AtACTIN) C Total fatty acids in seeds (%) 4 3 GmZF35-transgenic Arabidopsis plants ** ** ** * ** ** ** ** * ** GmZF35-transgenic Arabidopsis plants GmZF35-transgenic Arabidopsis plants Supplemental Figure S. Expression level of GmZF35 and total fatty acids contents of seeds in GmZF35-transgenic Arabidopsis plants. (A) Expression level of GmZF35 in Col- and GmZF35-transgenic Arabidopsis plants. At4g59 is used as internal control. Error bars indicate SD (n = 3). () Expression level of GmZF35 in Col- and GmZF35-transgenic Arabidopsis plants. AtACTIN is used as internal control. Error bars indicate SD (n = 3). (C) Total fatty acids contents in seeds of Col- and GmZF35-transgenic Arabidopsis plants. Asterisks indicate a significant difference compared with Col- (*P<.5 and **P<.). Error bars indicate SD (n = 4).
3 A Col- OE-3 OE-9 Col- OE-9 OE-3 Supplemental Figure S3. Morphology of GmZF35-transgenic Arabidopsis. (A) Rosettes of 3-week-old Col- and GmZF35-transgenic Arabidopsis plants (OE-3, OE-9 and ). () Rosettes of 5-week-old Col- and GmZF35-transgenic Arabidopsis plants (OE-3, OE-9 and ).
4 At3g96 F F F3 F4 F5 FaTA ChIP signal (%input).4. No Ab Anti-FLAG F F F3 F4 F5 FaTA KASIII Supplemental Figure S4. GmZF35 ChIP analysis of the At3g96 promoter and FaTA promoter. (A) The schematic diagrams depict the promoters of At3g96 and FaTA. At3g96 encoding Pkpα is regulated by WRI, and its promoter is not enriched in GmZF35 ChIP-Seq analysis. The right panel represents various fragments (F to F5, black rectangles) used for qpcr in kb promoter region (black line) of At3g96. The left panel represents the location of the enriched interval (black rectangle) identified by ChIP-Seq in kb promoter region (black line) of FaTA. The blue rectangles indicate CDS regions of At3g96 and FaTA. The primer sequences are shown in Supplemental table S4. () The graph indicates the percentages of DNA fragments that coimmunoprecipitated with FLAG antibody relative to the input DNAs in Col- and GmZF35-transgenic Arabidopsis plants (OE-3, OE-9, ). The no Ab samples are coimmunoprecipitated without antibody and served as negative control. The KASIII promoter fragment identified as a GmZF35 binding site is used as a positive control. Error bars indicate SD (n = 3).
5 C E G H Relative expression ative expression Rel Relativ ve expression expression Relative pression Relative ex CCP 4 TAG 3 Col- OE-3 OE-9 Relative expression D ative expression Col- OE-3 OE WRI Col- OE-3 OE-9 F Rel ve expression Relati KASIII Col- OE-3 OE-9 OLEO Col- OE-3 OE-9 Pkpα Pkpβ Col- OE-3 OE-9 Col- wri OE-9 GmZF35-OE wri PKpα PKpβ CCP KASIII TAG OLEO SUS ACP KASI Col- wri OE-9 GmZF35-OE wri PKpα PKpβ CCP KASIII TAG OLEO SUS ACP KASI Supplemental Figure S5. Expression levels of GmZF35 downstream genes in various Arabidopsis plants. (A-F) Expression levels of CCP (A), KASIII (), TAG (C), OLEO (D), WRI (E), PKpα and PKpβ (F) in 5-day-old seedlings of Col- and GmZF35-transgenic Arabidopsis plants. AtACTIN is used as internal control. Error bars indicate SD (n = 3). (G) Expression levels of PKpα, PKpβ, CCP, KASIII, TAG, OLEO, SUS, ACP and KASI in 5-day-old seedlings of Col-, wri, GmZF35-OE (OE-9 and ) and GmZF35-OE wri plants. AtACTIN is used as internal control. Error bars indicate SD (n = 3). For all panels, relative mrna levels of each gene in Col- are set to. (H) Expression levels of PKpα, PKpβ, CCP, KASIII, TAG, OLEO, SUS, ACP and KASI in 5-day-old seedlings of Col-, wri, GmZF35-OE (OE-9 and ) and GmZF35-OE wri plants. At4g59 is used as internal control. Error bars indicate SD (n = 3). For all panels, relative mrna levels of each gene in Col- are set to.
6 GmZF35 relative ex xpression.5..5 seeds (%) Total fatty acids in 4 3 * ** ** ** * ** GmZF35-transgenic soybean plants GmZF35-transgenic soybean plants Supplemental Figure S6. Expression level of GmZF35 and total fatty acids contents of seeds in GmZF35-transgenic soybean plants. (A) Expression level of GmZF35 in leaves of Jack, vector control and GmZF35-transgenic soybean plants. Error bars indicate SD (n = 3). () Total fatty acids contents in seeds of Jack, vector control and GmZF35-transgenic soybean plants. Asterisks indicate a significant difference compared with Jack (*P<.5 and **P<.). Error bars indicate SD (n = 4).
7 A Jack Vector control OE-34 OE-4 OE-73 Jack Vector control OE-34 OE-4 OE-73 Supplemental Figure S7. Morphology of GmZF35-transgenic soybean plants. (A) Morphology of transgenic soybean plants in V stage. Jack, vector control and GmZF35transgenic soybean plants (OE-34, OE-4 and OE-73) are observed. () Morphology of transgenic soybean plants in R3 stage. Jack, vector control and GmZF35-transgenic soybean plants (OE34, 34 OE-4 OE 4 and OE-73) OE 73) are observed. obser ed
8 Relative exp pression Relative expressio on Glyma5g3477 (GmWRI-like) Glyma5g44 (GmWRI-like) H H H3 H4 H5 H6 H H H3 H4 H5 H6 6.6 Glyma9g353 (GmCCP-like). Glyma5g55 (GmKASIII-like) H H H3 H4 H5 H6 H H H3 H4 H5 H6 Relative exp pression Relative expressio on ive expression Relati Relative exp pression Glyma8g4435 (GmKASIII-like) Glyma7g6 (GmTAG-like) Relative exp pression Relativ ve expression Glyma9g438 (GmKASIII-like3) H H H3 H4 H5 H6 H H H3 H4 H5 H6. Glyma3g656 (GmTAG-like) H H H3 H4 H5 H6 H H H3 H4 H5 H6.. Relative expression 3 Glyma9g36 (GmOLEO-like) Relative expression 5 5 Glyma6g78 (GmOLEO-like) H H H3 H4 H5 H6 H H H3 H4 H5 H6 Supplemental Figure S8. Expression pattern of potential downstream genes in six developmental stages of soybean seeds. The gray lines indicate WRI homologs, the blue line indicates CCP homolog, the green lines indicate KASIII homologs, the purple lines indicate TAG homologs, and the orange lines indicate OLEO homologs. Error bars indicate SD (n = 3).
9 .5 Jack Vector control OE-34 OE-4 OE-73 Relative expr ression.5 Glyma8g4435 (GmKASIII-like) Glyma9g438 (GmKASIII-like3) C Relative luminescenc ce Relative luminescenc ce Supplemental Figure S9. Overexpression of GmZF35 does not influence the expression of Glyma8g4435 and Glyma9g438. (A) The relative expression of Glyma8g4435 and Glyma9g438 are unchanged in GmZF35- transgenic soybean plants compared with Jack and vector control plants. The mrna levels (relative to GmTUULIN) of each gene in Jack are set to. Error bars indicate SD (n = 3). () Inclusion of 35S:GmZF35 effector can not activate the expression of Glyma8g4435 and Glyma9g438 by transient expression in tobacco leaves. The A. tumefacien harboring reporter vector is mixed with A. tumefacien harboring 35S:GmZF35-FLAG and infiltrated into tobacco leaves. A. tumefacien harboring pgw4 is used as negative control. (C) Quantitative analysis of luminescence intensity in (). LUC image is taken days after infiltration. Luminescence intensity is analyzed with IndiGo software. Error bars indicate SD (n =5).
10 3 GmZF35 GsZF35 Relative ex xpression HF47 ZYD3 Supplemental Figure S. Expression of GmZF35 and GsZF35 in seeds of HF47 and ZYD3 as determined by TaqMan RT-qPCR. GmZF35 or GsZF35 can not be detected in ZYD3 or HF47, respectively. The expression of GsZF35 allele in ZYD3 seed is set to. GmTUULIN is used as internal control. Error bars indicate SD (n = 3).
11 Total fatty acid ds in GsZF35- transgenic Arabi dopsis seeds (%) 4 3 ** Col- Fatty acids in GsZF F35-transgenic Arabidopisis seeds (%) ** ** ** 5 OE- OE-6 OE-5 35S:GsZF35 5 Col- OE- OE-6 OE-5 ** * ** ** ** ** ** ** ** ** ** ** ** **** ** ** ** ** ** ** ** **** 6: 8: 8: 8: 8:3 : Supplemental Figure S. Function of GsZF35 from ZYD3 on seed oil accumulation. (A) Total fatty acids contents in seeds of GsZF35-transgenic Arabidopsis plants (OE-, OE-6 and OE- 5). GmZF35-transgenic Arabidopsis plants () is used as positive control. Error bars indicate SD (n = 4). () Contents of fatty acid compositions in seeds of GsZF35-transgenic plants. GmZF35- transgenic Arabidopsis plants () is used as positive control. Error bars indicate SD (n = 4). For two panels, asterisks indicate a significant difference compared to Col- (*P<.5 and **P<.).
12 ds in SOMNUSabidopsis seeds (%) Total fatty aci overexpressing Ara 4 3 ** ** Supplemental Figure S. Overexpression of SOMNUS/TZF4 improves seed oil accumulation in transgenic Arabidopsis seeds. Total fatty acids contents in seeds of Col- and SOMNUS-overexpressing Arabidopsis plants (TZF4 OE and TZF4 OE) are measured. Error bars indicate SD (n = 4). Asterisk indicates a significant difference compared to Col- (**P<.).
13 A Col- OE-3 Jack Vector control OE-34 OE-9 OE-4 OE-73 Supplemental Figure S3. Morphology of GmZF35-transgenic Arabidopsis seeds and GmZF35-transgenic soybean seeds. (A) Morphology of seeds from Col- and GmZF35-transgenic Arabidopsis plants (OE-3, OE-9 and ). ar =.5 mm. () Morphology of seeds from Jack, vector control and GmZF35transgenic soybean plants (OE-34, OE-4 and OE-73). seeds are used for photographing. ar = cm.
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplemental Information. Spatial Auxin Signaling. Controls Leaf Flattening in Arabidopsis
 Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Supplementary Figures
 Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Supplementary Figures a miel1-2 (SALK_41369).1kb miel1-1 (SALK_978) b TUB MIEL1 Supplementary Figure 1. MIEL1 expression in miel1 mutant and S:MIEL1-MYC transgenic plants. (a) Mapping of the T-DNA insertion
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Potential use of High iron and low phytate GM rice and their Bio-safety Assessment
 Potential use of High iron and low phytate GM rice and their Bio-safety Assessment Dr. Karabi Datta University of Calcutta, India Background High iron rice and iron bioavailability Micronutrient deficiency
Potential use of High iron and low phytate GM rice and their Bio-safety Assessment Dr. Karabi Datta University of Calcutta, India Background High iron rice and iron bioavailability Micronutrient deficiency
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus
 Supplementary figures for: A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus officinalis Daisuke Tsugama, Kohei Matsuyama, Mayui Ide, Masato Hayashi, Kaien Fujino, and
Supplementary figures for: A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus officinalis Daisuke Tsugama, Kohei Matsuyama, Mayui Ide, Masato Hayashi, Kaien Fujino, and
Supplemental Fig. 1. Relative mrna Expression. Relative mrna Expression WT KO WT KO RT 4 0 C
 Supplemental Fig. 1 A 1.5 1..5 Hdac11 (ibat) n=4 n=4 n=4 n=4 n=4 n=4 n=4 n=4 WT KO WT KO WT KO WT KO RT 4 C RT 4 C Supplemental Figure 1. Hdac11 mrna is undetectable in KO adipose tissue. Quantitative
Supplemental Fig. 1 A 1.5 1..5 Hdac11 (ibat) n=4 n=4 n=4 n=4 n=4 n=4 n=4 n=4 WT KO WT KO WT KO WT KO RT 4 C RT 4 C Supplemental Figure 1. Hdac11 mrna is undetectable in KO adipose tissue. Quantitative
Soybean GmMYB73 promotes lipid accumulation in transgenic plants
 Liu et al. BMC Plant Biology 2014, 14:73 RESEARCH ARTICLE Open Access Soybean GmMYB73 promotes lipid accumulation in transgenic plants Yun-Feng Liu 1, Qing-Tian Li 1, Xiang Lu 1, Qing-Xin Song 1, Sin-Man
Liu et al. BMC Plant Biology 2014, 14:73 RESEARCH ARTICLE Open Access Soybean GmMYB73 promotes lipid accumulation in transgenic plants Yun-Feng Liu 1, Qing-Tian Li 1, Xiang Lu 1, Qing-Xin Song 1, Sin-Man
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
Supplementary Figure S1. Generation of LSL-EZH2 conditional transgenic mice.
 Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
The N-end rule pathway regulates pathogen responses. in plants
 SUPPLEMENTARY INFORMATION The N-end rule pathway regulates pathogen responses in plants Rémi de Marchi 1,2, Maud Sorel 1, Brian Mooney 1, Isabelle Fudal 3, Kevin Goslin 1, Kamila Kwaśniewska 4, Patrick
SUPPLEMENTARY INFORMATION The N-end rule pathway regulates pathogen responses in plants Rémi de Marchi 1,2, Maud Sorel 1, Brian Mooney 1, Isabelle Fudal 3, Kevin Goslin 1, Kamila Kwaśniewska 4, Patrick
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Light triggers PILS-dependent reduction in nuclear auxin signalling for growth transition
 In the format provided y the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 217.15 Light triggers PILS-dependent reduction in nuclear auxin signalling for growth transition Chloé
In the format provided y the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 3 ARTICLE NUMBER: 217.15 Light triggers PILS-dependent reduction in nuclear auxin signalling for growth transition Chloé
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Regulation of Floral Organ Identity. Dr. Chloe Diamond Mara
 Regulation of Floral Organ Identity Dr. Chloe Diamond Mara Flower Development Angiosperms (flowering plants) are the most widespread group of land plants Flowers are the reproductive organs that consist
Regulation of Floral Organ Identity Dr. Chloe Diamond Mara Flower Development Angiosperms (flowering plants) are the most widespread group of land plants Flowers are the reproductive organs that consist
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supporting Information
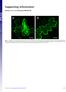 Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Supporting Information Harries et al. 1.173/pnas.9923916 A Fig. S1. Disruption of microfilaments within epidermal cells after treatment with 5 M Lat. Images of N. benthamiana cells are from plants expressing
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplemental Material for:
 Supplemental Material for: Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu, Vijay G. Sankaran, Min Ni, Tobias F. Menne, Rishi V. Puram,
Supplemental Material for: Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu, Vijay G. Sankaran, Min Ni, Tobias F. Menne, Rishi V. Puram,
Molecular mechanism of the priming by jasmonic acid of specific dehydration stress response genes in Arabidopsis
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Publications in the Biological Sciences Papers in the Biological Sciences 2016 Molecular mechanism of the priming
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Publications in the Biological Sciences Papers in the Biological Sciences 2016 Molecular mechanism of the priming
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supporting Information
 Supporting Information Lee et al. 10.1073/pnas.0910950106 Fig. S1. Fe (A), Zn (B), Cu (C), and Mn (D) concentrations in flag leaves from WT, osnas3-1, and OsNAS3-antisense (AN-2) plants. Each measurement
Supporting Information Lee et al. 10.1073/pnas.0910950106 Fig. S1. Fe (A), Zn (B), Cu (C), and Mn (D) concentrations in flag leaves from WT, osnas3-1, and OsNAS3-antisense (AN-2) plants. Each measurement
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Figure S1 Expression of AHL gene family members in diploid (Ler Col) and triploid (Ler
 Supplemental material Supplemental figure legends Figure S Expression of AHL gene family members in diploid (Ler ) and triploid (Ler osd) seeds. AHLs from clade B are labelled with (I), and AHLs from clade
Supplemental material Supplemental figure legends Figure S Expression of AHL gene family members in diploid (Ler ) and triploid (Ler osd) seeds. AHLs from clade B are labelled with (I), and AHLs from clade
Supplemental Data. Wang et al. (2013). Plant Cell /tpc
 Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Supplemental Data. Beck et al. (2010). Plant Cell /tpc
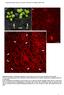 Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Supporting Information for
 Supporting Information for CuO Nanoparticle Interaction with Arabidopsis: Toxicity, Parent-Progeny Transfer and Gene Expression Zhenyu Wang,, Lina Xu, Jian Zhao,,, Xiangke Wang, Jason C. White, and Baoshan
Supporting Information for CuO Nanoparticle Interaction with Arabidopsis: Toxicity, Parent-Progeny Transfer and Gene Expression Zhenyu Wang,, Lina Xu, Jian Zhao,,, Xiangke Wang, Jason C. White, and Baoshan
Supplemental Figure 1. Small RNA size distribution from different soybean tissues.
 Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
Cloning and Expression of a Bacterial CGTase and Impacts on Phytoremediation. Sarah J. MacDonald Assistant Professor Missouri Valley College
 Cloning and Expression of a Bacterial CGTase and Impacts on Phytoremediation Sarah J. MacDonald Assistant Professor Missouri Valley College Phytoremediation of Organic Compounds Phytodegradation: Plants
Cloning and Expression of a Bacterial CGTase and Impacts on Phytoremediation Sarah J. MacDonald Assistant Professor Missouri Valley College Phytoremediation of Organic Compounds Phytodegradation: Plants
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Figure S1. (A) Schematic diagram of dnrar transgene allele. (B) X-Gal staining of testis from
 Figure S1. (A) Schematic diagram of dnrar transgene allele. (B) X-Gal staining of testis from germ cell mutants (dnrar flox/flox, Stra8-Cre +, RARElacZ) (A ), controls (dnrar flox/flox, RARElacZ) (B ),
Figure S1. (A) Schematic diagram of dnrar transgene allele. (B) X-Gal staining of testis from germ cell mutants (dnrar flox/flox, Stra8-Cre +, RARElacZ) (A ), controls (dnrar flox/flox, RARElacZ) (B ),
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Supplementary information
 Supplementary information Supplementary Figure 1: Components of Arabidopsis tricarboxylic acid (TCA) cycle. Schematic summary of the TCA cycle and the enzymes related to the reactions. The large text and
Supplementary information Supplementary Figure 1: Components of Arabidopsis tricarboxylic acid (TCA) cycle. Schematic summary of the TCA cycle and the enzymes related to the reactions. The large text and
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Supplementary Information
 Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
Supplementary Information Supplementary Figure 1! a! b! Nfatc1!! Nfatc1"! P1! P2! pa1! pa2! ex1! ex2! exons 3-9! ex1! ex11!!" #" Nfatc1A!!" Nfatc1B! #"!" Nfatc1C! #" DN1! DN2! DN1!!A! #A!!B! #B!!C! #C!!A!
DNA methylation Dots(+) CxxC EGFP DAP
 a 1.2 b 5mC PI Merge n level Relativ ve expressio 1.0 0.8 0.6 0.4 0.2 0 a b c Wildtype Gadd45 Gadd4 45b KO Figure S1. Gadd45b-deficiency does not affect paternal DNA demethylation (a) Relative expression
a 1.2 b 5mC PI Merge n level Relativ ve expressio 1.0 0.8 0.6 0.4 0.2 0 a b c Wildtype Gadd45 Gadd4 45b KO Figure S1. Gadd45b-deficiency does not affect paternal DNA demethylation (a) Relative expression
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation
 Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Supplementary Figure 1 ITGB1 and ITGA11 increase with evidence for heterodimers following HSC activation. (a) Time course of rat HSC activation indicated by the detection of -SMA and COL1 (log scale).
Nature Immunology: doi: /ni Supplementary Figure 1 33,312. Aire rep 1. Aire rep 2 # 44,325 # 44,055. Aire rep 1. Aire rep 2.
 a 33,312 b rep 1 rep 1 # 44,325 rep 2 # 44,055 [0-84] rep 2 [0-84] 1810043G02Rik Pfkl Dnmt3l Icosl rep 1 [0-165] rep 2 [0-165] Rps14 Cd74 Mir5107 Tcof1 rep 1 [0-69] rep 2 [0-68] Id3 E2f2 Asap3 rep 1 [0-141]
a 33,312 b rep 1 rep 1 # 44,325 rep 2 # 44,055 [0-84] rep 2 [0-84] 1810043G02Rik Pfkl Dnmt3l Icosl rep 1 [0-165] rep 2 [0-165] Rps14 Cd74 Mir5107 Tcof1 rep 1 [0-69] rep 2 [0-68] Id3 E2f2 Asap3 rep 1 [0-141]
Supplemental Data. Müller-Xing et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Phenotypes of iclf (clf-28 swn-7 CLF pro :CLF-GR) plants. A, Late rescue of iclf plants by renewed DEX treatment; senescent inflorescence with elongated siliques (arrow; 90 DAG,
Supplemental Figure 1. Phenotypes of iclf (clf-28 swn-7 CLF pro :CLF-GR) plants. A, Late rescue of iclf plants by renewed DEX treatment; senescent inflorescence with elongated siliques (arrow; 90 DAG,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Neocortex Zbtb20 / NFIA / Sox9
 Neocortex / NFIA / Sox9 Supplementary Figure 1. Expression of, NFIA, and Sox9 in the mouse neocortex at. The lower panels are higher magnification views of the oxed area. Arrowheads indicate triple-positive
Neocortex / NFIA / Sox9 Supplementary Figure 1. Expression of, NFIA, and Sox9 in the mouse neocortex at. The lower panels are higher magnification views of the oxed area. Arrowheads indicate triple-positive
Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70.
 Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70. peptide MH+ charge XCorr score Delta Cn score Protein ID KNAVVTVPAYFNDSQRQ 1680.83 2 2.22 0.37
Supplemental Table S1. The list of peptides identified in TAP-WPP1 LC/MS/MS analysis, corresponding to HSC70. peptide MH+ charge XCorr score Delta Cn score Protein ID KNAVVTVPAYFNDSQRQ 1680.83 2 2.22 0.37
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Peking-Yale Joint Center of Plant Molecular Genetics and Agrobiotechnology, College of Life Sciences, Peking University, Beijing , China b
 The Plant Cell, Vol. 20: 1437 1455, June 2008, www.plantcell.org ª 2008 American Society of Plant Biologists RESEARCH ARTICLES Arabidopsis DDB1-CUL4 ASSOCIATED FACTOR1 Forms a Nuclear E3 Ubiquitin Ligase
The Plant Cell, Vol. 20: 1437 1455, June 2008, www.plantcell.org ª 2008 American Society of Plant Biologists RESEARCH ARTICLES Arabidopsis DDB1-CUL4 ASSOCIATED FACTOR1 Forms a Nuclear E3 Ubiquitin Ligase
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning
 Lee_Jin Suk 1 Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning Jin Suk Lee, Takeshi Kuroha, Marketa Hnilova, Dmitriy Khatayevich, Masahiro M. Kanaoka, Jessica M. McAbee, Mehmet
Lee_Jin Suk 1 Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning Jin Suk Lee, Takeshi Kuroha, Marketa Hnilova, Dmitriy Khatayevich, Masahiro M. Kanaoka, Jessica M. McAbee, Mehmet
Evaluating Variability of Allergens in Commodity Crops (Peanuts)
 Evaluating Variability of Allergens in Commodity Crops (Peanuts) Peggy Ozias-Akins Laura Ramos Paola Faustinelli Ye Chu University of Georgia Manel Jordana Katherine Arias McMaster University Soheila Maleki
Evaluating Variability of Allergens in Commodity Crops (Peanuts) Peggy Ozias-Akins Laura Ramos Paola Faustinelli Ye Chu University of Georgia Manel Jordana Katherine Arias McMaster University Soheila Maleki
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
RNA-Seq Atlas of Glycine max: A guide to the Soybean Transcriptome
 RNA-Seq Atlas of Glycine max: A guide to the Soybean Transcriptome How do novel structures or functions evolve? A more relevant example Symbiosis and Nitrogen Fixation Limpens & Bisseling (23) Curr. Opin.
RNA-Seq Atlas of Glycine max: A guide to the Soybean Transcriptome How do novel structures or functions evolve? A more relevant example Symbiosis and Nitrogen Fixation Limpens & Bisseling (23) Curr. Opin.
Figure legends of supplementary figures
 Figure legends of supplementary figures Figure 1. Phenotypic analysis of rice early flowering1 () plants and enhanced expression of floral identity genes in.. Leaf emergence of,, and plants with complementary
Figure legends of supplementary figures Figure 1. Phenotypic analysis of rice early flowering1 () plants and enhanced expression of floral identity genes in.. Leaf emergence of,, and plants with complementary
Supplemental Data. Di Giorgio et al. (2016). Plant Cell /tpc
 Supplemental Figure 1. Synteny analysis of NIP4;1 and NIP4;2. Examination of the NIP4;1-NIP4;2 region in rabidopsis thaliana and equivalent evolutionary regions in other dicot genomes are shown on the
Supplemental Figure 1. Synteny analysis of NIP4;1 and NIP4;2. Examination of the NIP4;1-NIP4;2 region in rabidopsis thaliana and equivalent evolutionary regions in other dicot genomes are shown on the
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A.
 Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
Supplementary Figure 1. Genotyping strategies for Mcm3 +/+, Mcm3 +/Lox and Mcm3 +/- mice and luciferase activity in Mcm3 +/Lox mice. A. Upper part, three-primer PCR strategy at the Mcm3 locus yielding
SUPPLEMENTARY FIGURES AND TABLE
 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
Supplemental Data. Wu et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
The Ufm1-activating enzyme Uba5 is indispensable for erythroid differentiation in mice
 Supplementary information The Ufm1-activating enzyme Uba5 is indispensable for erythroid differentiation in mice Kanako Tatsumi 1, 2, Harumi Yamamoto-Mukai 2, Ritsuko Shimizu 3, Satoshi Waguri 4, Yu-Shin
Supplementary information The Ufm1-activating enzyme Uba5 is indispensable for erythroid differentiation in mice Kanako Tatsumi 1, 2, Harumi Yamamoto-Mukai 2, Ritsuko Shimizu 3, Satoshi Waguri 4, Yu-Shin
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplemental Data. Candat et al. Plant Cell (2014) /tpc Cytosol. Nucleus. Mitochondria. Plastid. Peroxisome. Endomembrane system
 Cytosol Nucleus 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT
Cytosol Nucleus 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 PSORT MultiLoc YLoc SubLoc BaCelLo WoLF PSORT
Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples.
 Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary Figures
 Supplementary Figures 9 10 11 Supplementary Figure 1. Old plants are more resistant to insect herbivores than young plants. (a) Image of young (1-day-old, 1D) and old (-day-old, D) plants of Arabidopsis
Supplementary Figures 9 10 11 Supplementary Figure 1. Old plants are more resistant to insect herbivores than young plants. (a) Image of young (1-day-old, 1D) and old (-day-old, D) plants of Arabidopsis
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION
 Supplementary igure 1. The expression level of PP4 (At4g16860) is diminished in the rpp4 mutant (SK017569). The mna was extracted to analyze the expression level of PP4 using quantitative PC (qpc). Gene
Supplementary igure 1. The expression level of PP4 (At4g16860) is diminished in the rpp4 mutant (SK017569). The mna was extracted to analyze the expression level of PP4 using quantitative PC (qpc). Gene
ddm1a (PFG_3A-51065) ATG ddm1b (PFG_2B-60109) ATG osdrm2 (PFG_3A-04110) osdrm2 osdrm2 osdrm2
 Relative expression.6.5.4.3.2.1 OsDDM1a OsDDM1b OsDRM2 TG TG TG ddm1a (PFG_3-5165) P1 P3 ddm1b (PFG_2-619) P2 531bp 5233bp P1 P4 P3 P2 F1 R1 F1 TG TG TG F1 R1 C ddm1a -/- -/- ddm1b -/- +/- +/- -/- D (PFG_3-411)
Relative expression.6.5.4.3.2.1 OsDDM1a OsDDM1b OsDRM2 TG TG TG ddm1a (PFG_3-5165) P1 P3 ddm1b (PFG_2-619) P2 531bp 5233bp P1 P4 P3 P2 F1 R1 F1 TG TG TG F1 R1 C ddm1a -/- -/- ddm1b -/- +/- +/- -/- D (PFG_3-411)
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Supplemental Information
 Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplemental Figures Legends and Supplemental Figures. for. pirna-guided slicing of transposon transcripts enforces their transcriptional
 Supplemental Figures Legends and Supplemental Figures for pirn-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear pirn repertoire Kirsten-ndré
Supplemental Figures Legends and Supplemental Figures for pirn-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear pirn repertoire Kirsten-ndré
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
Nature Neuroscience: doi: /nn Supplementary Figure 1. Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites.
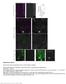 Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
s -1 (A) and (B) 2 s 1 in WT, riq1, and the riq1 mutant complemented by
 A 250-2 s -1 B 250-2 s -1 Supplemental Figure 1. The NPQ Phenotype in riq Mutants Was Complemented by Introduction of RIQ Genes. (A) and (B) 2 s 1 in, riq1, and the riq1 mutant complemented by the RIQ1
A 250-2 s -1 B 250-2 s -1 Supplemental Figure 1. The NPQ Phenotype in riq Mutants Was Complemented by Introduction of RIQ Genes. (A) and (B) 2 s 1 in, riq1, and the riq1 mutant complemented by the RIQ1
Rice in vivo RNA structurome reveals RNA secondary structure conservation and divergence in plants
 Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Generation and validation of mtef4-knockout mice.
 Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
Fig. S1. Weight of embryos during development. Embryos are collected at different time points (E12.5, E14.5, E16.5 and E18.5) from matings between
 Fig. S1. Weight of embryos during development. Embryos are collected at different time points (E12.5, E14.5, E16.5 and E18.5) from matings between Myod +/ or Myod / females and Myod +/ ;Igf2 +/ males and
Fig. S1. Weight of embryos during development. Embryos are collected at different time points (E12.5, E14.5, E16.5 and E18.5) from matings between Myod +/ or Myod / females and Myod +/ ;Igf2 +/ males and
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
Supplementary Figure S1 Targeted disruption and overexpression of Gpr43 in mice. (a) A targeting vector was constructed by ligation of 3 fragments:
 Supplementary Figure S1 Targeted disruption and overexpression of Gpr43 in mice. (a) A targeting vector was constructed by ligation of 3 fragments: the 5' and 3' homology recombination arms and a fragment
Supplementary Figure S1 Targeted disruption and overexpression of Gpr43 in mice. (a) A targeting vector was constructed by ligation of 3 fragments: the 5' and 3' homology recombination arms and a fragment
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
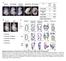 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
Supplementary Information and Figure legends
 Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
